Lus10029800 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029800 pacid=23159770 polypeptide=Lus10029800 locus=Lus10029800.g ID=Lus10029800.BGIv1.0 annot-version=v1.0
ATGTTGAGCTTCCATGTTGGTGGGAAAGTGGTGGATAAAGTTGATTTATTGAGGAAGAAGCATTGGACATGGCGACTGGATGTGTGGCCGTTTGCTATTA
TCTATGCTCTGTGGCTCACTACTGTCATCCCGAGCCTCGACATTGGTGATGCCTTCATTGTTTTCGGCGGCCTTGTTGCATTCCACATTCTGGTCTTGCT
CTTCACAGCTTGGTCTGTTGATTTCAGGTGCTTCGTTCAGTTCAGTAAGGCTAATGATATTCATCAAGCAGATGCATGTAAGATAACTCCAGCAAAGTTT
TCAGGATCCAAAGAAGTTGTGCCCCTTTATATTAGAAAACAAGCAGGAGGGTCTTCATCTGCAGAGGATCAGGATGAGATTTTCTTTGATTTCCGAAAGC
AGTGCTTTATATATTCAGATGAGAAGAAGACATTTTGCAAGCTTCCGTATCCAACGAAAGAAAAATTTGGTTACTATCTTAAATGCACCGGCCATGGCTC
TGAAGCCAAAGTACTCACTGCTAGTGAAAAATGGGGGCGAAATGTATTCGAATATCCGCAGCCAACATTCCAAAAGCTGATGAAAGAGCACTGCATGGAA
CCCTTTTTCGTGTTTCAGGTGTTCTGCGTGGGACTTTGGTGCTTGGATGAATATTGGTACTACAGCATATTCACCCTGTTTATGCTGTTTATGTTTGAAT
CAACAATGGCCAAAAGCAGGCTCAAGACTCTAAGTGAGCTTAGGCGTGTGAGAGTGGACAATCAAACACTTATGGTACACCGGTGTGGAAAGTGGGTGAA
GCTTTCTGGAACTGAACTACTTCCTGGGGATATTGTATCCATTGGTCGATCTGTTGGACAGAATGGAGAAGACAAGACAGTGCCTGCAGACATGCTTCTC
TTAGCAGGGAGCGCCATTGTGAATGAAGCCATTCTCACAGGCGAGTCAACTCCCCAGTGGAAGGTTTCAATTGCTGCTAGGGGAAAAGATGAGAATCTAT
CAGCTAGAAGGGATAAAACCCATGTCTTATTTGGTGGAACTAAGATATTGCAGCACACTCCAGATAAGACTTTCACCCTGCGTACCCCTGATGGTGGTTG
CCTGGCTGTTGTTCTAAGAACTGGGTTCGAAACAAGTCAAGGAAAGTTGATGAGGACTATTTTGTTCTCTACAGAAAGGGTAACTGCTAACAGCTGGGAA
AGTGGTCTTTTCATTCTGTTCTTGGTAGTGTTTGCTGTCATCGCAGCAGGTTATGTGCTGAAGAAGGGACTGGAGGACCCCACAAGGAGCAAGTACAAGC
TTTTCCTCAGTTGTTCATTCATCATAACATCCGTGATCCCACCTGAGTTGCCAATGGAACTATCAATAGCAGTTAATACATCTCTAATTGCCTTGGCACG
TCGTGGAATATTTTGCACGGAGCCTTTCCGTATTCCATTTGCCGGGAAGGTTGATATTTGTTGTTTTGACAAGACTGGAACACTCACTTCAGATGATATG
GAGTTCCGTGGGGTTGTTGGATTGGATGATACTACTGATTTAGAAAATGACATGACTAAAGTGCCTTCCCGTACAGTGGAAATTTTGGCAGCTTGTCATG
CATTGGTTTTTGTGGACAATAAGCTGGTTGGCGATCCTCTTGAGAAGGCTGCACTTAAAGGAATAGACTGGAGCTATAAATCTGACGAGAAGGCTATGCC
CAAGAGGGGAAGCGGGAATGCTGTTCAGATTGTTCAACGACACCATTTTGCATCTTATTTAAAGAGAATGTCAGTTGTCGTTCGTGTAGAGGAGACATTT
TTGGCTTTTGTAAAGGGTGCACCCGAGACTATTCAAGATAGGCTTACTAACTTGCCACCATCCTATGTCGATACATACAAGAAATACACACGTCAAGGCT
CCCGCGTTTTGGCCCTCGCTTTCAAGTCCCTTCCAGACATGACAGTCAGTGAAGCACGAAATATGGATAGGGATGAAGTGGAGAACGGTCTTACTTTTGC
TGGTTTTGCTGTGTTTAACTGCCCAATCAGAGCAGATTCAGCTACTATATTGTTAGAGTTGAAGAACTCATCGCATGACTTGGCAATGATTACTGGCGAC
CAAGCCTTGACAGCTTGCCATGTTGCTAACCAAGTGCACATAATCTCGAGACCAACATTGATACTTGTCAAATCGAGAAATGAAAGATACGAGTGGATTT
CACCAGATGAAACAGAAACCGTTCCGTACAGTGACAAAGAGGTTGAGGCTTTATCAGAAAGTCATGACCTGTGCATTGGAGGTGACTGTATGGAAATGCT
GCAACAGAGTTCAGCTGTTCTTCGAGTCATTCCTTTTGTCAAGGTCTTTGCAAGGGTCGCCCCTGAACAAAAAGAATTGATCATGACTACATTCAAGACA
GTGGGGAGAGTTACATTGATGTGCGGTGATGGCACAAATGATGTTGGAGCTCTGAAGCAGGCCCATGTCGGTGTTGCACTGTTGAATGCAGTACCTCCAT
CTTCAGAAACAGCTAAAGATAAAGATGGGAAAGAGGGGAAAGAGGGAAATCAAAAGGCTGTCAAACCCAAGAAAACAAATGCTACTGAGACAGGATCTTC
AAAGAGCAAAGCTCTGACTAAGGCAGAGTCAAGCGGTAGGTCTTCAGCTGGTAACCGTCATCTAACAGCTGTAGAAATGCAGCAACAGAAACTGAAGAAA
CTGATGGACGAGATGAACGAAGAGGGCGATGGTCGCTCTGCTCCCGTAGTCAAACTCGGTGATGCCTCAATGGCGTCACCTTTCACGGCTAAGCATGCGT
CAGTTGCCCCAACTACAGACATTATTCGTCAAGGTCGCAGCACTCTGGTGACTACCCTTCAGATGTTTAAAATCCTTGGTCTGAATTGCCTTGCCACTGC
CTATGTACTTAGTGTCATGTACTTGGATGGAGTAAAGCTCGGCGACGTACAGGCAACAATCAGCGGCATATTTACGGCCGCATTTTTCCTCTTCATCTCT
CACGCAAGGCCTCTCCCGACTTTATCCTCTGAGCGGCCGCACCCTAACATCTTCTGCTTCTACGTCTTCCTTTCTCTGATGGGACAGTTTGCAATGCACT
TGGTCTTCCTCATTTCTTCGGTGAAAGAAGCGGAAAAGCACATGCCGGAGGAGTGCATCGAGCCGGACTCTGATTTCCACCCGAATCTGGTGAACACGGT
TTCGTACATGGTGAGCATGATGCTTCAGGTGGCCACTTTTGCTGTGAACTACATGGGACACCCTTTCAACCAGAGTATCTCGGAGAACAAACCCTTCTTG
TATGCAATTCTTGGTGCTGGTGGTTTTTTTACCATCATAACCTCGGATCTGTTCAGGGATCTGAACGACTGGTTAAAGTTGGTGCCTTTGCCGCTGGGTT
TGAGGAACAAGCTTCTGATCTGGGCTCTGCTTACCTACTTCGCCTGCTATTCGTGGGATAGATTGTTGAGGTGGGCATTCCCTGGCAAGGTTCCAGCCTG
GCAGAAAAAGCAACGTGCTGCTGCTGCGAATTTGGAGAAGAAAAAACGGGTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029800 pacid=23159770 polypeptide=Lus10029800 locus=Lus10029800.g ID=Lus10029800.BGIv1.0 annot-version=v1.0
MLSFHVGGKVVDKVDLLRKKHWTWRLDVWPFAIIYALWLTTVIPSLDIGDAFIVFGGLVAFHILVLLFTAWSVDFRCFVQFSKANDIHQADACKITPAKF
SGSKEVVPLYIRKQAGGSSSAEDQDEIFFDFRKQCFIYSDEKKTFCKLPYPTKEKFGYYLKCTGHGSEAKVLTASEKWGRNVFEYPQPTFQKLMKEHCME
PFFVFQVFCVGLWCLDEYWYYSIFTLFMLFMFESTMAKSRLKTLSELRRVRVDNQTLMVHRCGKWVKLSGTELLPGDIVSIGRSVGQNGEDKTVPADMLL
LAGSAIVNEAILTGESTPQWKVSIAARGKDENLSARRDKTHVLFGGTKILQHTPDKTFTLRTPDGGCLAVVLRTGFETSQGKLMRTILFSTERVTANSWE
SGLFILFLVVFAVIAAGYVLKKGLEDPTRSKYKLFLSCSFIITSVIPPELPMELSIAVNTSLIALARRGIFCTEPFRIPFAGKVDICCFDKTGTLTSDDM
EFRGVVGLDDTTDLENDMTKVPSRTVEILAACHALVFVDNKLVGDPLEKAALKGIDWSYKSDEKAMPKRGSGNAVQIVQRHHFASYLKRMSVVVRVEETF
LAFVKGAPETIQDRLTNLPPSYVDTYKKYTRQGSRVLALAFKSLPDMTVSEARNMDRDEVENGLTFAGFAVFNCPIRADSATILLELKNSSHDLAMITGD
QALTACHVANQVHIISRPTLILVKSRNERYEWISPDETETVPYSDKEVEALSESHDLCIGGDCMEMLQQSSAVLRVIPFVKVFARVAPEQKELIMTTFKT
VGRVTLMCGDGTNDVGALKQAHVGVALLNAVPPSSETAKDKDGKEGKEGNQKAVKPKKTNATETGSSKSKALTKAESSGRSSAGNRHLTAVEMQQQKLKK
LMDEMNEEGDGRSAPVVKLGDASMASPFTAKHASVAPTTDIIRQGRSTLVTTLQMFKILGLNCLATAYVLSVMYLDGVKLGDVQATISGIFTAAFFLFIS
HARPLPTLSSERPHPNIFCFYVFLSLMGQFAMHLVFLISSVKEAEKHMPEECIEPDSDFHPNLVNTVSYMVSMMLQVATFAVNYMGHPFNQSISENKPFL
YAILGAGGFFTIITSDLFRDLNDWLKLVPLPLGLRNKLLIWALLTYFACYSWDRLLRWAFPGKVPAWQKKQRAAAANLEKKKRV
|
DESeq2's median of ratios [FLAX]
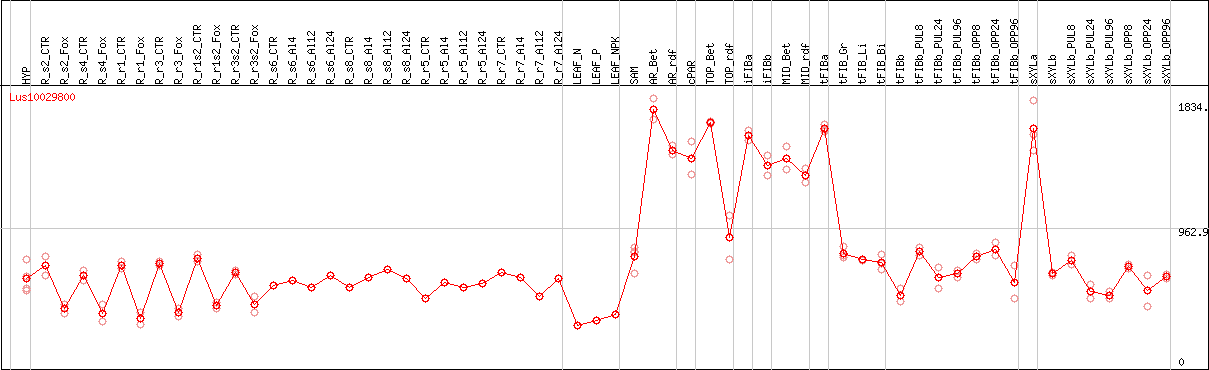
Coexpressed genes
Lus10029800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
