Lus10029327 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029327 pacid=23139869 polypeptide=Lus10029327 locus=Lus10029327.g ID=Lus10029327.BGIv1.0 annot-version=v1.0
ATGGCAGCTCCGTCTTCTAAGAAGAGGACTCAGAACAGGAATCCTAAAGAAGCTCCGAAGTTCAACAAGGATTCTAAGAAGCCATTCAAAGCTAAGAAGA
AGGACTCTACCGATGATGTTGCCAAAAAAACTGAATCCGCCGCCATGTCTTTGCAGCTGGAAGATGACGTCCCTGACTTTCCCCGAGGTGGAGGGAGCAC
TCTAAGCCAGAAAGAGCGTGATGATATACGATCGAAAGTTGATGCGGAGTTTGAGGCTGGCGAAATGGGTGTTAGAAAGAACAAGAAGGGTAAGAAACTG
CGAAAGAAACAAGTAGACCCAGACGATTTCGGTTCCCTCTTTGGAGATGGGTTAACTGGCAAGCTGCCCAAGTTTGCTAATAAGATCACAACAAAGAATA
TTTCTCCTGGAATGAAGCTCTGGGGAGTTGTTGCTGAGGTAAATGAAAAGGACCTTGTAATTAGTCTCCCTGGAGGACTGCGTGGATTAGTTCGTAACAC
TGACGCTGTTGATTCTATTTGGGATGATGAAACTAAGGAAAATGAGGGTTATCGTCCCAACACATTCCATGTGGGGCAGCTGGTTTCCTGTATTGTCTTG
CAGTTGGATGACGATAAAAAGGGGAAAAGAAAGATATGGCTTTCTCTGTGCCTTTCATTATTACACAAGGCGTTTTCACTGGACACAATACAGGAAGGGA
TGGTACTCACATCATATGTGAAGAGCATTGAAGATCATGGCTATATTCTTCACTTCGGAGTACCTTCTTTTTCTGGGTTTCTGCCTAAGAGCAGCCAGTC
TGACTCACGTGATACTGTGGTGAAGCCTGGTCAACTCTTGAAAGGGGTTATTACTAAGATTGATAAAAACCGGAAAGTTGTTTTTTTGAGTGCTGACAAG
GATATGGTGTCGAAATATGTGATGTCGGATGTTAAAGGAATGTCAATCGATCTTCTCATTCCTGGAATGATGGTGAATGCTCGTGTACAGACAACACTTG
AAAATGGTGTTATGTTCTCCTTCCTTACATACTTCACTGGAACTGTTGATATGTTCCATTTGCAAAGTTCTTTTCCTGCTACCAACTGGAAAGATGATTA
TAGCAAGAATAAGAAGGTTATTGCTCGTATATTATTTATCGACCCGTCTACTAGAGCTGTTGGCTTGACTCTCAATCGACATCTGGTTCAAAACATAAGT
CCTCCAACGCTTGTCAAAGTCGGAGATATTTGTGACAATGCAAAAGTGGTTTGGGTAGATAAAAATATGGGGCTCTTGCTTGAGATCCCTTCTATTCCAG
TATCTATTCCAGCTTTTGTTAGTATACCTGATGTTGCTGAAGCTGAAGTGCCAAAGCTGGAAAAGAAGTTTAAGGAGCGCAGTATTGTCCGTGTGAGAAT
AATTGGTTTTAGGCATTTGGAAGGGCTTGCTACAGGAACTCTGAAGGAAAGTGCTATTGAAGGCCCTGTTTTTACTCACTCGGAAGTTCAACCAGGAATG
GTGGTGAGGGGTAAAATCATAGCAGTGGATAGCTTTGGTGCTATCGTGCAGTTTCCTGGTGGTTTGAAGGCACTTTGCCCTCTTCAGCACATGTCTGAAT
TTGAAATTGCAAAGCCGAGGAAAAAGTTCCAGGTTGCTGCCGAACTAACTTTTCGTGTGCTTGGCTGCAAGTCGAAAAGGATAACTGTTACTCACAAGAA
AACACTTGTTAAGTCAAAACTTCCAATCATTAGTTCATATGCAGAAGCTATTGATGGACTAGTTGTACATGGTTGGATAACGAAGATTGAAAAGCATGGA
TGCTTCGTCCATTTTTACAATGGAGTCCAAGGATTTGCTCCAAGGCTAGTTATACAACTTCTATACATATTTTTAGCCTTGCTTATACTTGTTATGGAAA
CGGCAACACCATATGTGGAGCTTTGTGTTACCATTTTATCTGAGCTTGGCTTAGAACCAGGAAGTGAGCCAAACTCAATGTATCACCTTGGCCAAGTTAT
CAAATGCAGAGTAACAAGCTCTGTTCCTTCTTCTCGTCGAATCAACCTAAGCTTCATGATGAAGGCCACAAGCCTTTCTGAAGAAGATTCCTTGAAGCTG
GGGAATGTCGTTACAGGAGTGGTTGAGAAAGTTACTCCATCTGTTGTTGTTGTTAATATTAATGGGAAGCAATACCTCAAGGGTACCATTGCTACTGAAC
ATATGGCTGATCATCATGGACTAGCGGTCTTGATGAAGTCGGTCTTGAAGCCTGGACATGAATTTGACAAACTCGTAGTCCTTGATATTGAGGGCAATAA
TGTAATACTCTCGGCCAAATACTCGCTCATCAATTCAGCGCAAGAGCTTCCTTCAGATATAAGTCAGATTCGTCCACACTCAATTGTCCATGGTTACATC
TGCAACATAATTGAAACCGGCTGCTTTGTTCGCTTTCTTGACCGGTTGACTGGTTTCTCCCCTAGAAGCAAGGGAATGTATCTTTCAATGGATGATTCTA
GTGTTCAGCTTTCGGAAGCATTCCACATAGGGCAGTCTGTGCGTAGTAATATCGTTGAGGTGAATAGTGAAACAAATAGAGTCACATTGTCCCTGAAGCA
GTCAATTTGTGCATCAGCTGATGCATCTTTTCTTCAGGGTTATTTTCTCTCTGATGACAAGATTGCAAAGTTCCAGTCAACTGATGTCGAAGGTCCAGGC
TTCCAATGGGTTGAAGAATTTGGAGTTGGCAGCATCACTGAAGGAAGAATCCAAGAGTCCAAAGAGTTTGGAGTTGTCGTGAGCTTCGACAAATATAAAA
ATGTTCTTGGTTTTATTACAAATTATCAATTGGGCGGGACGACCATCGGCACAGGATCGATCATACAAGCAGCTATACTAGACGTTGCCAGGAGTGAACG
CCTAGTTGACTTAACCTTGAAACCAGAGCTCTTGAATAAATTCCGAGATCAGAGCTCCAGTAAAAAGAGTCAAAAGAAGAAACGCAAAAGTGAAGCGTGT
AAGGACTTGGAAGTGAATCAAAAAGTGAATGCCATTGTAGAGATTGTGAAGGAGAATTACATGGCAAGTGTTCTTTCACTCCCGGAATACAATTACGCAA
TAGGCTATGCTTCATTATCCGACTACAATTCTCAGAGACTGCCTCACAAGCAAATTGCGAATGGACAAAGTATTACAGCTACCATTGTAGCCCTTCCAAG
CCCTTCCACAACTGGCAGGCTGCTTTTGATGCTCAATTCTACCGGCCAGGTTACTGAGACATCCAGTTCAAAGAAGGCAAAAAAGAAGGCTAGCTATGGT
GTTGGATCACTTGTTCAAGTGGAGGTTAATGAAAAGAAGCCACTTGAAATGCGAGTGAAATTTGGTGCTGGTCTTCGTGGAAGAATCCATGCGACGGAGG
CAACTAATGCTGGAGTCATTGAAGACCCATTTAGTGAATTCCGAACTGGTCAAACTTTGACGGCTAGGATTGTTGCAAAGTCTGGTCAACCGAACATTAA
GAACGGTGTGCTGTGGGAACTGTCTCTGAAAGCTGAAGTGCTGGCAGATGAAAGTGACAACATATCTGACAACCATGAGTTTTCATCTGGACAATGTGTC
GGCGGCTATGTATATAAAGTAGACAGTGAATGGGTTTGGTTAGCCATATCTCGAACCGTAAATGCACAACTATTTATATTAGATAGTTCTTCTGAACCTG
GTGAGCTTCAGGAATTTCAGAACCTTTTTGGTGTGGGAAAGCTTGTCAGGGGGTATGTTTTAACATACAACAGAGCAAAGAAATTGCTTCGTCTTGGTAT
GCATCCATTACGTCCGCTCAAGCAAATTGACAATGTGGCCCCTCATATTCACGAAGGAGATGTGGTCGGGGGGAGAATTTCAAGGGCACTTCCAGGTGTA
GGTGGACTGTTTGTCCAAATTGGCCCTCATTTACATGGTAAAGTTCATTTTACTGAACTTAAAGATGAGTGGACTGCTGACCCTTTGGCTGAGTATCCTG
AAGGACAGTTTGTCAGATGCAAGGTTCTAGAGATCGGCCTGTCAATCAAGGGCACTGCTCATATAGATCTGTCACTACGTTCATCTTTTGATGGCATGCG
CGGTCTGAGCTCAGTGGAATCATCTGGAAACACGCATGTCGAAAAGATTGAGGATCTTCACCCAAATGATGTTGTTCAGGGTTATGTGAAGAGTATTACG
GCAAAAGGATGTTTCATTTTGCTTTCACGTAAGCTCGAAGCAAAAATACTTGTCTCAAATCTATCTGATGGATATATTGAGAAACCCGAAACAGAATTCC
CCGTGGGTAAACTTGTTGTTGGAAGGGTATTGTCCGTGGAGCCTTTGTCAAAACGAATTGAAGTTACTCTGAGGACAACAAAACCTGGCAATGAATCAAA
ATCTCAGGGGAATGATTTAAGCCACCTGCGTGTTGGAAGAATAGTTTCTGGCTCGATTAGGCGAGTGGAACCATATGGTCTATTTGTAACACTAAGCAAT
TCAAACATGGTTGGGTTGTGTCACATATCGGAGCTTCCAGCTGATCAGGCCAACGACACTCTAAGTAACTATAAAGCTGGGGAGAAGGTTACGGTGAAAA
TTCTACAGGTTGATGAGGAAAGACGTCGTATCTCTCTTGGAATGACAGACTTGGATATGGAAAACAACGACGATGAATCTGTTGAAGGTGATGAGGAAGT
CGGTGAGTCGGATGATGATGACACTGACACCAACATTACACTGCTACCCGAGAGAGACGATTCAGAATCAGAAGACGAAAGTAAAGAATGTCCATTTCCC
GAAGCAGAATCTAGAGCTTCCATTCCTCCTTTGGAGGTCACCCTGGATGATATGGAAGAGGATCAAGCAGACGATCGGGTAGCTTCAGATAACCAGGATC
ATAACGTTGAGCCTGGTACTAAAGACAAAAAGAAAAAGGAAAAATGTAAAACGAGGGGAGAAAGGGAGAAAGAGATCAGAGCTGCTGAAGAAAGACTGCT
GGAAAAGGATGTTGTACCAGAAACTGCTGATGAGTTTGAGAAACTCCTGCGTGGCTCTCCCAATAGTAGCTTTATTTGGATCAAGTACATGGCTTTCTTG
CTTAATACGTCTGATGTAGAGAAAGCTCGTTCAGTGGCTGAAAGGGCTCTGAGGACGATAAGTTTCCGTGAAGAGGGCGAGAAACTGAATATCTGGGTAG
CTTACTTCAATTTGGAAAACCAGTACGGAGTTCCTCGAGAGGAGGCGGTCAACAAATTATTCACAAGAGCATTGCAATTCTGCGATTCGAAGAAGGTTCA
TATGGCACTATTGGGGATGTACGAGAGGACTGAACAAGATAAGTCTGCTGATGAACTTGCCAGCAAAATGGTCAGGAAGTTTAAGCAATCATGCAAGGTT
TGGTTAGCGAGAATGCGGAGGGTTATAAGCCAAGAGGAAGACAAAGTTCAGTCAGTTGTACAACGAGCCCTAATGTGCCTTCCACGGCGTAAGCACATCA
AATTCATCACGCAAGCTGCACTCCTCGAGTTCAAAAGCGGGCTTGCTGATAGAGGCCGGAGCATGTTTGAAGGAGTTTTGAGAGAGTACCCGAAGCGAAC
GGACTTGTGGAGCATATACATCGACCAAGAAATCAAGCATGGAGATGCTGATATGATCCGTTCATTGTTTGAGAGAGCAATTACGCTGAGCCTAGCGCCT
AAAAAGATGACATTCTTGTTCAAGAAGTATCTCGAGTACGAGAAGTCAATGGGCGACGTTGAAAAAATCGAGGCTGTGAAGCAGAAAGCCCTCGACTATG
CTCGATCTGAACTGGTTTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029327 pacid=23139869 polypeptide=Lus10029327 locus=Lus10029327.g ID=Lus10029327.BGIv1.0 annot-version=v1.0
MAAPSSKKRTQNRNPKEAPKFNKDSKKPFKAKKKDSTDDVAKKTESAAMSLQLEDDVPDFPRGGGSTLSQKERDDIRSKVDAEFEAGEMGVRKNKKGKKL
RKKQVDPDDFGSLFGDGLTGKLPKFANKITTKNISPGMKLWGVVAEVNEKDLVISLPGGLRGLVRNTDAVDSIWDDETKENEGYRPNTFHVGQLVSCIVL
QLDDDKKGKRKIWLSLCLSLLHKAFSLDTIQEGMVLTSYVKSIEDHGYILHFGVPSFSGFLPKSSQSDSRDTVVKPGQLLKGVITKIDKNRKVVFLSADK
DMVSKYVMSDVKGMSIDLLIPGMMVNARVQTTLENGVMFSFLTYFTGTVDMFHLQSSFPATNWKDDYSKNKKVIARILFIDPSTRAVGLTLNRHLVQNIS
PPTLVKVGDICDNAKVVWVDKNMGLLLEIPSIPVSIPAFVSIPDVAEAEVPKLEKKFKERSIVRVRIIGFRHLEGLATGTLKESAIEGPVFTHSEVQPGM
VVRGKIIAVDSFGAIVQFPGGLKALCPLQHMSEFEIAKPRKKFQVAAELTFRVLGCKSKRITVTHKKTLVKSKLPIISSYAEAIDGLVVHGWITKIEKHG
CFVHFYNGVQGFAPRLVIQLLYIFLALLILVMETATPYVELCVTILSELGLEPGSEPNSMYHLGQVIKCRVTSSVPSSRRINLSFMMKATSLSEEDSLKL
GNVVTGVVEKVTPSVVVVNINGKQYLKGTIATEHMADHHGLAVLMKSVLKPGHEFDKLVVLDIEGNNVILSAKYSLINSAQELPSDISQIRPHSIVHGYI
CNIIETGCFVRFLDRLTGFSPRSKGMYLSMDDSSVQLSEAFHIGQSVRSNIVEVNSETNRVTLSLKQSICASADASFLQGYFLSDDKIAKFQSTDVEGPG
FQWVEEFGVGSITEGRIQESKEFGVVVSFDKYKNVLGFITNYQLGGTTIGTGSIIQAAILDVARSERLVDLTLKPELLNKFRDQSSSKKSQKKKRKSEAC
KDLEVNQKVNAIVEIVKENYMASVLSLPEYNYAIGYASLSDYNSQRLPHKQIANGQSITATIVALPSPSTTGRLLLMLNSTGQVTETSSSKKAKKKASYG
VGSLVQVEVNEKKPLEMRVKFGAGLRGRIHATEATNAGVIEDPFSEFRTGQTLTARIVAKSGQPNIKNGVLWELSLKAEVLADESDNISDNHEFSSGQCV
GGYVYKVDSEWVWLAISRTVNAQLFILDSSSEPGELQEFQNLFGVGKLVRGYVLTYNRAKKLLRLGMHPLRPLKQIDNVAPHIHEGDVVGGRISRALPGV
GGLFVQIGPHLHGKVHFTELKDEWTADPLAEYPEGQFVRCKVLEIGLSIKGTAHIDLSLRSSFDGMRGLSSVESSGNTHVEKIEDLHPNDVVQGYVKSIT
AKGCFILLSRKLEAKILVSNLSDGYIEKPETEFPVGKLVVGRVLSVEPLSKRIEVTLRTTKPGNESKSQGNDLSHLRVGRIVSGSIRRVEPYGLFVTLSN
SNMVGLCHISELPADQANDTLSNYKAGEKVTVKILQVDEERRRISLGMTDLDMENNDDESVEGDEEVGESDDDDTDTNITLLPERDDSESEDESKECPFP
EAESRASIPPLEVTLDDMEEDQADDRVASDNQDHNVEPGTKDKKKKEKCKTRGEREKEIRAAEERLLEKDVVPETADEFEKLLRGSPNSSFIWIKYMAFL
LNTSDVEKARSVAERALRTISFREEGEKLNIWVAYFNLENQYGVPREEAVNKLFTRALQFCDSKKVHMALLGMYERTEQDKSADELASKMVRKFKQSCKV
WLARMRRVISQEEDKVQSVVQRALMCLPRRKHIKFITQAALLEFKSGLADRGRSMFEGVLREYPKRTDLWSIYIDQEIKHGDADMIRSLFERAITLSLAP
KKMTFLFKKYLEYEKSMGDVEKIEAVKQKALDYARSELV
|
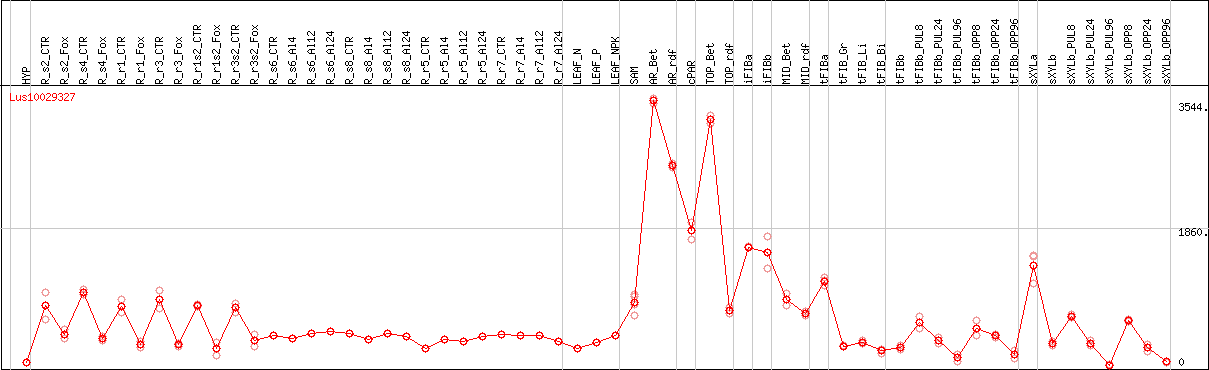
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029327 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
