External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G30340 168 / 6e-52
AS2
LBD13
LOB domain-containing protein 13 (.1)
AT2G40470 149 / 8e-45
AS2
ASL11, LBD15
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
AT5G63090 122 / 8e-35
AS2
LOBB, LOB
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT2G30130 121 / 1e-34
AS2
PCK1, LBD12, ASL5
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
AT3G27650 114 / 4e-32
AS2
LBD25
LOB domain-containing protein 25 (.1)
AT5G66870 117 / 8e-32
AS2
LBD36, ASL1
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
AT1G16530 112 / 2e-31
AS2
ASL9, LBD3
LOB DOMAIN-CONTAINING PROTEIN 3, ASYMMETRIC LEAVES 2-like 9 (.1)
AT1G07900 108 / 2e-29
AS2
LBD1
LOB domain-containing protein 1 (.1)
AT1G31320 107 / 2e-29
AS2
LBD4
LOB domain-containing protein 4 (.1)
AT2G23660 109 / 8e-29
AS2
LBD10
LOB domain-containing protein 10 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034208
248 / 6e-84
AT2G30340 201 / 9e-65
LOB domain-containing protein 13 (.1)
Lus10009930
161 / 1e-49
AT2G30340 230 / 5e-76
LOB domain-containing protein 13 (.1)
Lus10024183
143 / 1e-42
AT2G30340 230 / 4e-76
LOB domain-containing protein 13 (.1)
Lus10018275
121 / 1e-34
AT1G31320 243 / 2e-83
LOB domain-containing protein 4 (.1)
Lus10027676
120 / 3e-34
AT5G63090 233 / 8e-79
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Lus10040638
120 / 3e-34
AT1G31320 244 / 1e-83
LOB domain-containing protein 4 (.1)
Lus10031285
120 / 4e-34
AT2G30130 226 / 3e-76
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Lus10031191
119 / 4e-34
AT1G31320 199 / 2e-66
LOB domain-containing protein 4 (.1)
Lus10039940
119 / 1e-33
AT5G63090 236 / 5e-80
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G081200
161 / 1e-49
AT2G40470 249 / 2e-84
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
Potri.013G156200
155 / 2e-47
AT2G40470 207 / 2e-67
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
Potri.019G127300
144 / 3e-43
AT2G40470 225 / 1e-74
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
Potri.005G097800
125 / 2e-36
AT1G31320 193 / 6e-64
LOB domain-containing protein 4 (.1)
Potri.007G066700
124 / 4e-36
AT1G31320 216 / 3e-73
LOB domain-containing protein 4 (.1)
Potri.001G081400
120 / 3e-34
AT1G31320 228 / 3e-77
LOB domain-containing protein 4 (.1)
Potri.010G184400
119 / 8e-34
AT2G30130 227 / 7e-77
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Potri.003G149000
119 / 1e-33
AT1G31320 241 / 2e-82
LOB domain-containing protein 4 (.1)
Potri.015G082200
119 / 1e-33
AT5G63090 223 / 4e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Potri.001G281600
117 / 2e-33
AT2G30130 248 / 4e-85
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Lus10029061 pacid=23151837 polypeptide=Lus10029061 locus=Lus10029061.g ID=Lus10029061.BGIv1.0 annot-version=v1.0
ATGGAGAATTCAGATGATGTAGAAAGAGTGATGATGCTAAGTAGTAATAGTAGTCAAGCATCAGGAGGACTGAATACAATAACTCCATGTGCAGCCTGCA
AGCTGCTGAGGAGGAGATGTGCAGCTCAAGAGTGCCCTTTCTCGCCATACTTCTCCCCTCTCGAGCCCCACAAGTTTGCTTACGTCCACAAGGTCTTTGG
TGCCAGCAATGTCTCCAAGATGCTCACGGAAGTGGGAGAGAGCCAAAGGTGGGATGCGGCAAACAGCCTAGTATACGAGGCAAGTGTGAGGCTAAGAGAC
CCTGTTCATGGGTGCATGGGTGCAATCTCACTTCTCCAACACCAAATCCACTCTTTGGAATCCGAACTCACCACTCTCAGGGCCGAGATTCTTCATCATA
GATACAATGCTGCAGCAGCAACACCACCATCAACAATGGACATTGTTACTACTACTACTACATCACCTATGATCATCAATTCCCCTCCTCCTGTTGATTT
TAGCCATGATGTTCGAGGCCAAGGGGCAGTGCTCCTTCCTCCCGTCACACTGTCGACAGATTCTTCTTCCTCGTCCTCCCCTTCGAATCGTTATGGTGGC
ATTTCGAATAATGATCACAACATCCCCTTCTATGGTTAA
AA sequence
>Lus10029061 pacid=23151837 polypeptide=Lus10029061 locus=Lus10029061.g ID=Lus10029061.BGIv1.0 annot-version=v1.0
MENSDDVERVMMLSSNSSQASGGLNTITPCAACKLLRRRCAAQECPFSPYFSPLEPHKFAYVHKVFGASNVSKMLTEVGESQRWDAANSLVYEASVRLRD
PVHGCMGAISLLQHQIHSLESELTTLRAEILHHRYNAAAATPPSTMDIVTTTTTSPMIINSPPPVDFSHDVRGQGAVLLPPVTLSTDSSSSSSPSNRYGG
ISNNDHNIPFYG
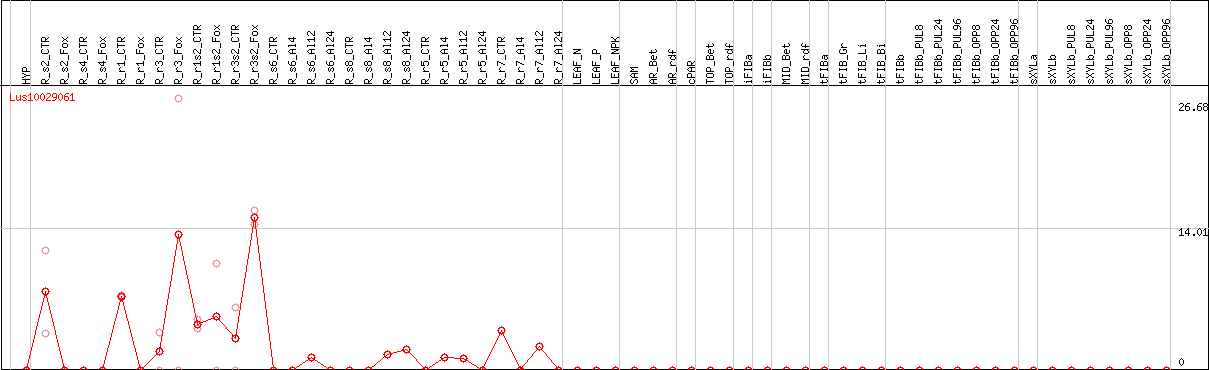
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029061 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.