External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G57670 292 / 1e-97
C2H2ZnF
WIP2, NTT
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
AT3G20880 287 / 3e-95
C2H2ZnF
WIP4
WIP domain protein 4 (.1)
AT1G51220 283 / 7e-95
C2H2ZnF
AtWIP5, WIP5
WIP domain protein 5 (.1)
AT1G08290 267 / 2e-88
C2H2ZnF
WIP3
WIP domain protein 3 (.1)
AT1G13290 265 / 3e-88
C2H2ZnF
WIP6, DOT5
WIP domain protein 6, DEFECTIVELY ORGANIZED TRIBUTARIES 5, C2H2-like zinc finger protein (.1)
AT1G34790 263 / 4e-87
C2H2ZnF
WIP1, TT1
WIP domain protein 1, transparent testa 1, C2H2 and C2HC zinc fingers superfamily protein (.1)
AT1G34370 101 / 2e-24
C2H2ZnF
STOP1
sensitive to proton rhizotoxicity 1, C2H2 and C2HC zinc fingers superfamily protein (.1.2.3)
AT5G22890 94 / 9e-22
C2H2ZnF
C2H2 and C2HC zinc fingers superfamily protein (.1)
AT5G03150 92 / 6e-21
C2H2ZnF
JKD
JACKDAW, C2H2-like zinc finger protein (.1)
AT1G55110 90 / 4e-20
C2H2ZnF
ARABIDOPSISTHALIANAINDETERMINATE(ID)-DOMAIN7, ATIDD7, ARABIDOPSISTHALIANAINDETERMINATE(ID)-DOMAIN7
indeterminate(ID)-domain 7 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039613
425 / 3e-151
AT3G57670 300 / 9e-101
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Lus10031838
294 / 2e-98
AT3G57670 386 / 4e-133
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Lus10031271
294 / 3e-98
AT3G57670 379 / 6e-131
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Lus10004887
286 / 3e-95
AT1G08290 357 / 9e-123
WIP domain protein 3 (.1)
Lus10020597
281 / 2e-93
AT1G08290 335 / 7e-114
WIP domain protein 3 (.1)
Lus10020923
275 / 6e-92
AT1G34790 312 / 2e-106
WIP domain protein 1, transparent testa 1, C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10033454
275 / 8e-92
AT1G34790 315 / 1e-107
WIP domain protein 1, transparent testa 1, C2H2 and C2HC zinc fingers superfamily protein (.1)
Lus10035044
275 / 8e-92
AT1G13290 338 / 1e-116
WIP domain protein 6, DEFECTIVELY ORGANIZED TRIBUTARIES 5, C2H2-like zinc finger protein (.1)
Lus10021700
275 / 9e-92
AT1G13290 333 / 2e-114
WIP domain protein 6, DEFECTIVELY ORGANIZED TRIBUTARIES 5, C2H2-like zinc finger protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G267900
310 / 2e-105
AT3G57670 328 / 2e-111
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Potri.009G062300
298 / 8e-101
AT3G57670 336 / 3e-114
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Potri.003G205000
297 / 2e-100
AT3G57670 377 / 2e-130
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Potri.016G052700
296 / 2e-99
AT3G57670 389 / 7e-135
WIP domain protein 2, NO TRANSMITTING TRACT, C2H2-type zinc finger family protein (.1)
Potri.001G018900
293 / 8e-99
AT1G51220 353 / 7e-122
WIP domain protein 5 (.1)
Potri.009G143700
290 / 1e-97
AT1G08290 364 / 7e-126
WIP domain protein 3 (.1)
Potri.004G183900
289 / 4e-97
AT1G08290 352 / 4e-121
WIP domain protein 3 (.1)
Potri.010G129000
280 / 1e-94
AT1G13290 352 / 1e-122
WIP domain protein 6, DEFECTIVELY ORGANIZED TRIBUTARIES 5, C2H2-like zinc finger protein (.1)
Potri.002G098200
271 / 8e-91
AT1G34790 311 / 3e-106
WIP domain protein 1, transparent testa 1, C2H2 and C2HC zinc fingers superfamily protein (.1)
Potri.013G114600
103 / 1e-24
AT1G34370 495 / 3e-172
sensitive to proton rhizotoxicity 1, C2H2 and C2HC zinc fingers superfamily protein (.1.2.3)
PFAM info
Representative CDS sequence
>Lus10029525 pacid=23149402 polypeptide=Lus10029525 locus=Lus10029525.g ID=Lus10029525.BGIv1.0 annot-version=v1.0
ATGGCAGACCCTTCTGCCTCTGAAACTTCTGCTGCCGTTGGATGGTTCAATTTCTTCCCTCTCACTACCAACACTACAAAAGGAGCCTCATTATCATCAT
CCATTGATCTTAATGGTCCTCCTCCTTCTACTCCGCTACTCATTGACTATTTCAGCCATGGCAAACAAAAGGAGCCTCGTTATCATCATCCTAATAGTAA
TAGTACCATTTCTCTGCACATTGGTCCTCCTCCTCCTCCTAGTTGGGTTTCTTCTTCAACTTCTTCTTCCTCGTCTTCCTCTTCTTCTGCTAATAAAACA
TGTAGCGGCAGTAACAACATCAATCGAGAAGATGCACATTTCTCATCTTTACTGGTAAACAAGAGTGAGCAATACTGGATTCCAACTCCAACCCAGATCC
TCTTTGGTCCTTCTCAATTCTCCTGCCATCTCTGCTCCAAGTCCTTCAACCGTTACAACAACTTGCAGATGCATATGTGGGGGCACGGTTCACAGTACAG
AAAAGGACCCGACTCCTTAAAAGGAAGCCAGCCAACCGCAATGCTGAGGCTCCCATGCTACTGCTGCGCAGCGGGGTGCAAGCACAACATAGATCACCCA
AGGGCTAAAGCCTTGAAGGACTTCAGGACCCTCCAAACTCACTACAAGCGTAAGCACGGCATCAAGCCCTTCATGTGCAGGAAATGCGGGAAGCCCTTTG
CAGTCAAAGGAGACTGGCGTACCCACGAGAAGAATTGTGGCCGCATCTGGTACTGCATTTGCGGCTCCGACTTCAAGCACAAGCGGTCGTTGAAAGATCA
TGTCAAGTCCTTTGGCCACTCCAGCCGTGATCGAGAGGAGGATGAGGATGAGGACGAGCTCGACGGTGATCCTACGGTAAATATATAA
AA sequence
>Lus10029525 pacid=23149402 polypeptide=Lus10029525 locus=Lus10029525.g ID=Lus10029525.BGIv1.0 annot-version=v1.0
MADPSASETSAAVGWFNFFPLTTNTTKGASLSSSIDLNGPPPSTPLLIDYFSHGKQKEPRYHHPNSNSTISLHIGPPPPPSWVSSSTSSSSSSSSSANKT
CSGSNNINREDAHFSSLLVNKSEQYWIPTPTQILFGPSQFSCHLCSKSFNRYNNLQMHMWGHGSQYRKGPDSLKGSQPTAMLRLPCYCCAAGCKHNIDHP
RAKALKDFRTLQTHYKRKHGIKPFMCRKCGKPFAVKGDWRTHEKNCGRIWYCICGSDFKHKRSLKDHVKSFGHSSRDREEDEDEDELDGDPTVNI
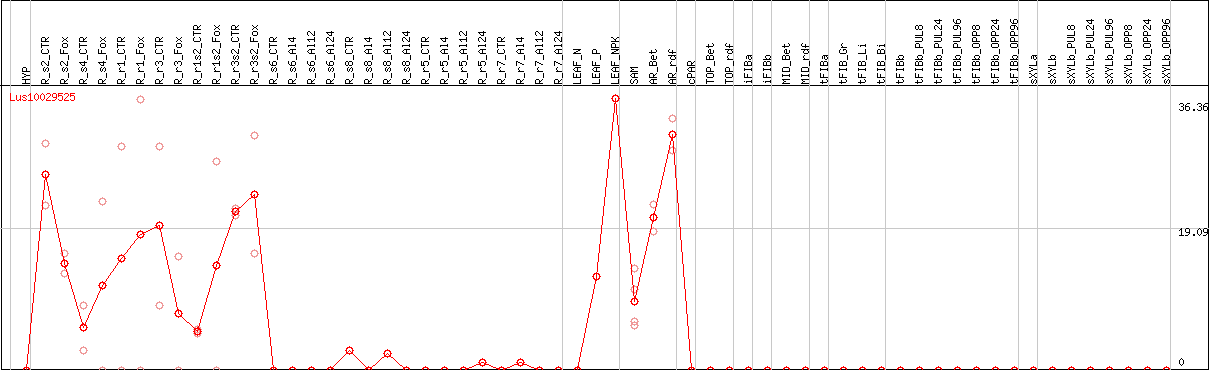
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029525 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.