Lus10029303 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029303 pacid=23139944 polypeptide=Lus10029303 locus=Lus10029303.g ID=Lus10029303.BGIv1.0 annot-version=v1.0
ATGTTCAGGCTGCACAAGATCAAGTCGGACCGGTTGTCTCTAGGGGAGCGGATCGATTTCAAGATCTCCCATTTCAAAGCACTCCAGGTACCAAGAGGTT
GGGACAAGCTGTATGTGACAGTGATATCGGTGGAGACAGGGAAGACTATAGCAAGGCTGAGCAAAGCAGTTGTGAAGAATGGTGGCAGCTGCCATTGGGA
AGAGAGCTTTACCGAATCGGTTTGGGTCTCGCCCGACGAATCGTCCAAGGTGCTTGAAGATTGCATCTTCAAGTTCGTCCTTGCAATGGGATCTGCGAGA
TCTGGTATTCTGGGAGAGGTGACTGTGAACATGTCGAGCTATATCAGCTCGAGTTCAGCAGTTTCGGTATCTTTTCCCTTGAAGAAATGCGATTATGCCA
CTGTTCTGCAGGCCACGATTCAGTGCTTGACACCGATGGCGAATCTCAGGGATGCAGAGGGCAAAGAAACAAACCTTACTGATAAAAATGGCAAAGAAGA
AGAATTACGAGACGATGAAGAAGGAACCAAGTCCGATCAGGAATCCGATACAGCTTCTCCGAAAGGCGGCCCTTCTTACGGAAGTAGTGACACTGGCTCT
CCACCGGTTCAGGCCGGTCCTGAAACCAGGGAAGCTAGTTCCTCCGCAACGAATTCTCCTGGCAGCGAAGATTTGGAAGAAGCTGCAACAGCAGGCAGGC
CTGTTCTGTTTCCTTCGAGACCAAGCCTGGTCCGAAGAGAAAGAAGAACCAGTTCCGGAAATGGAATTTCGAGTATGAACTATGTCGTGGACAGTCCTCG
ATCCGAAAGATCCGTACCTATGAGGAGAATCCCGCGTCCAGGAAGCCGCTCAGGGGAAATCCATCAGGAATTCGGCATTAGCAGGCTGAGAAATTCAGCT
TTACGAACCATGGATTCCTCCTCGACTTCGCAGACGTTCCTCGAAGCTGCGGAGGAAACGATTGGAGACCTCAGGACGCAGGCCAAGATGTGGGAGAGGA
CTGCCAGAAAGCTGATGCTTGAGATGGACTACCTGAAGAAAGAAAACGTGGAGCTTTCCAGGAATCAAGGGAATCAGGATATGGAGTTTACGGAGGCTTG
CGCCGAAAGGGATGGATTAAGGAAAGAAGTCGAGCAGCTGAAGCAGTTGTTAGCGAAGGCGAAGATGAAGCCGCCTCCTTTGGATAATCCGAGCGAAGGT
ACGTGGCAAATGGTTAAGGAGTTGGAAAGGGAGATCAGTTTCCAGAAGGAAACAAGTGGAAACCTGACGATGCAGCTTAAGAGGAGTCAAGAGTCGAATG
TTGAGCTCGTTACTGTTCTTCAGGATCTGGAAGAGACACTGGAGAAGCAGAAGATTGAAATCGAGAGTCTTTACGACATGGAGTCGAGATTCAGCAAGTT
GGAAGATGCTATGGAAGCGAGCGCTGCAAGTAACAAGAACCTGAAGGTTCAGGTTCTGAAACTTCAGGAATCTGAAAAGGATTTGCTCAACAAAGTTCGA
GAGCTGGAATCTGAAAAGGATTCCACCACTACAGACAGCGTCTCGATTCGAGAACGAGAAGTTGAGAGCATTAAAATGGAGATGCAAGGCAGAGTCAGAG
AGCTTACCGAGGAGCTAGATGGAGTAAGAGCAGAGGTGAAAAGACTTGAGGCAGGCTTGGTTTTGAAACAGGTGGAGAATAAGGATCTTCAGAGATGTAG
AACTGGAGCGGAGGCCAGGGTTTCGGTTCTGGAAAAGGAAAAGGAACATTTGGAGCTTGAGTTGGAAACAGTGACGAGAGAAAATGTAATTGCTTCAAAA
TGCTTGAATGACCTGCAAAAGGATTTAGCAACAATAAGCAGCAACCTTGAAACTCATGCTTCCGTCAATAAGAACCTGGAAAGAAAATCGTTGGAGCTAG
AACAGAACAATCAAGAAATAGAGCTTTGTTTATCAGAACTAAGACAGGAGAACGAAGATCTATCAACACACATTCTTGCGTTGGAAACTCGGTTAAGGTT
CGAAGCAAGTGAACATAAATCCCGGATCTGTGAGCTTGAGAAGGAGCTAAGGAAGGCTGAAAGTGATTTCAACGTTTCAGCTCAAGAGATTGAAAGGCTG
AAACTTGAGTTGGAAGTTAAAGTCAGTGAAATGTCAAAGCAGGTAGATGAAAGCAAGGCTGAGATTCAAAGTCTGGAAACCATTATGCAGGCTAATGAAG
AGGAGGTCAGAGATCTTCAGAGGTGGAAAAGTGAGTTAGCAGCCAAACTTTCTGTTATTCAGGGCGAGAAGGACCAACTGGAGGAGAAGATGGGAGCCAT
CATGACAGAAAACGATATCGCCACAAGACGCTCCAACGAGCTACAATCGGAGCTTACCACGACCAGAAGCAGCTTAGAAGAAAAGAGCTCTACTAATGAA
GGCCTTGAGAGGAAGCATCTGGAACTGGAAAAACAGAAGAATGACTTGGGGAAACAATTGTCAGGAATCGAGCAAGATAATCAGGAGTTGTTGATGAACA
TAACTGACCTGGAAACTCAGCTAAGGTCTGAAGCCAGTGACCACGAATCCCAACTGAATTCTGTAGAAGAGAGCTACAAAAAGCTGCTTATGCAGATTGA
TAGCGACTGCAACGATTCACTAGCAAAGGTTGAGAAATTGAGAATGGAGTTAGAAGCTGAAGTTACAGAACTGAAAGCAGAACTCAGTGACAGGAAGGCA
GAGATTGAGATACTTGATGCCTGTTTGCATGCCAAAGAAGAGGAGATCGGACAGCTTCAAAGGCACACAAGAGAATCAGAGGTCAAAATTTTCGATTTGA
AGGAGGAAAAATGCCAGCTAGAGAACAACATGGAAACAGTCATGGAAGAAAATGACGTTGCAGTGAGACGCTTGAATGACTCTCAACAGGAAATAATTAT
GCTCAGAAGCAGTGTTGAATCCCATGTTGCTTCAAACAATGCTCTCGAGACCAAATGCTCGGAGCTAGAAGGGAAGAAGCTAGAACTGGAGCTCCATTCT
TCGGAGCTAAAGCAGGGAAATGAACAGCTCGCCACACTTTTGTCAGAATTGGAAGCTGAACTAAGATCCGAGAGAGATAAACTCGAGTCCCTAAAGCGTG
TACTTGAAGAGAAGGAGGAGCAGCTAAAGGAAGTTGATAACAGATACAATATTTTGAGGCAAGAGTTGGAAAGTGCTAAAAAGGAGATGGAAGTTACAGT
CTCTGAGCATGCTAAAGAACTTACTGAGAAAATGGCTGAGATAAAAGGACTGGAGTCTTTGCTACAGTCACAAAAAGAAGAAAATGCGAGGGTTCAGTAT
TTGTTGCAGTCAAAAGAAGAAGAAAATAGGAGCCTTCAATCTCTACTGCAGTCAAAAGATGAAGAAATTGGGAGCCTTCAGTCTGTGTTGCAGTCGAAAG
AAGAAGACATTGTGAACCTTCAGGTTTTGATGCAGTCAAAAGAAGAAGAAATTGTTATCCTTCAGGCTTTGCTGCAGTCCAAAGAAGAAGAAACTCGGAG
CCTTCAGACTTCGCTGCAGTCAAAAGAAAACGAGACTGAGAACCTTCAGTCTTTGCTGCACTTAAAAGAAGAAGAAACAGGTAGCCTTCAGTTTCTGGTT
CAGTCGAAAGAAGAACAAATTGGGAGCCTTCAGACTTTCCTGCAGTCAAAAGAAGAAGAAATTGACAGCCTTCAGACTTTGCTGCAGTCAAAACAAGAAG
AAATTGGGGTCCTTCAGCCTTTGCTGGAGTTAAAAGAAGAAGAAATTGGTAACCTTCAGTCTCTGGTGCAGTCAAAAGAAGAAGAAATTGGGAACCTTCA
GTCTTTGGTGCATTCAAAAGAAGAGGAAATTGTTAACCTTCAGTCTTCTCTGCAATCAAAAGAAGAAGAAATTGATACCCTTCAGTCATTGCTGCAGTCA
AAGGAAGAAGTAGTGGTGAACCTTCAGTCTTTCTTGCAGTCAAAAGAAGAAGCAATTGTGAGGCTTCAGTCTTTGCTGCAGTCAAAAGAAGAAGAAACAG
GAAACCTTCAGTCTTTGTTACAGACAAAAGAAGAAGAAATTGGGAACCTTCAGATGTGCAGCAGAGAATTAGAGTCTGAACTTGTTAACCTGGAACAGGA
GAAGCATCAGCTCGAACACGACATGGAAACAGTCGTGAGAGAAAATGTTGAAGCCAGGAACCGGTTGAAAGACTTGTACAGTAGCATCGACTATCATGTT
ACTGCCAAGGAAGTTCCTGAAAGTGATCATTCATCAGAGAAGGAGGAAAGTCAAGGTTTGTCCTCCTCCATATCTGACCTGGAAGCTAAAATGTTGCACC
TCGGAAGAGAGAGGGAATCAACGGAGGAAGAAGTCAAGGACTCCAAGTCCGAGGCCAAGAGTTTGAAGGACGAGATTGTCAGATTGAGTGACGAAATTGA
TACCGAAAGGATGAACATGAAGAATCTGCTGATGCAGATCAATAATGAATGCATTACAGCACATGAGGAAAATGAGTGCCTGAAAAATGAGATTCAGAAG
TTACAAGCCACATATGAAAGCTTTAGAGAAGAGCATAATGCAATTCTGGTAAAGAATGAAGAGTTGGTAAGGGAAAACACTGATCTGGAAAGTCAACGCA
ACCATTTGGAGGCAACATTGAAGAGTACTCAATTAACCCTTGATGAGTTCTCCAAGAGAATTATAAACTTGGAAGATGAAAATGAATGTCTGGAAGAGAA
GAGTCACAGGTCACAAGCCACAACTGAAGACTTCATAGAAGAACATAATGCGTGTCTGGTGAAGAATGAAGAGTTGGCAATGATGAACACTGATCTAAAG
ACAGAGCTGAAGAATGCTCAGTTAAGCCTTGATGAGTTCGCCAAGAGGGTTATGGGTTTTGAAGAGAAAATTTATTCACTTGAGCAAGGGAGTGCCACCA
AGGTTAAGAATCTAGCTTCAAAAGTTGATCAATTGTGCAGAAAATTCGAGGAACAGGTAGAAAATCAGAAGGAAGCTAGACCGACTACTTCAGAAATGGA
ATTCAGCTTTGAAGAGCTAGCTGCTTTCAAGATAAATCAAGAGAAGCTACAGTCTGAGAATAGAAGAATGACATACCGGTTCGAAGACTACAAGTCATCT
GAAGCAAAATTGAGGACAACAATCAACGAACTCGAGTTAACGCTCACGGTTTCTGACTATGAGCACAACATTCTGAAAGAAGAAGCCATGAAGCTGAAGG
CCCAGTTGCTGAAACTAGGACCAGTCCACGAACAAGTGTTGGCGTTGAAGACCGAACTGAATGCTGCCAAGTCAGATAAAAATAAGTTCAAATCTTCGTT
GAATCTAGTCTCCACGGAATGTCAACATCTTGAGGCAGAGAAGAGCTCACTTATTGAAGAAATAGCTGTCCTCCATAAGTGTGTCCATCAACTAGAGGAA
CATAAGAGACAAAGATTCGCCTTGGAAGATAAGCTGAAAAATATGGAGAGGGATCTGATGGCCAAAGAAGCATTGTGGCAACGTGACGAGCAAGTCAACG
ATGAGCTCAACAAGATCAGGAAAGCCAATAAACAGCTCCATAAGATAATGGAGGAACTGGAGGATGAGAAAGATGATTATGCAGCAAAAGCCAGATCGCT
CGAGGAAGAGTTGGTGATGATTCAGGAAGAACAGTTGAACTTCTCCAGCAGTAGCAACGTGGATGAAAACGGAGATGCTGGATCATTTGAGCAGCAACTT
TCCGAGTCTAGGAATGCAAAACCCTCACTGAGAATGCACAGAAAAAGGAAGATAAAGATAAATTAG
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029303 pacid=23139944 polypeptide=Lus10029303 locus=Lus10029303.g ID=Lus10029303.BGIv1.0 annot-version=v1.0
MFRLHKIKSDRLSLGERIDFKISHFKALQVPRGWDKLYVTVISVETGKTIARLSKAVVKNGGSCHWEESFTESVWVSPDESSKVLEDCIFKFVLAMGSAR
SGILGEVTVNMSSYISSSSAVSVSFPLKKCDYATVLQATIQCLTPMANLRDAEGKETNLTDKNGKEEELRDDEEGTKSDQESDTASPKGGPSYGSSDTGS
PPVQAGPETREASSSATNSPGSEDLEEAATAGRPVLFPSRPSLVRRERRTSSGNGISSMNYVVDSPRSERSVPMRRIPRPGSRSGEIHQEFGISRLRNSA
LRTMDSSSTSQTFLEAAEETIGDLRTQAKMWERTARKLMLEMDYLKKENVELSRNQGNQDMEFTEACAERDGLRKEVEQLKQLLAKAKMKPPPLDNPSEG
TWQMVKELEREISFQKETSGNLTMQLKRSQESNVELVTVLQDLEETLEKQKIEIESLYDMESRFSKLEDAMEASAASNKNLKVQVLKLQESEKDLLNKVR
ELESEKDSTTTDSVSIREREVESIKMEMQGRVRELTEELDGVRAEVKRLEAGLVLKQVENKDLQRCRTGAEARVSVLEKEKEHLELELETVTRENVIASK
CLNDLQKDLATISSNLETHASVNKNLERKSLELEQNNQEIELCLSELRQENEDLSTHILALETRLRFEASEHKSRICELEKELRKAESDFNVSAQEIERL
KLELEVKVSEMSKQVDESKAEIQSLETIMQANEEEVRDLQRWKSELAAKLSVIQGEKDQLEEKMGAIMTENDIATRRSNELQSELTTTRSSLEEKSSTNE
GLERKHLELEKQKNDLGKQLSGIEQDNQELLMNITDLETQLRSEASDHESQLNSVEESYKKLLMQIDSDCNDSLAKVEKLRMELEAEVTELKAELSDRKA
EIEILDACLHAKEEEIGQLQRHTRESEVKIFDLKEEKCQLENNMETVMEENDVAVRRLNDSQQEIIMLRSSVESHVASNNALETKCSELEGKKLELELHS
SELKQGNEQLATLLSELEAELRSERDKLESLKRVLEEKEEQLKEVDNRYNILRQELESAKKEMEVTVSEHAKELTEKMAEIKGLESLLQSQKEENARVQY
LLQSKEEENRSLQSLLQSKDEEIGSLQSVLQSKEEDIVNLQVLMQSKEEEIVILQALLQSKEEETRSLQTSLQSKENETENLQSLLHLKEEETGSLQFLV
QSKEEQIGSLQTFLQSKEEEIDSLQTLLQSKQEEIGVLQPLLELKEEEIGNLQSLVQSKEEEIGNLQSLVHSKEEEIVNLQSSLQSKEEEIDTLQSLLQS
KEEVVVNLQSFLQSKEEAIVRLQSLLQSKEEETGNLQSLLQTKEEEIGNLQMCSRELESELVNLEQEKHQLEHDMETVVRENVEARNRLKDLYSSIDYHV
TAKEVPESDHSSEKEESQGLSSSISDLEAKMLHLGRERESTEEEVKDSKSEAKSLKDEIVRLSDEIDTERMNMKNLLMQINNECITAHEENECLKNEIQK
LQATYESFREEHNAILVKNEELVRENTDLESQRNHLEATLKSTQLTLDEFSKRIINLEDENECLEEKSHRSQATTEDFIEEHNACLVKNEELAMMNTDLK
TELKNAQLSLDEFAKRVMGFEEKIYSLEQGSATKVKNLASKVDQLCRKFEEQVENQKEARPTTSEMEFSFEELAAFKINQEKLQSENRRMTYRFEDYKSS
EAKLRTTINELELTLTVSDYEHNILKEEAMKLKAQLLKLGPVHEQVLALKTELNAAKSDKNKFKSSLNLVSTECQHLEAEKSSLIEEIAVLHKCVHQLEE
HKRQRFALEDKLKNMERDLMAKEALWQRDEQVNDELNKIRKANKQLHKIMEELEDEKDDYAAKARSLEEELVMIQEEQLNFSSSSNVDENGDAGSFEQQL
SESRNAKPSLRMHRKRKIKIN
|
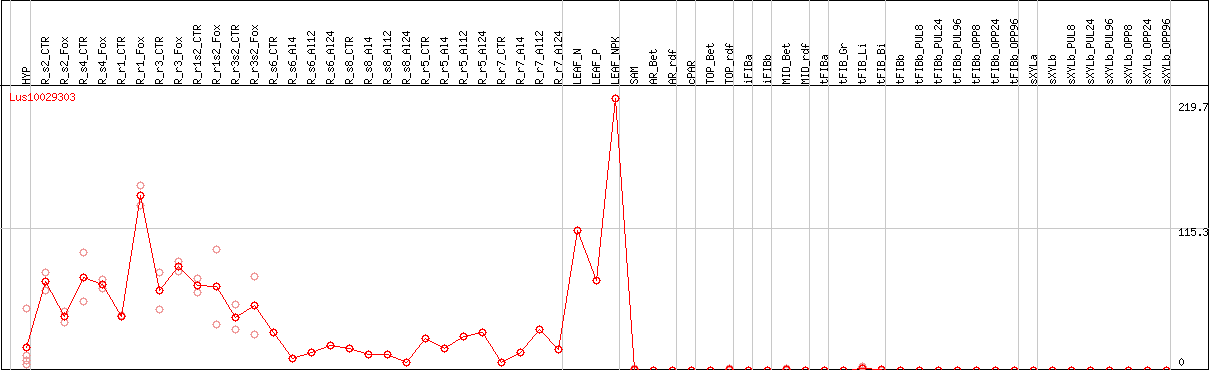
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029303 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
