External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G39050 77 / 2e-16
PMAT1
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
AT3G29635 69 / 2e-13
HXXXD-type acyl-transferase family protein (.1)
AT3G29590 67 / 5e-13
AT5MAT
HXXXD-type acyl-transferase family protein (.1)
AT3G29670 67 / 9e-13
PMAT2
phenolic glucoside malonyltransferase 2, HXXXD-type acyl-transferase family protein (.1)
AT5G61160 65 / 4e-12
AACT1
anthocyanin 5-aromatic acyltransferase 1 (.1)
AT3G29680 65 / 4e-12
HXXXD-type acyl-transferase family protein (.1)
AT1G03495 62 / 3e-11
HXXXD-type acyl-transferase family protein (.1)
AT1G03940 57 / 1e-09
HXXXD-type acyl-transferase family protein (.1)
AT5G39080 57 / 2e-09
HXXXD-type acyl-transferase family protein (.1)
AT5G39090 56 / 4e-09
HXXXD-type acyl-transferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020721
350 / 5e-120
AT5G39050 280 / 2e-89
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Lus10036582
276 / 5e-88
AT3G01090 834 / 0.0
SNF1-RELATED PROTEIN KINASE 1.1, SNF1 kinase homolog 10 (.1.2.3)
Lus10035802
264 / 4e-86
AT5G39050 276 / 2e-87
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Lus10033599
218 / 8e-69
AT3G29635 307 / 1e-99
HXXXD-type acyl-transferase family protein (.1)
Lus10020514
111 / 2e-28
AT3G29590 296 / 9e-96
HXXXD-type acyl-transferase family protein (.1)
Lus10005362
108 / 2e-27
AT3G29635 292 / 4e-94
HXXXD-type acyl-transferase family protein (.1)
Lus10021388
100 / 3e-24
AT3G29635 278 / 3e-88
HXXXD-type acyl-transferase family protein (.1)
Lus10021390
96 / 8e-23
AT3G29590 283 / 7e-91
HXXXD-type acyl-transferase family protein (.1)
Lus10021389
89 / 1e-20
AT3G29590 295 / 4e-95
HXXXD-type acyl-transferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G103200
114 / 1e-29
AT5G39050 374 / 1e-125
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G103300
113 / 4e-29
AT5G39050 352 / 3e-117
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G120700
112 / 9e-29
AT5G39050 343 / 1e-113
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.001G455150
108 / 9e-29
AT5G39050 191 / 7e-58
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G103100
110 / 6e-28
AT5G39050 366 / 1e-122
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G110400
103 / 1e-25
AT5G39050 363 / 2e-121
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G110700
103 / 2e-25
AT5G39050 369 / 1e-123
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G109300
103 / 2e-25
AT5G39050 365 / 3e-122
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G110640
102 / 3e-25
AT5G39050 365 / 2e-122
phenolic glucoside malonyltransferase 1, HXXXD-type acyl-transferase family protein (.1)
Potri.004G096132
102 / 4e-25
AT3G29670 308 / 2e-100
phenolic glucoside malonyltransferase 2, HXXXD-type acyl-transferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0149
CoA-acyltrans
PF02458
Transferase
Transferase family
Representative CDS sequence
>Lus10029797 pacid=23159687 polypeptide=Lus10029797 locus=Lus10029797.g ID=Lus10029797.BGIv1.0 annot-version=v1.0
ATGAATTCAGATGACATTGCGTTGGAGTACTGGAACAATTGGTTCCAACTCCGGCAGCAGCTTCCAGGTATGGATCCGAATCCAAATCCTAGAAGCTTGG
AGCTTCTATGGAGTTTGGTAGACAGCGAATCGAAACGATTGCGATCGGCATTCCACTTCTCCAGAGAGAGAATCGACAAATTGAAGGAAGCATTGATCCC
CAAAATAGAAAATCCTAATTACTTATCCAGCTTCGTAATCACTCTGGCTTACACGGCAGCTTGCATCGTGAAAACGGAACTGAGAGGCGAGTCGGCTACG
ATCCAGCTAGCGTTCACTGCAGGTTTCAGATCCCGATTGGATCCTCCGGTTCCGGATAATTTCTTCGGCAACTGCGTGTCCCCGTTCGAAGTGGTCCTGC
GTGCGGCGGAGGCGGTGGACGAAGACGGCGGCATGGCATACGCGGCGGGGAAGATCATAGAGACGATCAAGGTGTTGGAGCGAGGAGTTATGGTGGGACC
GAAGGAGAGGCTTGCCAATTTTCTGTTTACGGGAAAACCGGAATTTGCGGTCGGAGTGTCAGGGTGGCCAGGCTGGGGATGTATGGAGTGGATTTCGGGT
ACGGAAGGAAATGGATGTATTCTCTGCTGCATTCAGCGGAGAAAGTGA
AA sequence
>Lus10029797 pacid=23159687 polypeptide=Lus10029797 locus=Lus10029797.g ID=Lus10029797.BGIv1.0 annot-version=v1.0
MNSDDIALEYWNNWFQLRQQLPGMDPNPNPRSLELLWSLVDSESKRLRSAFHFSRERIDKLKEALIPKIENPNYLSSFVITLAYTAACIVKTELRGESAT
IQLAFTAGFRSRLDPPVPDNFFGNCVSPFEVVLRAAEAVDEDGGMAYAAGKIIETIKVLERGVMVGPKERLANFLFTGKPEFAVGVSGWPGWGCMEWISG
TEGNGCILCCIQRRK
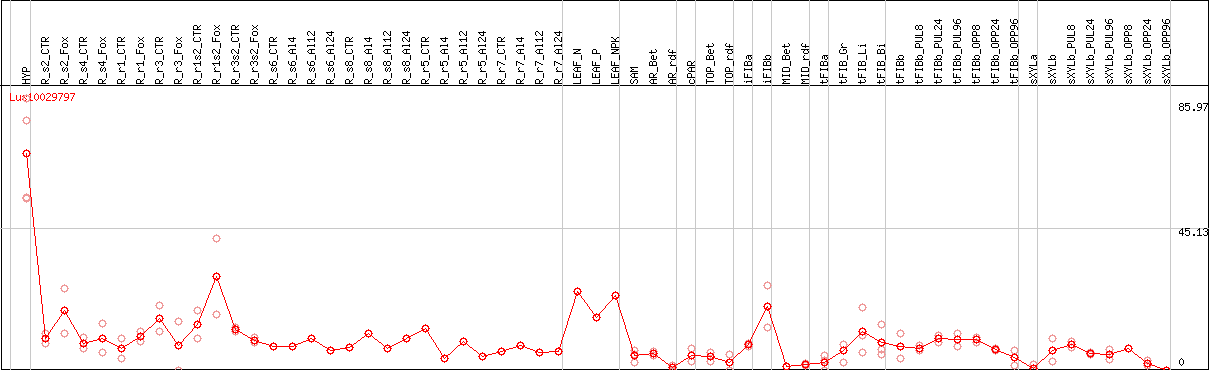
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029797 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.