External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G43700 193 / 7e-59
bZIP
SUE3, AtbZIP51, VIP1
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
AT2G31370 174 / 2e-51
bZIP
AtbZIP59, PosF21
Basic-leucine zipper (bZIP) transcription factor family protein
AT2G40620 174 / 2e-51
bZIP
AtbZIP18
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT1G06850 170 / 3e-51
bZIP
ATBZIP52
basic leucine-zipper 52 (.1.2)
AT1G06070 174 / 9e-51
bZIP
AtbZIP69
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT4G38900 171 / 1e-48
bZIP
AtbZIP29
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
AT2G21230 159 / 3e-44
bZIP
AtbZIP30
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
AT2G42380 95 / 5e-22
bZIP
ATBZIP34
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2)
AT3G58120 91 / 1e-20
bZIP
ATBZIP61
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT2G12940 91 / 1e-20
bZIP
UNE4
unfertilized embryo sac 4, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042802
405 / 7e-142
AT1G43700 313 / 1e-105
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Lus10028595
271 / 2e-89
AT1G43700 289 / 1e-96
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Lus10018899
266 / 1e-87
AT1G43700 287 / 7e-96
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Lus10008605
178 / 6e-53
AT2G40620 276 / 1e-90
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10009907
177 / 6e-53
AT1G06850 270 / 2e-89
basic leucine-zipper 52 (.1.2)
Lus10042211
176 / 3e-52
AT2G40620 277 / 4e-91
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10024204
175 / 3e-52
AT1G06850 271 / 5e-90
basic leucine-zipper 52 (.1.2)
Lus10042057
169 / 1e-47
AT4G38900 522 / 0.0
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
Lus10018061
169 / 1e-47
AT4G38900 529 / 0.0
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G069500
278 / 4e-92
AT1G43700 238 / 4e-76
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Potri.005G190700
264 / 7e-87
AT1G43700 219 / 2e-69
sulphate utilization efficiency 3, VIRE2-interacting protein 1 (.1)
Potri.013G091400
182 / 1e-54
AT2G40620 300 / 2e-100
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.019G130000
182 / 2e-54
AT2G40620 289 / 4e-95
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.013G156900
179 / 5e-53
AT2G40620 291 / 5e-96
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.017G027400
177 / 7e-52
AT1G06070 471 / 2e-165
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.007G130800
170 / 2e-49
AT1G06070 392 / 9e-135
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.004G163800
167 / 6e-47
AT4G38900 425 / 1e-142
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
Potri.009G125400
166 / 1e-46
AT4G38900 481 / 1e-164
Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
Potri.013G124400
93 / 2e-21
AT3G58120 293 / 2e-98
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0018
bZIP
PF07716
bZIP_2
Basic region leucine zipper
Representative CDS sequence
>Lus10029773 pacid=23152069 polypeptide=Lus10029773 locus=Lus10029773.g ID=Lus10029773.BGIv1.0 annot-version=v1.0
ATGGATTCCGCCAAGTACGGCGGCAAGCGGCCGTTGACAGATGACATAGACCAGATGCCGGAAGCCCCTCAGCGTGGGTGGCACCATCGCCGGGCTCACT
CCGACACTTCCTTCCGCTTCGACGACCTCCTGCTCTTTGACCCTTCCGACCTCGACCTATCGTCACTTGACTTCCCCAACGCCGCTCCTCCACGTGGAGC
TCCCATGGCCGTCGATTCTCCGTCGATCTCCGACGACTCGACCTCCAACGGCTGTGCTGCCCCTCCGCCTAATCCGAGGGCTAGACCGATCAATCATCTC
CGTAGCTTGTCCGTGGACTCCGACTTCTTCGACGGGTTCGGCCTAGGGTATGGAGATGAGAAGCCTGGCGAGAAAGGGATTGCTGCTGCTGCGGGAACTG
GAGGAGGTGAAAGGAGAGTCCATCACAGGCACAGCAGCTCCATGGACGGTTCTAGCTGTTCGGCTTCGTTTGAGGGGGATTCTTTTATGGATGATGGTAT
GAAGAAGGCCATGGCTCCTGATAGACTTGCTGAGCTCTCTCTTATTGATCCCAAGAGAGCTAAAAGGATTCTTGCTAATAGGCAATCTGCAGCTCGGTCA
AAGGAGAGGAAAATACGGTATACGGGTGAGCTTGAGAGGAAGGTTCATACACTTCAGACCGAAGCAACCACCCTCTCTGCCCAAGTCACGGTTCTCCAGA
GGGATACAAGTGGATTAAATACTGAGAACAGAGAGTTGAAACTGCGGTTGCAAGCTATGGAGCAACAAGCGCATCTTAGAGATGCACTGAATGAAGCACT
AAGGGAAGAAGTGCAGCGTCTGAAGATAGCAGCTGGTCAAATGCCAGTTTCAAATGGAAACCCATTCCACCGCCCCCAATATTCTTCACAACAGCCACCT
CCGCAGCAAACCCAGCAGCAGGGTCTTCCGCCACAGTCACAGCCTCCTAACAGCCAGAACTCAAATGGGCAACCCCGCCGTGGCTTCCCCAACTTTGGGC
AGAGGGTGTAG
AA sequence
>Lus10029773 pacid=23152069 polypeptide=Lus10029773 locus=Lus10029773.g ID=Lus10029773.BGIv1.0 annot-version=v1.0
MDSAKYGGKRPLTDDIDQMPEAPQRGWHHRRAHSDTSFRFDDLLLFDPSDLDLSSLDFPNAAPPRGAPMAVDSPSISDDSTSNGCAAPPPNPRARPINHL
RSLSVDSDFFDGFGLGYGDEKPGEKGIAAAAGTGGGERRVHHRHSSSMDGSSCSASFEGDSFMDDGMKKAMAPDRLAELSLIDPKRAKRILANRQSAARS
KERKIRYTGELERKVHTLQTEATTLSAQVTVLQRDTSGLNTENRELKLRLQAMEQQAHLRDALNEALREEVQRLKIAAGQMPVSNGNPFHRPQYSSQQPP
PQQTQQQGLPPQSQPPNSQNSNGQPRRGFPNFGQRV
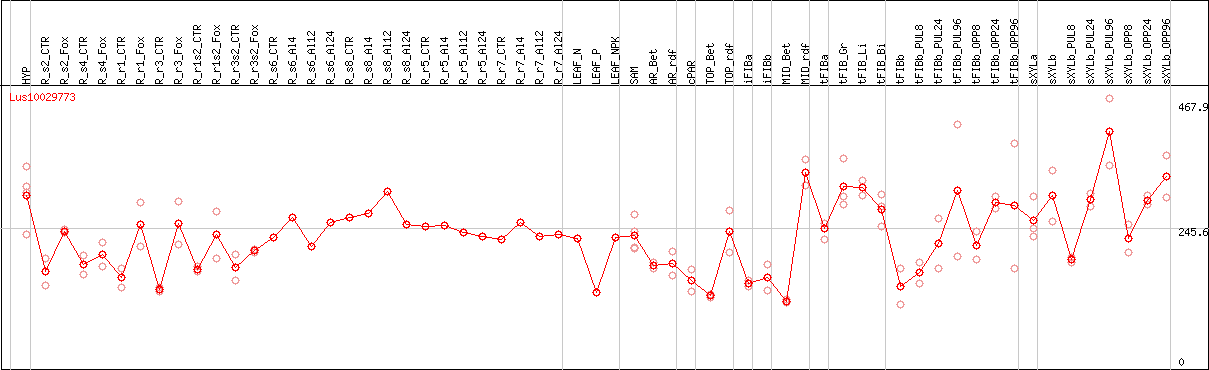
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029773 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.