External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G44080 160 / 2e-47
bZIP
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT1G03970 135 / 3e-38
bZIP
GBF4
G-box binding factor 4 (.1)
AT2G41070 60 / 2e-10
bZIP
DPBF4, ATBZIP12, EEL
ENHANCED EM LEVEL, Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
AT3G56850 59 / 5e-10
bZIP
DPBF3, AREB3
ABA-responsive element binding protein 3 (.1)
AT1G49720 59 / 8e-10
bZIP
ABF1
abscisic acid responsive element-binding factor 1 (.1.2)
AT2G36270 57 / 4e-09
bZIP
EEL, GIA1, ABI5
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT4G34000 55 / 2e-08
bZIP
AtABF3, ABF3, DPBF5
DC3 PROMOTER-BINDING FACTOR 5, abscisic acid responsive elements-binding factor 3 (.1.2.3)
AT1G45249 55 / 2e-08
bZIP
AtABF2, ATAREB1, AREB1, ABF2
ABSCISIC ACID RESPONSIVE ELEMENTS-BINDING PROTEIN 2, abscisic acid responsive elements-binding factor 2 (.1.2.3)
AT3G19290 54 / 4e-08
bZIP
AREB2, ABF4
ABA-RESPONSIVE ELEMENT BINDING PROTEIN 2, ABRE binding factor 4 (.1.2.3)
AT2G17770 49 / 9e-07
bZIP
FDP, ATBZIP27
FD PARALOG, basic region/leucine zipper motif 27 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018857
248 / 8e-83
AT5G44080 126 / 4e-35
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10029784
168 / 4e-50
AT5G44080 166 / 7e-49
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10042811
94 / 1e-22
AT5G44080 86 / 2e-19
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10028889
72 / 4e-14
AT3G56850 273 / 7e-91
ABA-responsive element binding protein 3 (.1)
Lus10022720
67 / 2e-12
AT2G36270 312 / 2e-103
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10017049
60 / 5e-10
AT2G36270 321 / 1e-106
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10021369
60 / 6e-10
AT2G36270 322 / 2e-107
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10014194
57 / 4e-09
AT2G36270 342 / 2e-114
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10002399
57 / 6e-09
AT3G19290 203 / 2e-61
ABA-RESPONSIVE ELEMENT BINDING PROTEIN 2, ABRE binding factor 4 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G192900
163 / 2e-48
AT5G44080 142 / 5e-40
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.002G067400
161 / 1e-47
AT5G44080 141 / 2e-39
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.008G010800
59 / 7e-10
AT3G56850 246 / 3e-80
ABA-responsive element binding protein 3 (.1)
Potri.010G248100
59 / 1e-09
AT3G56850 243 / 2e-79
ABA-responsive element binding protein 3 (.1)
Potri.002G090800
57 / 3e-09
AT3G56850 136 / 9e-38
ABA-responsive element binding protein 3 (.1)
Potri.009G101200
57 / 3e-09
AT4G34000 358 / 2e-120
DC3 PROMOTER-BINDING FACTOR 5, abscisic acid responsive elements-binding factor 3 (.1.2.3)
Potri.009G018500
56 / 4e-09
AT3G56850 160 / 1e-47
ABA-responsive element binding protein 3 (.1)
Potri.002G125400
54 / 3e-08
AT1G45249 306 / 4e-100
ABSCISIC ACID RESPONSIVE ELEMENTS-BINDING PROTEIN 2, abscisic acid responsive elements-binding factor 2 (.1.2.3)
Potri.001G220700
53 / 5e-08
AT3G56850 152 / 2e-44
ABA-responsive element binding protein 3 (.1)
Potri.009G164500
53 / 7e-08
AT3G44460 160 / 2e-46
basic leucine zipper transcription factor 67, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0018
bZIP
PF00170
bZIP_1
bZIP transcription factor
Representative CDS sequence
>Lus10028575 pacid=23164548 polypeptide=Lus10028575 locus=Lus10028575.g ID=Lus10028575.BGIv1.0 annot-version=v1.0
ATGGCGACGTCGAAGGTGATTTCGTCGTCAAAGGCGATGTCTTCATCGTCGAAACCGAAGGATTCCGATCCATCCGCTTCTTTTTCTTCAGAAGCTGCAC
GATCGCATCGATTTTCTTCCTCCGATCACCACCGCATCACAACCAACACCGGCAACGGCTTGTCAACGGCGATGACAGTCGACGGAGTGTTGCGCAGCGT
GTACTCAGCTCCTCCTCATCCATTATCGGAGTCCGCACTCCCAAACGATCACTTATCCCTGATTACCAGCAACGAGAACGGGTTTGCACCGAGCAAGACG
GCGGACGACATATGGAGGGATATAGTGTCGACAGGGAGGAAAGAACTGAAGGAGGAGGAGCCCGACGAGATGATGACGCTGGAGGATTTCCTGGCGAAGG
CTGGAACATGCGAGGAAGAGGATGGTGAAGATAATGAGGTGAAGATGATTCCTATGCTATCGTTTGATTCCGCGAATGCGATTCAAATGTTGGATAAGGT
GGAAGGCTCGATCGTCGGGTTGGGGAACAGTGGGAGAGGGAAGAGAGGTAGAGGTCCGGCAATGGAGCAGCAGTTGGATAAGGCTGCTCAGCAGAGGCAG
AAGAGGATGATCAAGAATCGGGAGTCGGCTGCTAGGTCCAGAGAGAGGAAGCAGGCTTATCAAGTGGAACTGGAGTCTTTGGCAGTGAAGCTTGAGGAGG
AGCATGAACAGCTACTGAAGGAAAGGGAAGAGCGTGCCAAGGAAAGGTTAAAGCAGATTATGGAGAATGTGGTGCCTGTCGAGGAGAGGGGAAGACCTCC
ACGAAAGCTTCAAAGAGCTCGATCCTGGACGTGGTAG
AA sequence
>Lus10028575 pacid=23164548 polypeptide=Lus10028575 locus=Lus10028575.g ID=Lus10028575.BGIv1.0 annot-version=v1.0
MATSKVISSSKAMSSSSKPKDSDPSASFSSEAARSHRFSSSDHHRITTNTGNGLSTAMTVDGVLRSVYSAPPHPLSESALPNDHLSLITSNENGFAPSKT
ADDIWRDIVSTGRKELKEEEPDEMMTLEDFLAKAGTCEEEDGEDNEVKMIPMLSFDSANAIQMLDKVEGSIVGLGNSGRGKRGRGPAMEQQLDKAAQQRQ
KRMIKNRESAARSRERKQAYQVELESLAVKLEEEHEQLLKEREERAKERLKQIMENVVPVEERGRPPRKLQRARSWTW
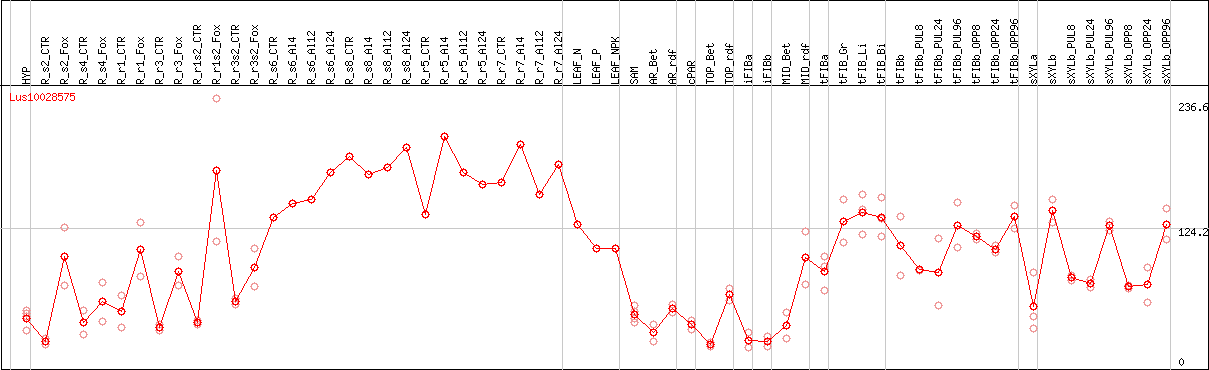
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028575 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.