Lus10029842 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029842 pacid=23159642 polypeptide=Lus10029842 locus=Lus10029842.g ID=Lus10029842.BGIv1.0 annot-version=v1.0
ATGGGTTTCATTTCTCGGAAGATTATCCCTTCTTGTGGCACGATGTGCGTGTGCTGCCCTGCAATGCGCTCCAGATCTCGCCAACCTGTTAAAAGATACA
AGAAATTGCTTGCTGATGTTTTCCCCAAGTCTCCTGATGCTCCTCCAAATGACAGAAAGATCACAAAGTTATGTGAATATGCAGCCAAAAACCCAATTCG
GATTCCCAAGATTGCCCAGTATCTTGAAGAAAGATGCTGCAAAGAACTGCGGAGCGGCCATACCAAGTTTATCAATGTTGCTGTGGAGGCTTACAATAAG
TTGCTCTGCATTTGTAAGAACCAGATGGTATATTTTGCCACTAGCTTGTTGAATGTGGTCACAGAACTGCTAGACAAACTTAAACAAGAGGCTTTGGTGA
TTCTTGGATGCCAGACTTTGACGAAGTTCATCTATAGCCAGGTTGATGGAACTTATGCTCATAGCATCGAAAAATATGCTCCTAGAATATGTACACTTGC
ACGCAAGAGTGGGGAGGAGGGTAATAGTAGCCGTTTGAGGGCATCAAGCCTGCAGTGCCTCTCAGCCATGGTATGGTTCATGTCAGAGTTCTCGTATATT
TTCCCTGCATTGGATGAGATTGTGCATTCCACCTTGGATAACTACGAGCCTGAATCTCCTATTGAAAATGATGGCGAGAGAGGGGAAGCTCACAATAATT
GGCTGAATGAAGTTGTTAGAAGTGAGGGAAGAAGTACAGCTGTAGGAAATGACCCTAGCTCTATGGCCATTATCAGAGCACGGCCAGAAAAGAAAGATCC
TTCTCTTTTGACTGGGGACGAAATTGAGCAACCAAGAGTGTGGGCACAAATTTGTATTCAAGGAATGGTGGATCTGGCAAAGGATAGTTCCACAATGCGA
TGCGTATTGGATCCTATGTTTGTCCACTTTGACACTAGACATCTCTGGGCTCCTCAACAAGGACTTGCTCTGATAGTTTTGTCTGCCATGTCATATTTTC
TGGAGAATTCAGGACATCAACAGTCAGTGTTATCTGCTGTTATCCGCCATTTAGATCACAAAAATGTTGCACACGATCCACTGCTCAAGTCTTATGTCAT
ACAAGCTGCAACTAGTTTAGCTAGGCAACTTCGATCAGTTACAGTTCTGGCTGAAGTTGGATTTGTCAGTGACTTGTGTAGGCATTTGAGAAAAAGCCTT
CAAGCCTCAGCTGAACCTTTAGGAGACCAGGAATTGAATTTGAATGTCTTGCTCCAAAACTCTATTGAAGATTGCTTACTTGAGATTGCCAAGGGGCTTG
GTGATGCCAAACCTCTACTTGACCACATGGCAATGTCACTGGAGAAGCTACCATCCTGTGGAATTGTTGCTCGGGCAACCATTAGATCACTGATGATTCT
TGCGCATGTGATCTCGTCGTCGTCAGTTTCTTCAACCAAGCAGACTTTCCCCGAATCGCTTCTTGTTCAACTTCTAAAAGCAATGCTGCATGCAAATGTT
GAGGTGCGGGTTGAAGCACACCAGATTTTCACAGTTCTTCTCATTCCAAATGCTAGTTGTCTTGGACGGGAAGCTTCTGTTCGAACTAGTTACATCTGTG
AACCAAGAAGATGGCGTTCTGGTACACCATCTACATTTGCTTCAATTTCTGCACTGCTCGAAAAGCTCCGGAAAGGGAAGGATGGGTCTGAAGTGGAAAT
GCATGGAAATGGAGTTCAAGATGACTGCAAAGACAGAGACAACTCCGAAGAAGATTGGAAATATGGACGGGCTAGAAAGAACTCCCCAAACTTTTACAAG
ATTAGTTCAATTATTGAGAGAACTACGGGGACACCAAGCTTGGCTGAGGCTGCTGAACCTTGTGCTTTGGAGCTAAATGAGGATCAGATATCACAGTTGC
TTTCTGCCTTCTGGATGCAAGCCAGCCTTCCAGACAACACTCCTCCCAATCTGGAGGCGATTGCTCATTCCTTTACACTAACACTTATTTCTTCACAACT
GAAGGAGACAAATGCAAACCTTGTGGTCCAGTTCTTTCAGCTTGCTCTCTCTGTTAGACAATTGTCCTTGGACTCTCATAATGGGATGTTACAGCCAGCA
TGCAGAAGATCAGCTTTTGTTTTAGCAATGGGCATGTTAATGTTCACTGCCAAGACATATCAAGTTCCCGATTTGAATGACTTGCTCAAGCCGTTGCTCT
CGGACAACGTTGATCCTTATTTGGGAATTAGTGATGATCTTCAAATATACGTTAAGCCCCAAGCCGATTTGAAAGAATTCGGATCATTCACAGATAACTC
ATTGGCTCTGTCAGTGCTAACTGAGTTGCAGAGGAAAATATGTGAATCTGATACAGCTGTAATGAACAGCTTGGTTCTAAACCTATCAACAACCACCGAG
ATGGACCCGGATGCAGTGGCCATGCAATTGTTGGAACCCTTCACACCTGATGACATGTTAATGTTTGGTTCACGATCAATGGGTGACCACAACCGGAAGC
TTTCTCATTCAAAGGAATCACTTTCCTTTGACGAGGATCTTCCATCTAATGCAGTGGCCGAGGATGATGCCACTAGCGATGCATCAGTCTCGGACTTGTC
ACGTTTCATCCCCAAAATTCCTTCATCTCCTTCTGTCTCGCACGTTATTAGCATCGGGCAGCTACTAGAATCCGCTCTTGAGGTAGCTGGTCAAGTGGCA
GGAACATCTGTTTCCATGTCACCCCTCTCGTACGACTCGATGACGAAACAGTGTGAAGCTCTGGGGACAGTGACGAGAAAGAAGCTCTCGAATTGGTTGG
CTCATGAGAATCAGCAATCTAAGGGAGCAGCTGACAAGTACTTCTTGCCACCTGTTGTGGAAAGTGGAATTCCTGCTGTGGCACCATGGCAGATGGGATT
GGTGGAGGGGGAGATGATGAAGCAGGAGGGAATGGAGCAGTATTTGCCAATGAAGCTACCACCTGCAAGTCCTTATGATAATTTCCTGAAAGCGGCTGGA
TGCTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029842 pacid=23159642 polypeptide=Lus10029842 locus=Lus10029842.g ID=Lus10029842.BGIv1.0 annot-version=v1.0
MGFISRKIIPSCGTMCVCCPAMRSRSRQPVKRYKKLLADVFPKSPDAPPNDRKITKLCEYAAKNPIRIPKIAQYLEERCCKELRSGHTKFINVAVEAYNK
LLCICKNQMVYFATSLLNVVTELLDKLKQEALVILGCQTLTKFIYSQVDGTYAHSIEKYAPRICTLARKSGEEGNSSRLRASSLQCLSAMVWFMSEFSYI
FPALDEIVHSTLDNYEPESPIENDGERGEAHNNWLNEVVRSEGRSTAVGNDPSSMAIIRARPEKKDPSLLTGDEIEQPRVWAQICIQGMVDLAKDSSTMR
CVLDPMFVHFDTRHLWAPQQGLALIVLSAMSYFLENSGHQQSVLSAVIRHLDHKNVAHDPLLKSYVIQAATSLARQLRSVTVLAEVGFVSDLCRHLRKSL
QASAEPLGDQELNLNVLLQNSIEDCLLEIAKGLGDAKPLLDHMAMSLEKLPSCGIVARATIRSLMILAHVISSSSVSSTKQTFPESLLVQLLKAMLHANV
EVRVEAHQIFTVLLIPNASCLGREASVRTSYICEPRRWRSGTPSTFASISALLEKLRKGKDGSEVEMHGNGVQDDCKDRDNSEEDWKYGRARKNSPNFYK
ISSIIERTTGTPSLAEAAEPCALELNEDQISQLLSAFWMQASLPDNTPPNLEAIAHSFTLTLISSQLKETNANLVVQFFQLALSVRQLSLDSHNGMLQPA
CRRSAFVLAMGMLMFTAKTYQVPDLNDLLKPLLSDNVDPYLGISDDLQIYVKPQADLKEFGSFTDNSLALSVLTELQRKICESDTAVMNSLVLNLSTTTE
MDPDAVAMQLLEPFTPDDMLMFGSRSMGDHNRKLSHSKESLSFDEDLPSNAVAEDDATSDASVSDLSRFIPKIPSSPSVSHVISIGQLLESALEVAGQVA
GTSVSMSPLSYDSMTKQCEALGTVTRKKLSNWLAHENQQSKGAADKYFLPPVVESGIPAVAPWQMGLVEGEMMKQEGMEQYLPMKLPPASPYDNFLKAAG
C
|
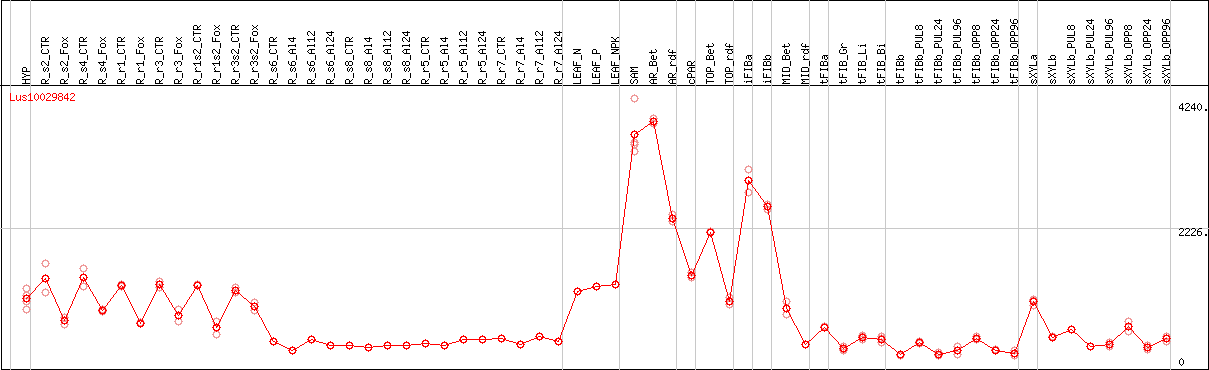
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10029842 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
