Lus10028705 [FLAX]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Poplar homologues |
|
||||||||||
| PFAM info | |||||||||||
|
Representative CDS sequence |
>Lus10028705 pacid=23164496 polypeptide=Lus10028705 locus=Lus10028705.g ID=Lus10028705.BGIv1.0 annot-version=v1.0
ATGGAGAAACCGTCGCCACCTTCGTCGCCGTCGTCGAGCTCATTGGACGGTTCCAAAGAGCTGCAGTGCGTTGGTAGGTTGGAGATCGTAAGGCCGAAGC
CAGTAGGTTTCCTCTGCGGTTCCATTCCTGTGCCGACTGATAAGTCATTTCATGCCATTGATTCGGCTCTTGTTCCTTCGAGCCAGAAGGTGAGTGCGCC
GAGGTATAGAATGCTTCCGACGGAGACTGATCTCAACACTCTTCCAGTTGTTTCTAACTTGCCTGAGAAGGTTCTTCAGATCGGTGCTGTACAGTCTAAA
ACCACCGGTGAACTGCTATGGGAAGGGGATGCCGTCTCATCAAACATTAGAAGGAAATGTGAAGCCCTTGCTGTAACTGGTCTAGTTGAATATGGAGATG
AGATAGATGTGATAGCTCCAGCTGATATCCTGAAGCAAATCTTTAAGATGCCCTATTCCAAAGCTCGGTTGTCTATTGCAGTGCGTCGTATTGGACAAAC
GCTTGTATTGAATTCTGGGCCTGATGTTGAAGAGGGAGAGAAGCTAGTCAGAAAATATGGAACTCAATCAAAATGTGCTGATCAATATCTATTTCTGAAC
TTTGCTATGCATTCGGTGAGAATGGAGGCCTGTGATTGTCCGCCATCTCATCATATGCCCGAAGGCCAATCAAACTCTTCTTTGCCCGAGGGAAGTACTT
TACAGTCTGATAGGAAGGGTGATGATGCTGTATCGAATAAAGAAGTGAACCATAGTTCACACTACTCAGAGGTCAAACAAGGTGGTTTCTTATTAGAAAG
AAAGAAGAACAAGAGAAACAAAGGTCGCCATCCTATTAAGAAAACATCACATGTCAGTGAGAAGTCCAAAAGCACAGTGCAAGAATCCGAGAAACATAAA
AGAGTGAGACATGATGGGTTTCTAAGGGCTTTGTTCTGGCAGTTCCATAACTTCCGCATGCTTCTGGGCAGTGACTTACTTCTGTTCAGTAATGAAAAGT
ACCTTGCCGTGAGCTTGCATTTATGGGACACGACAGGAATCTCCGAAGATGGTACACCAGCTTTTCATCCTCATGTTGTTCAGCAGAATGGTCTTTCTGT
GCTGAGATTCCTTCAAAAAAATTGCAAGCAAGATCCCGGTGCTTATTGGCTTTACAAAAGCGCCGGCGAAGACATGATTCAACTTTTTGATCTTTCTATC
ATTCCTGACAATCACTCTTCCAGTGACTATAATGATGATTCAAGCTCCCTGCCATCACTGCATAGAGGGTGTGATTCCTCGTTCTCATTAGGAACTCTCC
TTTATCGAGTTGCACACAGGCTCTCCCTTTCAATGGCTCCTAATAGAAGAGCAAAGTGTGCTAGGTTCTTCAGAAAATGCTTCGAATTCCTCGAAGAGCC
AGATCATCTGGTTGTACGTGCATCAGCTCATGAAGAGTTTGCACGGCTTCTTTTGACTAATGTTGAAGAGCTGGAATTGAGCTCAGAATCTCTTGCTATT
GACTATGAAGCTACAGTCCCAGTATTGCCTCAGGATCCATTTGATTCCCCCCCCCCCGAATATGTTGTGGAGGAAGGCGACATTGAAACAGCTGGAAAAA
GTATAGGGACTCTAGGCAAAACTGAAATATTCGAAGCTACAGAGAGGACATTGGAGGTGGATGAACCTGTTCCCCGCAGTTTGATGCCTTTGGGTTACAA
AGAATTGAGAGGCCCAGTAGAAAACCTACCAACTGTCGATTCTGATGAAAGCCTCGTAGTCTGTGAGATACCTAAGCCATCTACCCAGGCGATTCAAACT
GTTGGTGATCCCATATCCACTAAGCTAGCTGCTGTTCATCATGTTTCACAAGCCATTAAGTCGTTCAGATGGGTGCGCCAGCTACAACCTTCTGATGCAG
AATTTATTGATGAAGACAGCATTGCTTATAATCAGCCTCCTGTAGAGTTCTCCGTCTGCGCTTGTGGGGATCCTGACTGCATTGAAGTTTGTGACATCCG
TGAATGGCTTCCAACGTCAAAAATAGATCATAAACTGTGGAAACTCGTTCTTCTACTTGGAGAATCCTATCTGGCCCTTGGGCAAGCATATAAAGAAGAT
AAACAGTTGCATCAGGCTCTCAAGATCGTAGAGTTAGCATCTTCTGTTTATGGATCAATGCCACAGCATCTGGACGATGCTCGGTTTATGTCCTCAATGG
TTAAGTATTCGTCCTCAAGGCAGTCTACTGACGAAGATGGCAAGCTAATATCTTATATTGGTAATGTAAAGGGAGTGAAATCTGGTACTAATGGTGACAA
GCTTGCCTTTGAGCACATGTCTTCGACATATCTATTCTGGGCAAAGGCATGGACATTGGTTGGAGATGTCTATGTCGACTTTCATTTCATGAAAGGTTCA
GAACTCTTGTTGCAAACGGAGAAGAAAAAATCTCTTGCAGAATTGCGAATGCCAGGTGAGGTAGTGAAGGAGGTCCAAAGGCTGAAGAAAAGGCTGGGTG
AGTCTGTTCAGAATTGCAGTTCTTGTTCTTTGGTGAATTGTAGCTGCCGGAGCGATAGGGCCAGTAGTGGTAGCAGTGCAAGTAGCAGCAGTGGAAATTC
TAAGCCTTCTGTATCTTATGGCAGAAAGCACGGGAAAAGATCCGGTTCTAAGAAAGTTAAATCGCCAATACTTCTAGACAGCAAGGATGATCCTGCTAAT
GGAGATATGGTAACTAAGAATAAGCCAGATGGAAGGCTTTCTCAACCCGGCCAATATGGTGAAATCCCGGCAGGAGATTCGGGTATTTCCATGGAGCAGC
CTAGGAAGAATCCTGATACAGTTGGCTTGGAGCGCCGTAAATCTGCTACAGATTCAAAAAACAACCCTAGAAGCCAGGATGGTGGGATATTTAAGTACCT
TCGGGGAACCATTGTTGGAGGCACGGAGGATAACCTTTCTACTGCCCGAACTTGCTACCAAGAAGCTAGAAAAGCACTAACTGGTCTTTCATCCGCTTCG
CTGGAACTGCAGTCTTTGTTAAAGAAGATAGGGTGGGTCTGCAACGAAATCGGCCGAAAGAGTCTGGAAAGGAATCAATTGACAGAGGCAGAATGTGCTT
TTGCCGATGCCATCAGTGCATTCAAGGAAGTGTCTGACCACGCAAACATCATATTGATTAATTGTAACCTTGGCCATGGCAGACGGGCATTGGCCGAGGA
AATGGTGACCAAAGCAGAAAGTCTCAGACATCATGCGATCTTCCAGAAAGCACACAAGCAAGCTCTAGAAACTGCAAAGCTCGAATACCGTGAATCACTT
GGGTACTATGGGGCAGCGAAGGCCGAACTGAATGCGATATCAGATTATGCAAACAATGTGTCGGTTGGTCTTAGAAATGAAGTGTGCACTCAGCTGGCTC
ATACATATCTGAGACTCGGCATGCTTTTGGCAAGAGAAGATATCACCGCTGAAGTAAACGAAAATGAGTTGTCCCGTGCCAGTAGAACTCGAAAAGATGG
GAGGATGCACGAGATTTCTGCCAATGATGCTATCAGGGAGGCGTTGTCTCTGTACGAGTCCCTAGGTGAAATTCGCAAACAAGAGGCTGCATATGCCTAC
TTTCAGCTTGCTTGCCATCAAAGGGATTGCTGCTTGAAATTTCTTGGGACGGACCGGAATACCAAATGTGAAAACAGCAGTCTACAGCGGGTGAAGCAAT
ATGCTTCTCTGGCCGAGAGGAACTGGCAAAAATCGGTCGATTTTTACATTTCAGAGACACACCCTTCCATGTATCTCACCATCGTCATAGAGTGA
|
||||||||||
|
AA sequence
|
>Lus10028705 pacid=23164496 polypeptide=Lus10028705 locus=Lus10028705.g ID=Lus10028705.BGIv1.0 annot-version=v1.0
MEKPSPPSSPSSSSLDGSKELQCVGRLEIVRPKPVGFLCGSIPVPTDKSFHAIDSALVPSSQKVSAPRYRMLPTETDLNTLPVVSNLPEKVLQIGAVQSK
TTGELLWEGDAVSSNIRRKCEALAVTGLVEYGDEIDVIAPADILKQIFKMPYSKARLSIAVRRIGQTLVLNSGPDVEEGEKLVRKYGTQSKCADQYLFLN
FAMHSVRMEACDCPPSHHMPEGQSNSSLPEGSTLQSDRKGDDAVSNKEVNHSSHYSEVKQGGFLLERKKNKRNKGRHPIKKTSHVSEKSKSTVQESEKHK
RVRHDGFLRALFWQFHNFRMLLGSDLLLFSNEKYLAVSLHLWDTTGISEDGTPAFHPHVVQQNGLSVLRFLQKNCKQDPGAYWLYKSAGEDMIQLFDLSI
IPDNHSSSDYNDDSSSLPSLHRGCDSSFSLGTLLYRVAHRLSLSMAPNRRAKCARFFRKCFEFLEEPDHLVVRASAHEEFARLLLTNVEELELSSESLAI
DYEATVPVLPQDPFDSPPPEYVVEEGDIETAGKSIGTLGKTEIFEATERTLEVDEPVPRSLMPLGYKELRGPVENLPTVDSDESLVVCEIPKPSTQAIQT
VGDPISTKLAAVHHVSQAIKSFRWVRQLQPSDAEFIDEDSIAYNQPPVEFSVCACGDPDCIEVCDIREWLPTSKIDHKLWKLVLLLGESYLALGQAYKED
KQLHQALKIVELASSVYGSMPQHLDDARFMSSMVKYSSSRQSTDEDGKLISYIGNVKGVKSGTNGDKLAFEHMSSTYLFWAKAWTLVGDVYVDFHFMKGS
ELLLQTEKKKSLAELRMPGEVVKEVQRLKKRLGESVQNCSSCSLVNCSCRSDRASSGSSASSSSGNSKPSVSYGRKHGKRSGSKKVKSPILLDSKDDPAN
GDMVTKNKPDGRLSQPGQYGEIPAGDSGISMEQPRKNPDTVGLERRKSATDSKNNPRSQDGGIFKYLRGTIVGGTEDNLSTARTCYQEARKALTGLSSAS
LELQSLLKKIGWVCNEIGRKSLERNQLTEAECAFADAISAFKEVSDHANIILINCNLGHGRRALAEEMVTKAESLRHHAIFQKAHKQALETAKLEYRESL
GYYGAAKAELNAISDYANNVSVGLRNEVCTQLAHTYLRLGMLLAREDITAEVNENELSRASRTRKDGRMHEISANDAIREALSLYESLGEIRKQEAAYAY
FQLACHQRDCCLKFLGTDRNTKCENSSLQRVKQYASLAERNWQKSVDFYISETHPSMYLTIVIE
|
DESeq2's median of ratios [FLAX]
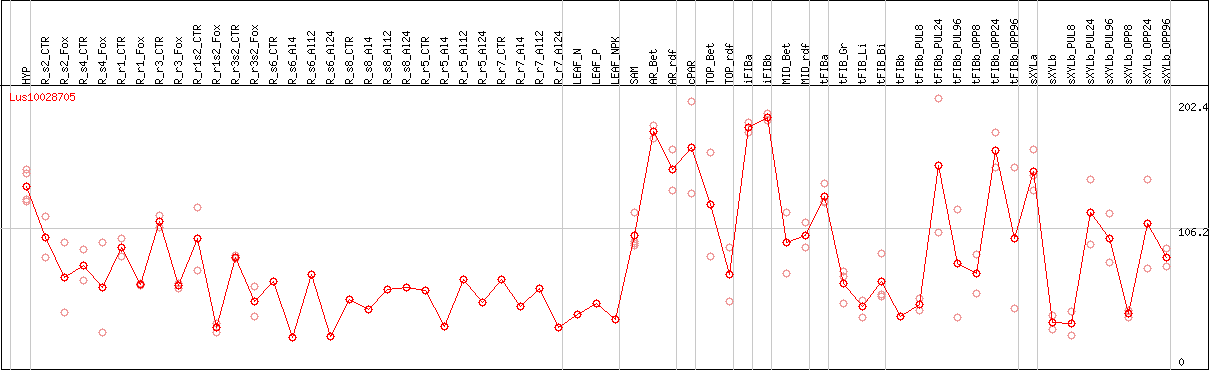
Coexpressed genes
Lus10028705 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
