External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G01110 117 / 2e-32
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G61990 82 / 5e-20
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G64100 79 / 7e-19
pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
AT1G19290 78 / 2e-18
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63150 77 / 3e-18
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63330 76 / 7e-18
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63630 75 / 7e-18
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G05670 76 / 9e-18
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
AT1G12300 76 / 9e-18
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G39710 76 / 1e-17
EMB2745
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007468
170 / 3e-51
AT5G01110 808 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019524
86 / 4e-21
AT3G54980 716 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013077
76 / 1e-17
AT5G40400 587 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10027916
74 / 4e-17
AT5G39710 1038 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10028943
73 / 5e-17
AT5G01110 299 / 2e-97
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10027914
73 / 1e-16
AT5G39710 1025 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10032789
73 / 1e-16
AT1G09820 250 / 4e-78
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10012068
73 / 2e-16
AT5G39710 1028 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10013283
72 / 2e-16
AT1G05670 785 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G276500
139 / 6e-40
AT5G01110 863 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G250700
84 / 9e-21
AT3G54980 781 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.014G117600
79 / 7e-19
AT1G12700 330 / 1e-105
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.009G105600
79 / 8e-19
AT5G39710 1022 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G035700
77 / 3e-18
AT1G05670 922 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Potri.013G032600
76 / 9e-18
AT1G12700 493 / 2e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.016G025600
76 / 2e-17
AT3G22470 426 / 3e-141
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G105700
74 / 5e-17
AT4G19440 758 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.018G038300
74 / 6e-17
AT1G12700 490 / 8e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.015G105400
73 / 1e-16
AT5G61990 773 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10028942 pacid=23177310 polypeptide=Lus10028942 locus=Lus10028942.g ID=Lus10028942.BGIv1.0 annot-version=v1.0
ATGGAAAGCCAAGGGTTGTTGCCGAATATTGTCACTTATAACGCAATCCTGAATGGGTTTTGTAGTCTAGGCAGAATGCAAGATGCTGAATCGGTGCTTC
AGAAAATGATTGAGAAGGGCGTATATCCTGATCGATCCACATACACAGCTCTTATAAATGGACATGTCAGCCAGGAGAACTTAAAGGAGGCGTTTCGATT
CCATGATGAAATGTTACTCAGAGGTTTTGTACCAGATGACGATACTTGA
AA sequence
>Lus10028942 pacid=23177310 polypeptide=Lus10028942 locus=Lus10028942.g ID=Lus10028942.BGIv1.0 annot-version=v1.0
MESQGLLPNIVTYNAILNGFCSLGRMQDAESVLQKMIEKGVYPDRSTYTALINGHVSQENLKEAFRFHDEMLLRGFVPDDDT
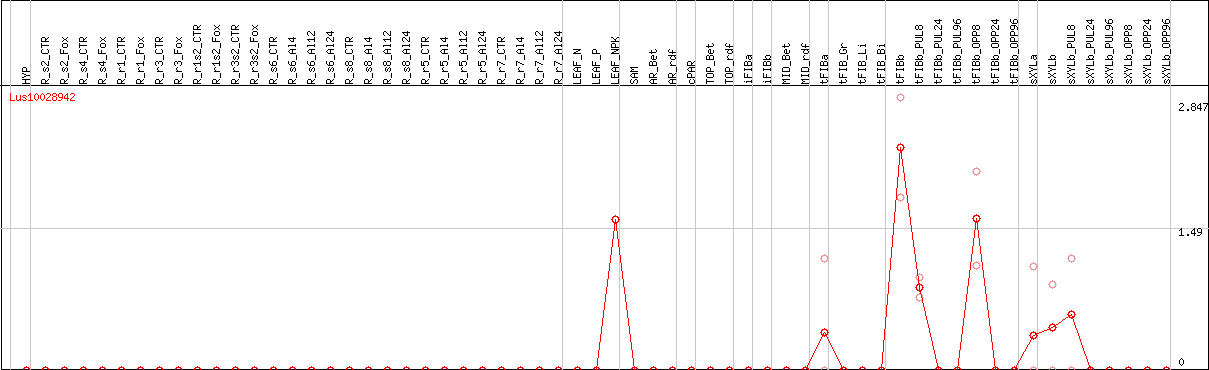
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028942 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.