Lus10028597 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10028597 pacid=23164506 polypeptide=Lus10028597 locus=Lus10028597.g ID=Lus10028597.BGIv1.0 annot-version=v1.0
ATGGAAGCGAATGCCGGCATGGTTGCCGGCTCCTACCGCCGGAACGAGCTCGTCCGGATTCGCCACGATTCCGACGGCGGGCCGAAACCACTGAAGAACT
TGAGTGGCCAAACGTGCCAGATCTGTGGTGATAGTGTTGGACTTACTGCCAATGGGGATGTATTCGTTGCTTGCAATGAATGTGCCTTCCCGGTTTGTCA
TCCTTGTTACGAGTACGAGCGAAAAGATGGGACCCAGGCTTGCCCCCAGTGCAAGACTCGGTACAAGAGGCACAAAGGGTGCCCTCGGGTTGATGGAGAT
GAGGATGAAGACGATGTTGATGATCTTGAAGACGAGTTCAGCTATCAAGCCACTGGGAAAGCTGGCCGACACTGGCAGGGTGATGATGTAGAGCTCTCTT
CCTCGTCGAGGCATGAATCTCAGATGCCGATTCCTCTTCTGACTAATGGCCAGCAGATTTCGGGTGAAATTCCATGCGCTACACCGGATACACAGTCTGT
TCGGACTACATCAGGCCCCTTAGGCCCTACTGACAAGAATGTACAGTCGTCTCCTTATCTCGATCCAAGGCAACCAGTTCCTGTTAGAATTGTGGATCCA
TCAAAGGACTTGAATTCCTATGGTCTTGGTAATGTGGACTGGAAGGAAAGAGTGGAGGGTTGGAAAATGAAGCAGGAGAAGGGAATGATGCATATGAGTA
ACCGACATAGTGAAGGAAAAGGAGACATTGAAGGCACAGGTTCAAATGGAGAAGAGCTTCAAATGGCTGACGATGCTCGGCAACCTATGAGCCGGGTGGT
CCCTATCCCATCTTCTCACATGACCCCATATCGCGTAGTCATCATTCTGCGATTCATCATCTTGTGTTTCTTCTTGCAATATCGTGTGACACACCCGGTG
AAGGATGCATACCCATTGTGGCTTATTTCCATCATTTGCGAGATCTGGTTTGCTTTGTCTTGGCTTCTGGATCAGTTCCCCAAATGGCAACCCATCAATC
GCGAGACTTATCTGGACAGGCTTGCACTGAGATTTGATCGTGAAGGGGAGCCATCGCAGCTAGCTCCTGTTGATGTGTTTGTCAGTACCGTGGATCCTAT
GAAAGAGCCTCCTCTAATCACTGCGAACACTGTCTTGTCCATACTTGCTGTGGATTACCCGGTTGACAAAGTATCATGCTATGTATCTGATGATGGATCA
GCAATGTTGTCATTTGAATCCCTCTCGGAGACTGCTGAATTCGCTAGGAAGTGGGTGCCTTTCTGCAAGAAGCACAATATAGAGCCTAGAGCTCCTGAGT
TCTACTTCGCCCAGAAGATAGATTACTTGAAGGACAAGATACAGCCTTCCTTTGTGAAAGAGAGACGAGCTATGAAGAGAGAATATGAAGAGTTCAAGGT
GCGGATCAATGCCCTTGTTGCTAAAGCACAAAAGATGCCTGAAGAAGGGTGGACAATGCAAGATGGCACTCCCTGGCCCGGGAATAATTCCAGGGACCAT
CCCGGAATGATTCAGGTCTTCCTAGGCCACAGTGGAGGACTCGATACCGATGGAAATGAACTGCCTCGGCTTGTGTACGTTTCTCGTGAAAAGAGACCGG
GATTCCAACATCACAAGAAAGCTGGAGCCATGAATGCATTGATTCGAGTTTCGGCTGTTTTGACCAATGGTGCCTATCTGCTGAATGTGGATTGTGATCA
TTACTTCAATAACAGCAAAGCTCTTAAAGAGGCAATGTGCTTCATGATGGATCCTGCATACGGAAAGAAGACATGCTATGTGCAGTTCCCACAACGTTTC
GATGGCATTGACTTGCACGATCGATATGCTAATCGCAATATTGTCTTCTTCGATATCAATTTGAAGGGGCTGGATGGTATTCAGGGACCGGTGTACGTCG
GTACTGGTTGTTGTTTCAACAGACAAGCATTATATGGTTACGACCCCGTTCTTACTGAAGAGGATCTAGAACCAAACATCGTTGTCAAGAGTTGTTGCGG
AACGAGAAAGAAGAGCAAGAGTGGGAGCAAAAAGTACTCTGACAAAAGAAAGAACATGAAGAGAACTGAATCATCAGTCCCGATGTTCAATCCAGAAGAC
ATTGAGGATGGTATCGAAGGATATGATGATGAGAGGTCACTCTTAATGTCTCAGAGAAGTCTCGAGAAGCGATTCGGTCAGTCACCAGTCTTTATCGCTG
CTACTTTCATGGAGATGGGAGGCATTCCACCGTCCACGAATCCCGCGACCCTTCTAAAGGAAGCTATCCATGTCATCAGCTGCGGGTATGAAGACAAGAC
TGAATGGGGCAAAGAGATCGGGTGGATCTACGGTTCGGTTACGGAAGATATTTTGACCGGTTTCAAGATGCATGCACGTGGTTGGATTTCAGTTTACTGC
ATGCCTCCACGCCCAGCGTTCAAGGGTTCTGCTCCAATCAATCTTTCTGATCGTTTGAACCAAGTTCTTCGGTGGGCCCTTGGATCGATTGAGATTTTGT
TGAGCAGGCATTGCCCTATATGGTATGGCTACAATGGGAAGTTAAAGCTCTTGGAGAGGCTCGCTTACATTAACACGATCGTCTATCCCATCACCTCTCT
TCCGCTGCTCGCCTACTGTATGCTTCCCGCGTTCTGTCTTCTCACGAACAAATTCATCATTCCCGAGATAAGCAACTTTGCCAGTATGTGGTTCATTCTC
CTTTTCGCATCCATTATCGTGACCGGAATCCTCGAGCTGAGGTGGAGCGGAGTCAGTCTGGAGGACTGGTGGAGGAACGAACAGTTCTGGGTCATTGGTG
GCACTTCGGCTCATCTCTTTGCTGTGTTCCAAGGTCTACTCAAGGTCCTCGCCGGAATCGACACCAACTTTACCGTCACTTCAAAGGCATCAGACGAGGA
TGGCGATTTCGCTGAGCTCTACCTCTTCAAATGGACCTCCTTGCTCATCCCTCCGACCACCGTCCTTATCGTGAACTTGGTAGGCATCGTGGCTGGAGTT
TCTTATGCCATAAACAGCGGATATCAGTCTTGGGGACCACTCTTTGGAAAGCTCTTCTTCGCCTTGTGGGTCGTCGCCCATTTGTATCCTTTCTTGAAAG
GTCTTTTGGGTCGCCAGAATCGTACACCTACCATTGTCATCGTCTGGTCGATTCTCCTCGCCTCGATATTCTCGTTACTGTGGGTGCGTATCGATCCTTT
CACTACCAACTCCACGAAGACTGTGTCGAATGGTCAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10028597 pacid=23164506 polypeptide=Lus10028597 locus=Lus10028597.g ID=Lus10028597.BGIv1.0 annot-version=v1.0
MEANAGMVAGSYRRNELVRIRHDSDGGPKPLKNLSGQTCQICGDSVGLTANGDVFVACNECAFPVCHPCYEYERKDGTQACPQCKTRYKRHKGCPRVDGD
EDEDDVDDLEDEFSYQATGKAGRHWQGDDVELSSSSRHESQMPIPLLTNGQQISGEIPCATPDTQSVRTTSGPLGPTDKNVQSSPYLDPRQPVPVRIVDP
SKDLNSYGLGNVDWKERVEGWKMKQEKGMMHMSNRHSEGKGDIEGTGSNGEELQMADDARQPMSRVVPIPSSHMTPYRVVIILRFIILCFFLQYRVTHPV
KDAYPLWLISIICEIWFALSWLLDQFPKWQPINRETYLDRLALRFDREGEPSQLAPVDVFVSTVDPMKEPPLITANTVLSILAVDYPVDKVSCYVSDDGS
AMLSFESLSETAEFARKWVPFCKKHNIEPRAPEFYFAQKIDYLKDKIQPSFVKERRAMKREYEEFKVRINALVAKAQKMPEEGWTMQDGTPWPGNNSRDH
PGMIQVFLGHSGGLDTDGNELPRLVYVSREKRPGFQHHKKAGAMNALIRVSAVLTNGAYLLNVDCDHYFNNSKALKEAMCFMMDPAYGKKTCYVQFPQRF
DGIDLHDRYANRNIVFFDINLKGLDGIQGPVYVGTGCCFNRQALYGYDPVLTEEDLEPNIVVKSCCGTRKKSKSGSKKYSDKRKNMKRTESSVPMFNPED
IEDGIEGYDDERSLLMSQRSLEKRFGQSPVFIAATFMEMGGIPPSTNPATLLKEAIHVISCGYEDKTEWGKEIGWIYGSVTEDILTGFKMHARGWISVYC
MPPRPAFKGSAPINLSDRLNQVLRWALGSIEILLSRHCPIWYGYNGKLKLLERLAYINTIVYPITSLPLLAYCMLPAFCLLTNKFIIPEISNFASMWFIL
LFASIIVTGILELRWSGVSLEDWWRNEQFWVIGGTSAHLFAVFQGLLKVLAGIDTNFTVTSKASDEDGDFAELYLFKWTSLLIPPTTVLIVNLVGIVAGV
SYAINSGYQSWGPLFGKLFFALWVVAHLYPFLKGLLGRQNRTPTIVIVWSILLASIFSLLWVRIDPFTTNSTKTVSNGQ
|
DESeq2's median of ratios [FLAX]
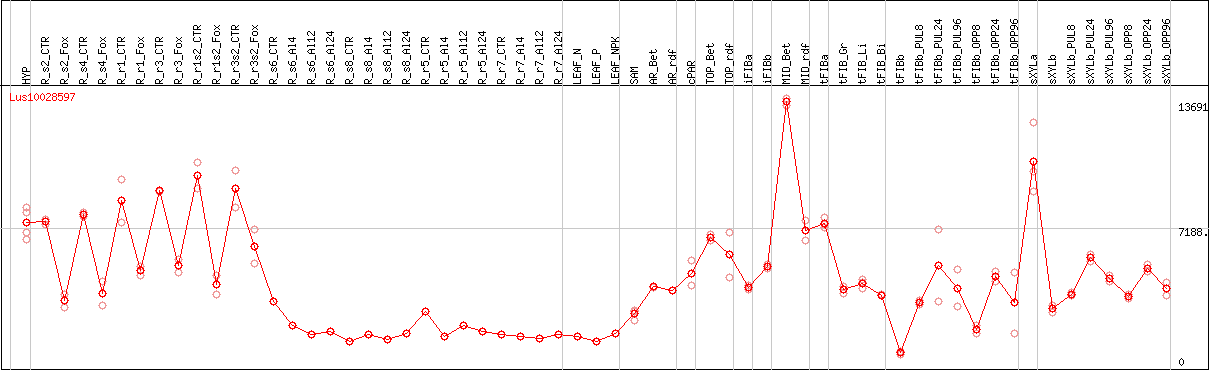
Coexpressed genes
Lus10028597 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
