Lus10028662 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10028662 pacid=23164553 polypeptide=Lus10028662 locus=Lus10028662.g ID=Lus10028662.BGIv1.0 annot-version=v1.0
ATGGGGTCGTGTGAGGATGCTGTTGTAGTCCACGACCAGGTTAATGCACCTGTAATCGGGCAGCATTCTTCCTGCTCGGAGTTCGATGAGGAATTTGGCG
CTGGAAGCTTGGCGGCTCCTGATGTGAATGCTACCAGTGCGGTGGGCTGCCTGGATCGGAAGGATAATGCGGGTAGTGTGAGTTCTAGAGACGAGTTAGA
GACTACATCTGTTGAAGGAGGGGTCGGGGAGACTTGTCCGAGTTTCACTGGCATGGGGTGTGTGGATTATAATGGGGATGCTGCCGGGGAATGTTCAGGG
AAGGTGGAGGAGAATGGAGGTGAGAGTGATGAATGGCTGAATGATGGACCTGCGGAATTGATGTCTCCTGATTTGGTTGATGGTTTGGGTTCAGGGAAGG
GTATGGAGGGTGATGAAAGTGGTGGTAATGTCCGTATTGATGTGGAAGAGGAGATGAGTGACTTCTTTGAAGGTGAGATGGAACGAGGCTTGCTTCCGTC
ACTGGGTAAGGATACAGATAGTCAATGCCGAAATTTGGATTCGTTACCTTCAGCATTGGGAACTGTGGAGAGTATGCGTCCTACTTGTATTCAAGTTGAT
AGCCGGAAAGACGAGCAGCCCGCTACTTCTTCTTCTTCCTCGAAAGGGTTAGCTGATGCTGCAGAAGGAAGAGGGGATGACTTAGAGGGGTTGGAGACCA
CTAGTGGGCAGTTGTCTTGGCAGGGTTCTTTGGAGTTGTCAAAGTTAAGCTTCAGAGCCGAGTCGGTGCAAAATGTTGTTCACGAGAATGTCGATTACCC
TGCTGATTATCCCCTTAAAGAAGATGTAGAAGGAATTAAGGAAGAAGAACATGTTCCTTTGGTATTTTCAGAGAATGTTCAAACTGGCCAAGCTTCATCT
TCAATGGAGCAAGTGAAGTGCACCGCAGGAAGTCCAGTGTCAACTGATTGTCCCAAACAATTTGACGATGGTGAAAGTGATGCTTCTGCCGACAGAGAAG
TTGATATATGTGCATGGATTGATGAGTTGCCTGTCAAAGAACCAGTAAAAAGTGTTGGTTTGATTGGTTATGAGAAGAAAACATATGAGGAGAATGACAG
ACCCGGTGCTTACCCTTTTATAAAGGTTCAAAGAAATAATGACGAAGTTAGCAGATTTGTCAATGTTGCACCGTTACTTGCTTGTGAGAGGTCACTGGAG
TATTTATCTACAGCCAGGTCGCTCAGTATTTGTAATCAACCGGATGGGGAAATAAATGAGAGGAACATAGACATACCTTCTGCAGAAAGCACTTCGGAAG
AGACTTTCGGTACTTTAACTGCTAAGATGGAAGATATCCGTCATAGTTTGTGCATGGAAGGAAACGACTGCTTTCTCAAGGAAGAAATGACCGAAATTCC
CACAGCTGTGTGTGTTAAGGAACTTGTTGTTTCTCCAACTCGACAGCCCTGTGAAAGTGAGGCTAAAAACCATGTAAGAACTGATTGTATTTCAAATACT
GAATTGCCACATAGTCAATCATATGCTCCCCTACAGAACAGCAGACCTTGCAGATCAAGTGGCAAGTCCCGGTCAACAAAGGGTAAAAATAAGGGCGAAA
AGGTAGCTCTGATACTGAGCTTTCAGAAGAGGAAAAGAAGTTTCCTACCTAAACGGACTCGAGTTTGTGAATGGGGATTACTGGGGAAAGTCATGGAATA
TTTCAAGGCGATTAATAAGCTGGGTGATGATGCAAAACAGGTACATAGATCACGAAAAGAAATGGCAAGACAAGGAAGGCAGAGGCAGAACAAGAATAAA
GCTGGCGGATCTGGTGGAGAAGGCCATACTTTTACTACTCGAATTCGTTTAAAAGTTAATGTGAGCAATGCTGTTAACACAGGTGTACCCATGGATGCTG
GTAATGCTACCATCAATGGTGACAGAGATGAAGCTATAGAACAGAAAAGAAAAGATCATCAGCGCAAGCAGAAAAATGTTACATCTGAGTCTGATGCTTC
TGTTGTAGATGCATGTTATCAGAACATCCAGCTTGAGGGAACTATATCTGATGAGAAATCAATCGAAGTTGCGGCAGGGGATTATCTTGATCCTGCTAAT
TTAGGGGGTTTGTTAGAAGTATCGAGAGAGAGGACATGTGCAGGTTCAGGAACTTCACCAGAATCAGAAGTAATTAATCTAGTACCTGAGGTTCAGGCTG
GATCACAATGTCTGCATCATATTCCTGATCGTGGTTCGGCGAACTTACAAATCTTGGCTTCTTCCGGGCTTCTTCCAAAAAATACTGCTGGGAAGAAAAA
GATGAAGAATAAGAAGAGTAAATTCCCTTGCGTAGATAATTGTTCCAAGGAAGATGACTTACCTGTTGCGGGAAGCACACACAAAGAAGCTAAATCAGCA
AAGAAAAAAAGCGTCAGGCAGAGGAAGTCAGAGAAATCTTCATCAATTGGCAAGGATGTTTCTATTGAACAGGTACCTTCTTCTGAAGAAACTGAACTGA
TGGTCCCCAGAGAAGCATGTGATGTTGATTCTTCAGTAAAAACTAAGGCGACTGGAACTCGGAGAACATCAGATGGATGTACTCAGAGAAGGTCTAAAAA
ATCATCTTCAGGGAAAGGCAGAAGATCAAGCAGTTCTATACAAAACTTGAACAGGGAGGGGATTGTTAGTAAAGTTGATGAAACTATTTGTGGTCCTTCT
GATAATGGAGTGGAAGTTGACACGGAGCAAGGTAACGGATTTGTTACGATCCTCTGCTTAGGATTTGCATCAGTTGGGACTATTCAAGAATCCACGAGTT
TAACTAATGTGGGAAGTGTGGATGATCTGGGCAAGGCATTGGATTCCGTACCAAAGCAGCTTTTGCCTGCGGATAGTGCTTGGGCGCACTGTGATAAGTG
TCATAAATGGAGACGTATTCCAGTTGAACTGGTACAGACAATTGGACAGAATGACCAAATATGGACATGCAAGGACAACGTGGATAAGGAGTACGCAGAT
TGTTCCATTCCCCAGGAGAAGTCCAATTCAGAAATTAATGCGGAGTTGGGCATATCAGATGGAGATGAAGATGCCTTCGATATACCTTCGAACAGTAAAG
GTCTAGAGAGTAGGCAGATAGTTTCCAAGGAACATGAGTTTACTCGGATTCGTACTAACCAGTTTATGCACCGTCGCCGGAAGACTCAAAACATTGATGA
GGTGATGATATGCCATTGTAGGCCACCAGTAGATGGAAGGCTTGGCTGTGGAGACGAATGCTTAAATCGAATGCTTAACATTGAGTGTGTTCAAGGCACT
TGCCCTTGTGGGGACGTCTGTTCAAATCAACAGTTCCAAAGACAAACTTATGCTAAGATGAGCTGGGGTCGATGTGGGAAAAAAGGGTTTGGGCTGCGGA
TGCAAGAAGATATTTCCAAAGGGAAATTTCTTATTGAGTACGTTGGAGAGGTGCTTGATATGCGAGCGTATGAGGCACGGCAAAAAGAGTATGCTGCTAA
GGGTCACAAACATTTCTACTTTATGACGTTAGATGGCAGTGAGGTTATAGATGCGTGTGCAAAAGGAAATCTGGGACGTTTCATTAACCATAGCTGTGAT
CCCAATTGCCGTACTGAAAAGTGGGTTGTAAATGGAGAAATTTGTATTGGAATGTTTGCAGTGAGGAATATTAAGAAGGGTGAAGAGCTGACATTTGACT
ACAACTATGTAAGGGTTTTTGGGGCCGCTGCCAAGAAATGTTATTGTGGCTCATCTCTGTGCCGCGGTTACATTGGTGGTGACCCAACTAACTATGAAGT
AATTGATCAAGTTGATTCCGATGAAGAGTTTCCTGAACCTGTGATGTTTGAGGAGGGGAGAAACGCCACGAGAAAAACACCTGCGAAAATGAAGCAGATT
GCAGAAAGTATTTCAGCGGAGAGGGATAATTTGGATGGCACTGTGGAATCTCTTGTTAAAGGGGACGTCTCTTCAAAAATTGGCACTGCTAGGAGTAAAT
CTCCTGATGTCTCTCAGTCTCTTGGTTCGTTGGAAGTGACTGATATAAAAACAAATTATCAACCTCGGCAGTCAGTAGAGATCTCTTGCCAAGCAGAAGA
AACAATGAGAATTCCCTCTGAGATTGAGAAAGATACTTCAAAGGAGGAAGCTGTTGATAAACCTACTCCTGCAACAACTGTGAAATCATCAAGTGACGGT
ACTGATGCAAAGAAGAAGACAAAATCTTCCGAAGATAAGCGGGTATTTGTGAAATCCCGTTTCCTAATTAAAGGTCAATCTGGATCCACTAAGAAAGGGA
AGCTTAATGTTATTCCTCAAACTGCAAATAAATCGACGGTGGCTAACAAATTGCCAGTGCAACCTGTGAATATTAAAAGGCCAATGGAAGGTGTCCCTAA
TGGTCGGCTTGAGACAGTTGAGGAGAAACTTAATGAGTTGTTGGATGATGAGGGTGGCATAACAAAGCGAAAAGATGCTGTTATTGGCTACCTAAAGCTT
CTTCTTCTTACCACTGCTTCGGGTGCTGGTGCGAATGGTGAAGCAATTCAAAGTAATCGAGATCTGTCCATGATCCTTGATGCAATCTTGAAAACCCGAT
CGCGAGCTGTCTTGATTGATATAATCAGCAAGAATGGTTTGCAGATGCTTCACAATATAATTAAGCGGTATAGATCGGACTTCAACAAGATTCCCATTGT
CCGTAAACTTCTAAAGGTCATAGAGTATTTGGCAGGGAGGGAGATACTCACAAACAACAATATATATGGCGGTCCTCCTCGCCCGGGGATGGAGAGCTTT
CGAGTATCTATGCTGTCCCTGACAGAGCATGATGATAGACAGGTACATCGAACTGCACGGAGTTTCAGAGACAAATGGTTTCCTCGGCAAAGATTCGAGG
TTAAACGTCACTCTTCTGATAAGGATGAGGGAAGGAATGAATTCCGAAGAGGATTCAACGACAACAAGACATCAACAATGCACAGTCTATCTCTCGATAA
CGGCCCAATCCAATCGAAGGTCGCATCAACTTCATCAGGCAGCGCAGTTGTACAGTCAGAGATTCCATCTAGTTCAGTGGACCGAAGTGTGAACGAAGTA
GGGTGTTCTTCTGCACTTTCCGACGAAAGGACCCGTAAAAGGAAAAGCCGATGGGATCAACCGTTTCGAATCAAACCTGACTCTAAGAAGACCGAATCAG
ATGTCAAAGTACCGGATGATGTGGACACATGCCATACAAATCTCGACGAGGATCTTCCACCCGGCTTCTCATCCAATCCGGGCTCCTCCGTGGTTTCCTG
TGACAGCTCACCGGCCCCGAAACCAAATGCCTGTGGGCTAAAGTGTCCTGTGGAAGAAGACATTAATGTAGGGGGTCCTCGGAAGAAGTTCGACTCTTGC
TTGCCTGTCTCGTTTGGAATCCCGTGGAACATTGTACAGCAATTCGGTACTCCTGCAGACGAAGCCGGAAGTAGTTGGGTTGTTGCTCCGGGGATACCGT
TTCGTCCATTCCCACCTTTACCGCCTACTCCCTACCAAAAGTTCGAAACCCCAGATTGCAAGACAGCGACTTTTAACAACGCTGTTGACTCGGCAAGCCG
GGTTGTTAGCTGCCGGAACGGAGGTTCGTTGGGGAAGAGGTACTTCAAGCAACAGAAGTGGAATCGGAGGACTCCGTGGGATTGGAAACACGACGGCAAT
CAGACCCGAAATGGAGATTGTGGCTCTTCTAACGGGAATGGAAGCTTACCGAGTCACGAGAACGCAAGTTCGTATACGAATACGAGTACTACTACTTCAC
AATCCAATGCACAGGCACCCAATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10028662 pacid=23164553 polypeptide=Lus10028662 locus=Lus10028662.g ID=Lus10028662.BGIv1.0 annot-version=v1.0
MGSCEDAVVVHDQVNAPVIGQHSSCSEFDEEFGAGSLAAPDVNATSAVGCLDRKDNAGSVSSRDELETTSVEGGVGETCPSFTGMGCVDYNGDAAGECSG
KVEENGGESDEWLNDGPAELMSPDLVDGLGSGKGMEGDESGGNVRIDVEEEMSDFFEGEMERGLLPSLGKDTDSQCRNLDSLPSALGTVESMRPTCIQVD
SRKDEQPATSSSSSKGLADAAEGRGDDLEGLETTSGQLSWQGSLELSKLSFRAESVQNVVHENVDYPADYPLKEDVEGIKEEEHVPLVFSENVQTGQASS
SMEQVKCTAGSPVSTDCPKQFDDGESDASADREVDICAWIDELPVKEPVKSVGLIGYEKKTYEENDRPGAYPFIKVQRNNDEVSRFVNVAPLLACERSLE
YLSTARSLSICNQPDGEINERNIDIPSAESTSEETFGTLTAKMEDIRHSLCMEGNDCFLKEEMTEIPTAVCVKELVVSPTRQPCESEAKNHVRTDCISNT
ELPHSQSYAPLQNSRPCRSSGKSRSTKGKNKGEKVALILSFQKRKRSFLPKRTRVCEWGLLGKVMEYFKAINKLGDDAKQVHRSRKEMARQGRQRQNKNK
AGGSGGEGHTFTTRIRLKVNVSNAVNTGVPMDAGNATINGDRDEAIEQKRKDHQRKQKNVTSESDASVVDACYQNIQLEGTISDEKSIEVAAGDYLDPAN
LGGLLEVSRERTCAGSGTSPESEVINLVPEVQAGSQCLHHIPDRGSANLQILASSGLLPKNTAGKKKMKNKKSKFPCVDNCSKEDDLPVAGSTHKEAKSA
KKKSVRQRKSEKSSSIGKDVSIEQVPSSEETELMVPREACDVDSSVKTKATGTRRTSDGCTQRRSKKSSSGKGRRSSSSIQNLNREGIVSKVDETICGPS
DNGVEVDTEQGNGFVTILCLGFASVGTIQESTSLTNVGSVDDLGKALDSVPKQLLPADSAWAHCDKCHKWRRIPVELVQTIGQNDQIWTCKDNVDKEYAD
CSIPQEKSNSEINAELGISDGDEDAFDIPSNSKGLESRQIVSKEHEFTRIRTNQFMHRRRKTQNIDEVMICHCRPPVDGRLGCGDECLNRMLNIECVQGT
CPCGDVCSNQQFQRQTYAKMSWGRCGKKGFGLRMQEDISKGKFLIEYVGEVLDMRAYEARQKEYAAKGHKHFYFMTLDGSEVIDACAKGNLGRFINHSCD
PNCRTEKWVVNGEICIGMFAVRNIKKGEELTFDYNYVRVFGAAAKKCYCGSSLCRGYIGGDPTNYEVIDQVDSDEEFPEPVMFEEGRNATRKTPAKMKQI
AESISAERDNLDGTVESLVKGDVSSKIGTARSKSPDVSQSLGSLEVTDIKTNYQPRQSVEISCQAEETMRIPSEIEKDTSKEEAVDKPTPATTVKSSSDG
TDAKKKTKSSEDKRVFVKSRFLIKGQSGSTKKGKLNVIPQTANKSTVANKLPVQPVNIKRPMEGVPNGRLETVEEKLNELLDDEGGITKRKDAVIGYLKL
LLLTTASGAGANGEAIQSNRDLSMILDAILKTRSRAVLIDIISKNGLQMLHNIIKRYRSDFNKIPIVRKLLKVIEYLAGREILTNNNIYGGPPRPGMESF
RVSMLSLTEHDDRQVHRTARSFRDKWFPRQRFEVKRHSSDKDEGRNEFRRGFNDNKTSTMHSLSLDNGPIQSKVASTSSGSAVVQSEIPSSSVDRSVNEV
GCSSALSDERTRKRKSRWDQPFRIKPDSKKTESDVKVPDDVDTCHTNLDEDLPPGFSSNPGSSVVSCDSSPAPKPNACGLKCPVEEDINVGGPRKKFDSC
LPVSFGIPWNIVQQFGTPADEAGSSWVVAPGIPFRPFPPLPPTPYQKFETPDCKTATFNNAVDSASRVVSCRNGGSLGKRYFKQQKWNRRTPWDWKHDGN
QTRNGDCGSSNGNGSLPSHENASSYTNTSTTTSQSNAQAPN
|
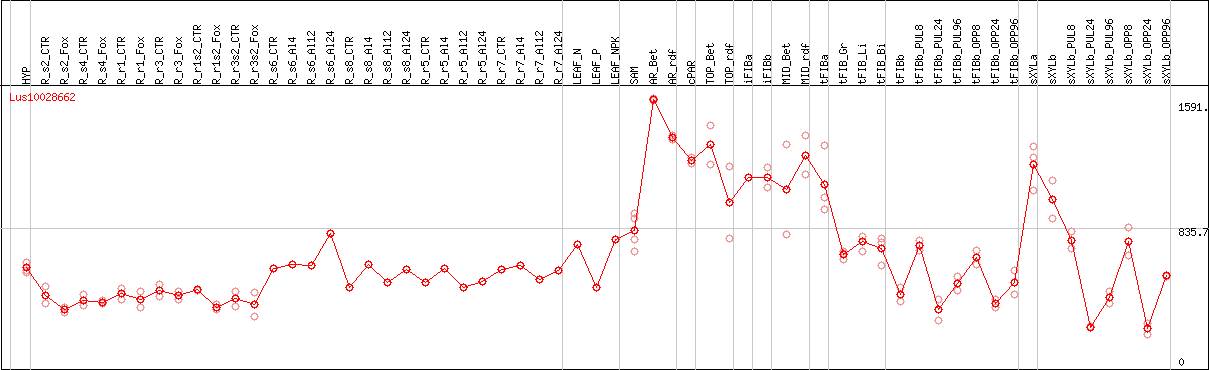
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028662 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
