External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G05550 96 / 4e-25
Trihelix
sequence-specific DNA binding transcription factors (.1.2)
AT3G11100 90 / 7e-23
Trihelix
sequence-specific DNA binding transcription factors (.1)
AT3G58630 67 / 6e-14
Trihelix
sequence-specific DNA binding transcription factors (.1)
AT3G14180 67 / 9e-14
Trihelix
ASIL2
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
AT1G54060 57 / 2e-10
Trihelix
ASIL1
6B-interacting protein 1-like 1 (.1)
AT3G24490 47 / 8e-07
Trihelix
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025406
79 / 2e-18
AT3G14180 202 / 3e-63
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Lus10015281
75 / 6e-18
AT3G58630 79 / 2e-18
sequence-specific DNA binding transcription factors (.1)
Lus10025404
75 / 4e-17
AT3G14180 197 / 1e-61
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Lus10029220
68 / 8e-16
AT3G14180 74 / 4e-17
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Lus10007273
72 / 9e-16
AT3G14180 200 / 4e-62
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Lus10029889
62 / 3e-12
AT3G14180 147 / 1e-40
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Lus10035893
62 / 4e-12
AT3G24490 214 / 1e-66
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
Lus10025772
60 / 3e-11
AT3G24490 223 / 3e-70
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
Lus10008141
58 / 1e-10
AT3G14180 271 / 3e-87
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G071300
124 / 8e-36
AT5G05550 209 / 3e-67
sequence-specific DNA binding transcription factors (.1.2)
Potri.010G186200
119 / 3e-34
AT5G05550 149 / 3e-44
sequence-specific DNA binding transcription factors (.1.2)
Potri.001G113600
75 / 7e-17
AT3G58630 138 / 1e-38
sequence-specific DNA binding transcription factors (.1)
Potri.003G118600
75 / 7e-17
AT3G58630 139 / 5e-39
sequence-specific DNA binding transcription factors (.1)
Potri.003G200000
68 / 2e-14
AT3G14180 172 / 2e-49
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Potri.T126206
68 / 2e-14
AT3G14180 173 / 9e-50
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Potri.001G026000
66 / 3e-13
AT3G14180 186 / 1e-54
Arabidopsis 6B-interacting protein 1-like 2, sequence-specific DNA binding transcription factors (.1)
Potri.001G179800
57 / 3e-10
AT3G24490 206 / 2e-63
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
Potri.003G056000
56 / 6e-10
AT3G24490 205 / 8e-63
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
Potri.018G075500
52 / 1e-08
AT3G24490 252 / 1e-81
Alcohol dehydrogenase transcription factor Myb/SANT-like family protein (.1)
PFAM info
Representative CDS sequence
>Lus10028985 pacid=23177200 polypeptide=Lus10028985 locus=Lus10028985.g ID=Lus10028985.BGIv1.0 annot-version=v1.0
ATGGAGGACGTGAGCGGCTCCGACGGAGCTGCTTGTAAGGAATTGGCGAGGGCGATACTGAAATTCGCGGAGATTTACGAGAAAGTCGAGAGCTTGAAGC
AGAAGCAGATGGTGGAGCTGGAGAAGCAGAGGATGGAGTTCACTAAAGAGCTCGAATTTGAGAGGATGAACATGTTCATGGCTGCCCAATTGGACATGAA
GAAGAAGAAGTCGTTTAAACGGCAGAAAGTCGCTTCTGGATCAGTCAACTTAGATGCTGCAAATGGTGATGCTAACTTGATTTACTTCCAGGTGGAGAGT
AATGCCTGCAAATCTGGATGCTATTTAGGAACCAAGTCGACTCTTTTTTCTGAATCGGGGAGAGCTTTCACGTTTCCAAACTCGTCTGTCGCATGA
AA sequence
>Lus10028985 pacid=23177200 polypeptide=Lus10028985 locus=Lus10028985.g ID=Lus10028985.BGIv1.0 annot-version=v1.0
MEDVSGSDGAACKELARAILKFAEIYEKVESLKQKQMVELEKQRMEFTKELEFERMNMFMAAQLDMKKKKSFKRQKVASGSVNLDAANGDANLIYFQVES
NACKSGCYLGTKSTLFSESGRAFTFPNSSVA
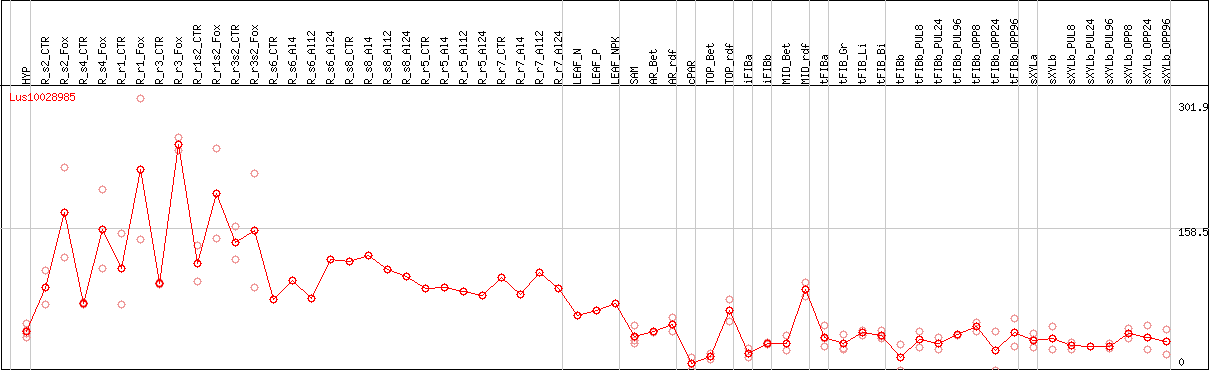
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028985 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.