External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G29330 316 / 2e-109
ATERD2, AERD2, ERD2
ARABIDOPSIS THALIANA ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ARABIDOPSIS ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ER lumen protein retaining receptor family protein (.1)
AT3G25040 258 / 1e-86
ERD2B
endoplasmic reticulum retention defective 2B (.1)
AT1G19970 87 / 5e-20
ER lumen protein retaining receptor family protein (.1)
AT3G25160 87 / 7e-20
ER lumen protein retaining receptor family protein (.1)
AT1G75760 87 / 9e-20
ER lumen protein retaining receptor family protein (.1)
AT4G38790 84 / 8e-19
ER lumen protein retaining receptor family protein (.1)
AT2G21190 84 / 1e-18
ER lumen protein retaining receptor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022785
260 / 2e-87
AT3G25040 382 / 9e-137
endoplasmic reticulum retention defective 2B (.1)
Lus10011846
249 / 3e-83
AT3G25040 364 / 7e-130
endoplasmic reticulum retention defective 2B (.1)
Lus10043149
242 / 2e-80
AT3G25040 332 / 2e-117
endoplasmic reticulum retention defective 2B (.1)
Lus10007314
174 / 4e-55
AT1G29330 179 / 2e-58
ARABIDOPSIS THALIANA ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ARABIDOPSIS ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ER lumen protein retaining receptor family protein (.1)
Lus10021097
92 / 8e-22
AT1G75760 475 / 8e-172
ER lumen protein retaining receptor family protein (.1)
Lus10007570
92 / 1e-21
AT4G38790 466 / 2e-168
ER lumen protein retaining receptor family protein (.1)
Lus10017213
91 / 4e-21
AT1G75760 481 / 5e-174
ER lumen protein retaining receptor family protein (.1)
Lus10026285
88 / 4e-20
AT4G38790 473 / 7e-171
ER lumen protein retaining receptor family protein (.1)
Lus10012176
87 / 1e-19
AT2G21190 397 / 2e-140
ER lumen protein retaining receptor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G071200
297 / 5e-102
AT1G29330 341 / 1e-120
ARABIDOPSIS THALIANA ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ARABIDOPSIS ENDOPLASMIC RETICULUM RETENTION DEFECTIVE 2, ER lumen protein retaining receptor family protein (.1)
Potri.017G056000
268 / 1e-90
AT3G25040 383 / 3e-137
endoplasmic reticulum retention defective 2B (.1)
Potri.001G315800
266 / 4e-90
AT3G25040 355 / 3e-126
endoplasmic reticulum retention defective 2B (.1)
Potri.004G167400
98 / 7e-24
AT4G38790 472 / 1e-170
ER lumen protein retaining receptor family protein (.1)
Potri.009G128900
97 / 1e-23
AT2G21190 476 / 2e-172
ER lumen protein retaining receptor family protein (.1)
Potri.002G022300
92 / 9e-22
AT1G75760 464 / 2e-167
ER lumen protein retaining receptor family protein (.1)
Potri.002G246700
84 / 1e-18
AT3G25160 400 / 5e-142
ER lumen protein retaining receptor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0141
MtN3-like
PF00810
ER_lumen_recept
ER lumen protein retaining receptor
Representative CDS sequence
>Lus10029263 pacid=23139939 polypeptide=Lus10029263 locus=Lus10029263.g ID=Lus10029263.BGIv1.0 annot-version=v1.0
ATGAATATCTTCAGATTCCTCGGTGACATGACTCACCTGATCAGCATCCTAATCCTTCTCCTCAAAATCTACGCCACCAAATCTTGCTCAGGTTGCAATC
GCGCGAAATTGTTAGTCGAGCTGCGTTGGATTAGGAGGAAAACCTATTTGATTCATTTAGTTTTGCTTGAGCTAACGGCCATTGTGATGCGTTTTTTTCT
AGGGGTTTCAAGGAAGACCCAGGAGCTGTACGCGCTGGTGTTCCTGGCGAGGTACTTGGATCTGTTCACCACCTTTGTGTCCGTATACAACTCTGTTATG
AAGGTTGTGTTCGTAGCCAGCAGTCTCGCCATTGTCTGGTGCATGCGGAGGCATCCGCTCGTTCGTCGTTCTTACGATAAGGACCTCGATACCTTCCGTC
ACTATTTCCTCGTTGGTGGGAGCTTTGTGCTGGCACTCTTGTGGCACGAGAAGTTCACGTTTCAGGAGATCTTCTGGGCCTTCTCAATATATTTGGAAGC
AGTGGCTATCCTTCCTCAGTTAGTGTTGCTCCAAAGAAGCGGGAATGTTGATAACTTAACAGGGCAATATGTCATCTTCCTTGGGGCTTACCGTGGACTC
TACATCTTGAATTGGATCTATCGTTACGTGACTGAGGCACATTTTGGTCGTTGGATTGCTGTTGTCTCTGGTGTCATCCAAACAGCCTTGTACGCCGACT
TCTTCTACTACTACTTCATGAGGTACTTGGAAGAACAAGACGAAGCTCCAGCTGCCAGCTTGAATCCGAACATACTTCAAGCGAAGGTTGCATATCTTAT
AAGCGTTTTTGAGGGGGTGAGATTGGATCCTTTGGCGTCTGGTAGAGTAGAAAAATTCTAG
AA sequence
>Lus10029263 pacid=23139939 polypeptide=Lus10029263 locus=Lus10029263.g ID=Lus10029263.BGIv1.0 annot-version=v1.0
MNIFRFLGDMTHLISILILLLKIYATKSCSGCNRAKLLVELRWIRRKTYLIHLVLLELTAIVMRFFLGVSRKTQELYALVFLARYLDLFTTFVSVYNSVM
KVVFVASSLAIVWCMRRHPLVRRSYDKDLDTFRHYFLVGGSFVLALLWHEKFTFQEIFWAFSIYLEAVAILPQLVLLQRSGNVDNLTGQYVIFLGAYRGL
YILNWIYRYVTEAHFGRWIAVVSGVIQTALYADFFYYYFMRYLEEQDEAPAASLNPNILQAKVAYLISVFEGVRLDPLASGRVEKF
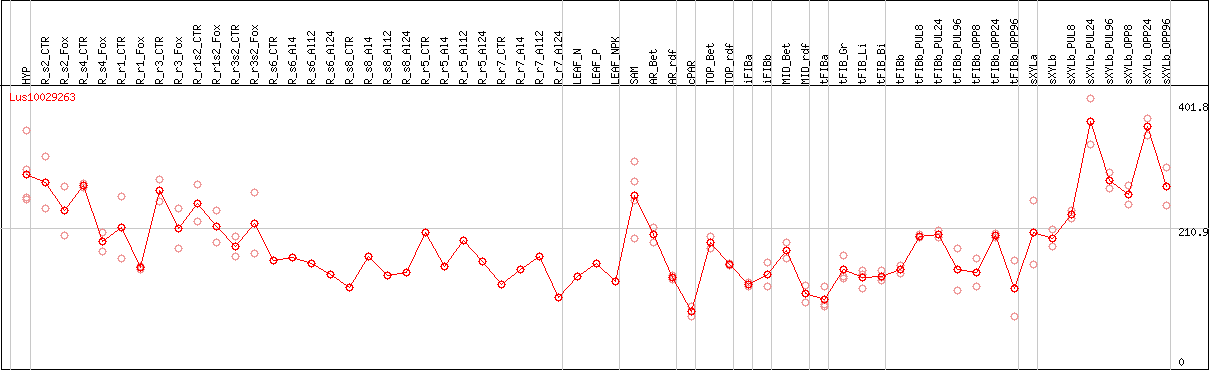
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029263 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.