External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G26330 122 / 7e-33
ATSBT3.18, UNE17
UNFERTILIZED EMBRYO SAC 17, Subtilisin-like serine endopeptidase family protein (.1)
AT2G04160 119 / 6e-32
AIR3
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
AT5G59810 117 / 4e-31
ATSBT5.4
Subtilase family protein (.1)
AT1G04110 116 / 7e-31
SDD1
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
AT1G32980 109 / 1e-30
Subtilisin-like serine endopeptidase family protein (.1)
AT3G14240 115 / 2e-30
Subtilase family protein (.1)
AT3G14067 115 / 3e-30
Subtilase family protein (.1)
AT4G10510 114 / 8e-30
Subtilase family protein (.1)
AT5G67360 112 / 2e-29
ARA12
Subtilase family protein (.1)
AT5G51750 112 / 2e-29
ATSBT1.3
subtilase 1.3 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G006570
129 / 3e-35
AT1G04110 568 / 0.0
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
Potri.001G113700
129 / 3e-35
AT1G04110 538 / 0.0
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
Potri.003G118500
129 / 4e-35
AT1G04110 557 / 0.0
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
Potri.003G118800
125 / 4e-34
AT1G04110 585 / 0.0
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
Potri.003G118700
124 / 2e-33
AT1G04110 585 / 0.0
STOMATAL DENSITY AND DISTRIBUTION, Subtilase family protein (.1)
Potri.011G165900
123 / 3e-33
AT2G04160 882 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.014G026600
122 / 6e-33
AT5G67360 546 / 0.0
Subtilase family protein (.1)
Potri.014G171600
122 / 8e-33
AT2G04160 986 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.011G050300
122 / 1e-32
AT2G04160 786 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.001G002200
120 / 3e-32
AT4G26330 891 / 0.0
UNFERTILIZED EMBRYO SAC 17, Subtilisin-like serine endopeptidase family protein (.1)
PFAM info
Representative CDS sequence
>Lus10029569 pacid=23152107 polypeptide=Lus10029569 locus=Lus10029569.g ID=Lus10029569.BGIv1.0 annot-version=v1.0
ATGTCGTGTCCACATCTTAGCGGTGTTGCAACTTTAATCAAGGCAATGCATCCAGACTGGTCACCTGCTGCAATAAAGTCAGCGATTTTAACCACTGCTT
ACACAAAGGACCTTAGTGGTAAGCCGATCTCCAATGAGCAGAATTCTCCTGCAACTGACTTTGATATGGGTGCTGGACATGTGAACCCACCTAAAGCTAT
GGATCCAGGTTTGATATATGATATCCAACTGGATGATTATGTTGCGTATATCTGCAGTTTGGGTTACAACGATTCATTTGTGGCCTTACTGCCTAAAAAC
TTGATCCTGGAATGTAAACCTCGTGTTATGATAATAGACGTTGTGCTTTTCAGGTGCATCGTGTTCAAGCTTCTAAAGATACTCAGTCAACCACCACAAC
AAGATACTGCTGTGGAGAGAAAATGGGTTACAATATGTTTTAAGAAGATCAGTTTGCTTTCCAGAAGATGGTAA
AA sequence
>Lus10029569 pacid=23152107 polypeptide=Lus10029569 locus=Lus10029569.g ID=Lus10029569.BGIv1.0 annot-version=v1.0
MSCPHLSGVATLIKAMHPDWSPAAIKSAILTTAYTKDLSGKPISNEQNSPATDFDMGAGHVNPPKAMDPGLIYDIQLDDYVAYICSLGYNDSFVALLPKN
LILECKPRVMIIDVVLFRCIVFKLLKILSQPPQQDTAVERKWVTICFKKISLLSRRW
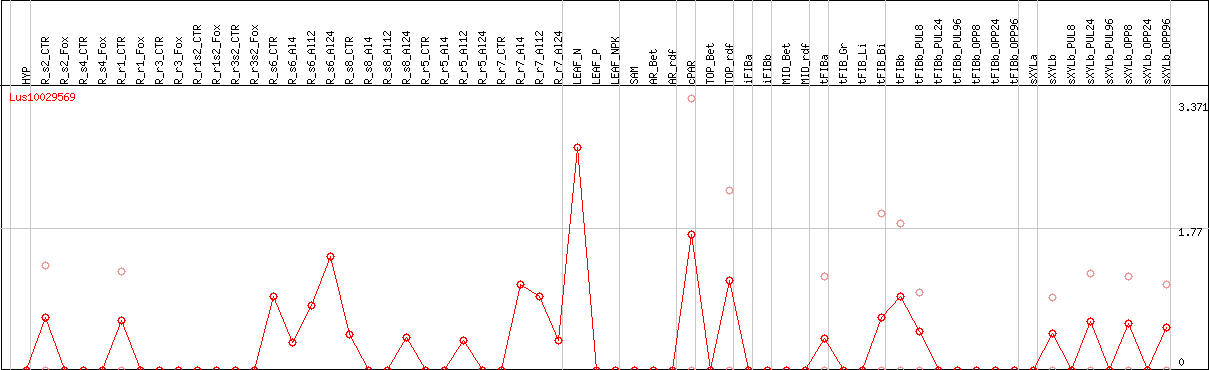
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029569 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.