Lus10028753 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10028753 pacid=23177269 polypeptide=Lus10028753 locus=Lus10028753.g ID=Lus10028753.BGIv1.0 annot-version=v1.0
ATGACTGAGCACAAGGAAGAGTCACTGATGGATAAGATATCGGAGAAGATCCACGGAGGCGATGATTCTTCTTCTTCGTCTTCCGATTCCGATCACGACG
ATAAGCCCAAGAAGTCGTCCTCTCCGGCTTCTCCTTCATCGATCAAGTCCAAAATCTATCGTCTCTTTGGACGTGAGCAGCCTGTTCACAAGGTTCTCGG
TGGTGGCAAACCCGCCGATGTTTTCCTGTGGAGGAACAAGAAGATCTCGGCAGGAGTGCTTGGTGGCGCAACTGCTGCATGGGTTCTCTTTGAATTGCTC
GAGTACCACCTTCTGACACTGGTTTGCCATGCTTTGATCCTCGCGTTGGCAGTTTTGTTTCTGTGGTCCAATGCCTCCACGTTTATCAACAAATCAACCC
CGCATATTCCGCAGGTCCACATCCCTGAGGATGCAGTTCTGCAATTTGCTTCTGGACTTAGGTATGAGATCAACCGCGGGTTGGCTGCTCTTCGTGATAT
AGCCTCAGGAAAGGACCTCAAAAAGTTCCTTGCTGTGATTGCTGGTTTATGGGTCCTATCAATCTTGGGGAGCTGTTGCAACTTCCTGACTCTGTTCTAT
ATCGTATTCGTGCTTCTCCACACAGTGCCAGTCTTGTATGAGAAGTATGAAGATAAAGTAGATGCATTTGGGGAGAAGGCATGGATCGAGATCAAGAAGC
AGTATGCAGTGTTCGACGAGAAGGTGCTAATTGGAGGGTATGTCGAGTGTCGGATGCAGATGAGGTTGGAAGTCGGGTATTTGGGGACACGGAAGTCCAA
AGGAACAGTTAGTTCATGTGTTCTCACTGTTTCTCAGCTTAGAATGTCGAGACAGCTAAGTTCAAAGTTCCCACTTGGAGAGCTAGCTGTTCACTCTTTT
TTGTGTGGTGTGAGACTTCCATTTCATCGAAACTCTCAGTGTCCCTCGACATTCACACTCCACCATCTCCAAACATCTTGGAAAACCATAGGCAAGCACA
AGAAATCAAGAGTAGGAGGAATGATTGTACAGTCTGTACCTAAGCGAGACTGTACATCTGCTTCCTCGTCTGTAGATACCGAAGCTGAGAATCAGGCGGA
GTGTGCAATCTCCCTTCCCCCAACTGTCCCTCGTTTCTATCGACCAATTTCTTCTTCACTCCTCAGTTTAATTCTTTCTATTCTCGCCATTATTAAACCC
CCTGAAGCTCATTTTCCCTCTTTCGCTCACTCTCAATTTCAGCTGTCTCTTTATTCTGACTTTGGCATTGGAACTCGCATTCCGAGAGCTCAACGATTTG
TCTACATACCCCGGTTTCCCTGCTTACTATCGGAGATGGAGGCCACATTGCCGGGTTGTAAATCAGTTACCTCCGTTCCTGGTTTAATTGTGAGAACAAC
ACCAGGAATAAGAAGTTCACAATGTAGCCTTTTGGTGGGGACTAAGGTCAACTTTCCTAGACTAGGAGTTCATGCTGCTAAGGTTGCTCATAAAAGTAAT
AAGAGAGGTGGAGCGCCACGTGTAACATGCCATGCAGAAAAGATTCTGGTCGCGAACAGAGGGGGAAGTGCTGTTCGTGTGATACGGACTGCCCACGAAT
TAGGAATACCATGTGTTGCTGTTTACTCAACCATCGACAAGGATGCCCTTCACGTCAAATTGGCTGATGAATCAGTTTGTATCGGTGAAGCACCAAGCAA
TCAGTCGTACTTAGTGATAGCCAATGTCTTATCTGCTGCTGTTAGTCGCCATTGTACAATGCTGCATCCTGGATATGGTTTTCTTGCTGAGAATGCCACA
TTTGTAGAAATGTGCCGGGCACACGGGATAAATTTTATTGGACCCAATCCGGATAGCATCAAAACTATGGGTGATAAAGCCACTGCCAGAGAAACAATGA
AGAATGCAGGAGTTCCAACTGTTCCAGGAAGTGATGGATTGTTGCAGAGCACAGAGGAAGCAATTAGGCTGGCTGAAGAAATAGGTTATCCTGTGATGAT
CAAGGCAACGGCTGGTGGTGGTGGGCGTGGAATGCGTCTTGCTAAAGAACCAGAGGAATTTGTGAAGTTGCTTCAGCAAGCCAAGAGCGAGGCTGCTGCT
GCATTTGGAAATGATGGAGTTTATTTAGAAAAATACATTCAAAATCCAAGGCACATCGAGTTCCAGGTCCTTGCAGATAAGTATGGTAATGTCATCCATT
TTGGCGAGAGAGACTGTAGTATCCAGAGACGTAACCAGAAGCTTCTGGAAGAGGCTCCCTCCCCAGCACTGACCCCAGAGTTGAGGAAAGCCATGGGTGA
TGCAGCAGTTGCAGCTGCCGCATCCATAGGGTACATTGGTGTGGGAACAGTGGAGTTCCTTCTGGATGAGAGGGGTTCCTTCTACTTCATGGAAATGAAT
ACTCGTATCCAGGTTGAGCATCCTGTAACTGAAATGATTTCATCTGTGGACTTGATTGAGGAACAAATTAGAGTAGCTATGGGAGAAAAACTTCGTTATA
AACAGGAAGAAATTGTGCTCAGAGGTCATTCAATCGAGTGCCGGATCAATGCAGAGGATGCATTTAAGGGATTTAGACCGGGACCAGGAAGAATAACAGC
ATACTTGCCATCTGGAGGACCCTTTGTCAGAATGGACAGTCATATTTATCCAGATTATGTTGTCCCTCCAAGCTATGATTCTCTGCTTGGAAAGCTTATT
GTTTGGGCCCCGACTAGAGAAAAGGCGATTGAGCGTATGAAGAGGGCCCTTAATGACACTATTATTACGGGAGTTCCTACAACAATTGATTATCATAAGT
TGATCCTTGAGATTGAGGACTTCAAAACTGGAAACGTCGATACAGCATTCATTCCAAAGCACGAAGATGAATTACAAGCGCCAACCCAGATAGTGCCAGC
AAACGGGATGGCCGCTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10028753 pacid=23177269 polypeptide=Lus10028753 locus=Lus10028753.g ID=Lus10028753.BGIv1.0 annot-version=v1.0
MTEHKEESLMDKISEKIHGGDDSSSSSSDSDHDDKPKKSSSPASPSSIKSKIYRLFGREQPVHKVLGGGKPADVFLWRNKKISAGVLGGATAAWVLFELL
EYHLLTLVCHALILALAVLFLWSNASTFINKSTPHIPQVHIPEDAVLQFASGLRYEINRGLAALRDIASGKDLKKFLAVIAGLWVLSILGSCCNFLTLFY
IVFVLLHTVPVLYEKYEDKVDAFGEKAWIEIKKQYAVFDEKVLIGGYVECRMQMRLEVGYLGTRKSKGTVSSCVLTVSQLRMSRQLSSKFPLGELAVHSF
LCGVRLPFHRNSQCPSTFTLHHLQTSWKTIGKHKKSRVGGMIVQSVPKRDCTSASSSVDTEAENQAECAISLPPTVPRFYRPISSSLLSLILSILAIIKP
PEAHFPSFAHSQFQLSLYSDFGIGTRIPRAQRFVYIPRFPCLLSEMEATLPGCKSVTSVPGLIVRTTPGIRSSQCSLLVGTKVNFPRLGVHAAKVAHKSN
KRGGAPRVTCHAEKILVANRGGSAVRVIRTAHELGIPCVAVYSTIDKDALHVKLADESVCIGEAPSNQSYLVIANVLSAAVSRHCTMLHPGYGFLAENAT
FVEMCRAHGINFIGPNPDSIKTMGDKATARETMKNAGVPTVPGSDGLLQSTEEAIRLAEEIGYPVMIKATAGGGGRGMRLAKEPEEFVKLLQQAKSEAAA
AFGNDGVYLEKYIQNPRHIEFQVLADKYGNVIHFGERDCSIQRRNQKLLEEAPSPALTPELRKAMGDAAVAAAASIGYIGVGTVEFLLDERGSFYFMEMN
TRIQVEHPVTEMISSVDLIEEQIRVAMGEKLRYKQEEIVLRGHSIECRINAEDAFKGFRPGPGRITAYLPSGGPFVRMDSHIYPDYVVPPSYDSLLGKLI
VWAPTREKAIERMKRALNDTIITGVPTTIDYHKLILEIEDFKTGNVDTAFIPKHEDELQAPTQIVPANGMAA
|
DESeq2's median of ratios [FLAX]
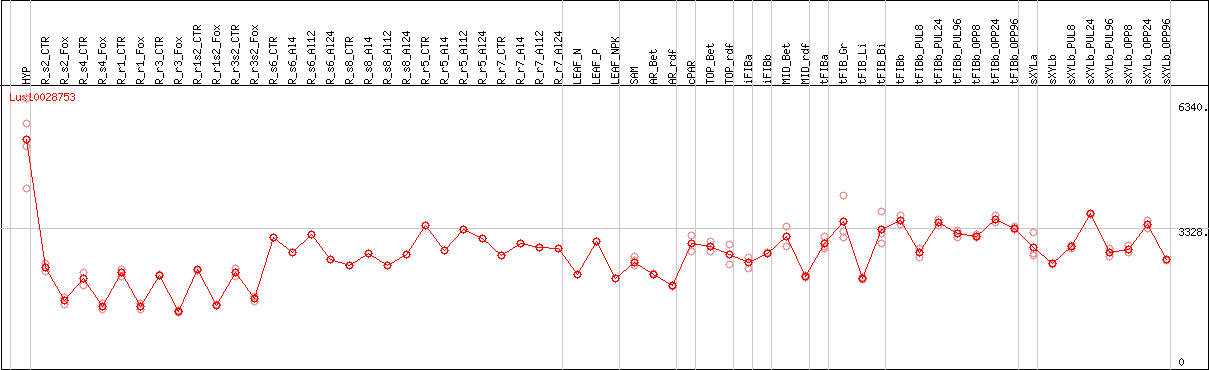
Coexpressed genes
Lus10028753 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
