External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G47090 133 / 3e-37
Leucine-rich repeat protein kinase family protein (.1)
AT3G47570 132 / 5e-37
Leucine-rich repeat protein kinase family protein (.1)
AT5G39390 127 / 7e-36
Leucine-rich repeat protein kinase family protein (.1)
AT5G20480 126 / 1e-34
EFR
EF-TU receptor (.1)
AT3G47110 125 / 1e-34
Leucine-rich repeat protein kinase family protein (.1)
AT3G47580 124 / 5e-34
Leucine-rich repeat protein kinase family protein (.1)
AT4G20140 95 / 1e-23
GSO1
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
AT5G44700 92 / 7e-23
GSO2, EDA23
GASSHO 2, EMBRYO SAC DEVELOPMENT ARREST 23, Leucine-rich repeat transmembrane protein kinase (.1)
AT5G46330 86 / 1e-20
FLS2
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT4G39400 81 / 5e-19
DWF2, CBB2, BIN1, BRI1, ATBRI1
DWARF 2, CABBAGE 2, BRASSINOSTEROID INSENSITIVE 1, BR INSENSITIVE 1, Leucine-rich receptor-like protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006445
202 / 1e-62
AT3G47570 468 / 1e-152
Leucine-rich repeat protein kinase family protein (.1)
Lus10011386
204 / 5e-62
AT3G47570 787 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10011388
203 / 8e-62
AT3G47570 757 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10011387
202 / 3e-61
AT3G47570 782 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10030903
182 / 1e-54
AT3G47570 798 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10016896
181 / 7e-54
AT3G47570 714 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10030845
179 / 3e-53
AT3G47570 796 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10014500
179 / 3e-53
AT3G47570 663 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Lus10030846
177 / 2e-52
AT3G47570 749 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10028479 pacid=23165974 polypeptide=Lus10028479 locus=Lus10028479.g ID=Lus10028479.BGIv1.0 annot-version=v1.0
ATGGCGGAATGTAAAGTATTGAAAAACATTAGGCACAAAAATCTTGTGAGGATAGTAACAGTATACTTAAGTGTTGATCATGAAGGGAATGATTTCAAGG
CTCTCATCTATGACTTCTTAGTTAGTGAAAGCTTAGAAGACTGGCTTCATCGGCCAATTGAAACTATAGATGAGTCGACAACAAAAAGCTTGAATTTCAC
CCAAAGGCTACATGTTGCTATTGAGATAGCTTTCAGAGTGGATTATCTTTACAACAATTGTGGAACGCCAATTGTTCATTGTGATCTCAAGCTTAGCAAT
GTACTTCTTGACAAGGATATAATTGCTCATGTTGGTGATTTTGGATTGGCACGATTCCTTTAA
AA sequence
>Lus10028479 pacid=23165974 polypeptide=Lus10028479 locus=Lus10028479.g ID=Lus10028479.BGIv1.0 annot-version=v1.0
MAECKVLKNIRHKNLVRIVTVYLSVDHEGNDFKALIYDFLVSESLEDWLHRPIETIDESTTKSLNFTQRLHVAIEIAFRVDYLYNNCGTPIVHCDLKLSN
VLLDKDIIAHVGDFGLARFL
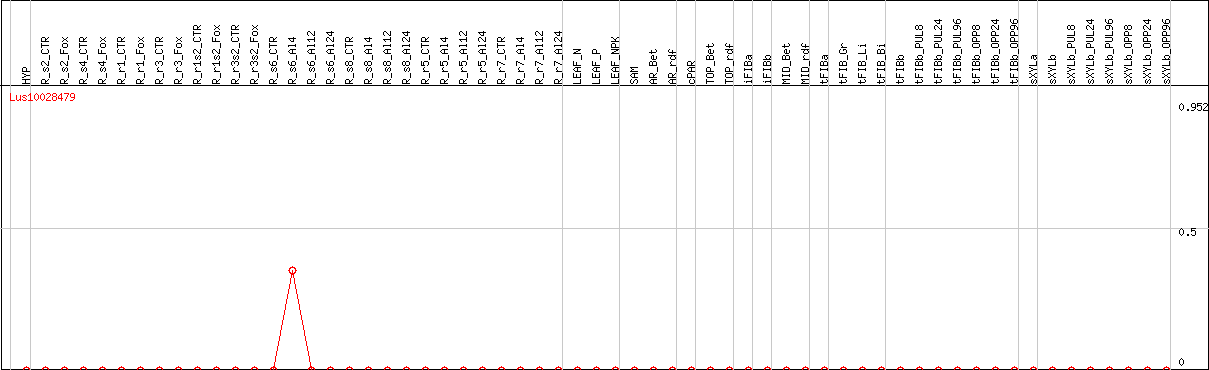
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028479 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.