Lus10029849 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029849 pacid=23159818 polypeptide=Lus10029849 locus=Lus10029849.g ID=Lus10029849.BGIv1.0 annot-version=v1.0
ATGGAAACAGAAACAATCGCCCCCGCGACGTTGGTAGATCAACAGCAGAAGAAGAGAGTGATATTCCTTGTGGCGGTGGACGAGAGCGAGGAAAGCCTGC
ACGCTCTCTCTTGGTTCCTCAACAATACTCTGCCCTCAGCTTCCTCCTCCGACCACCCACCAACTCTCGTCCTTCTATACGTCAAGCTCACACCTTCTCC
GCCTCTCTTCTTCACCCCACACGGGTATATGTTTTCACCGGAAGTAATGGCGACGGTGAACCAGTACAAGAACGAGGTTGCCGAATGTGTAATGGAGAAG
GCCAAAAGGGTTTGCAAGGAAGCAGTTAATGACAAGGTTAAGGTGGAGACTGTCATAGAAAGAGGGGATGCAAGGGATGTTATATGCCAGGTTGCTGATA
AGCTCTCCGTTGATCTTCTTGTCATGGGCAGCCATGGCTACGGCCCAATCAAGAGGGCGTTTCTGGGGAGCGTAAGCAACCATTGTGCTCAGAATGTGAA
GTGTCCAGTTCTTATTATCAAGAGACCCAAGATCCAATCCAACTCTCTCGGTGTTGTCGGCACGATTGTTTTGATCAAGTCATGCCTTCTTACCAACACG
GGCGCAGTCAACGCTAACTCGAAGATGACCGGAGAGAGCTACACGTTCTACTCCGCAATCGAAAACTCCGGCAGCCATATCTTCCCCGGCAAGGAAACCA
CCGCCAATTTAGCCACTCTGCTCGCTTCATCCTTCGTCAAAAGCAACGAGGATTCTCTTCCTTCCTTCGCCAACGTTATGAAAAACGTATCGCCGAAACT
GTTCCTAGCAATGTCGATAATTCCTAACCTATCCCAAGCGACCTTCGCAACCAGAGACTGCTTGTTCTTCTCCTACTTTAGAAAAGACTATGGAACTTTC
GCTGTATACTCCAACACTTCGTTCTCTCCCGTTTGGTACACCCAGCCGGTGAACGAAGATACCGGAGAGCTATACGGTGTTGCTTTGGAATCCAATTCCA
CGCTCTCCTCGAATTCCAGCTGGTTCCTGAATGGTCTGGAAACGAACGATGTCGTTTCCTCCGTGAGGACCAGCTGGAACAGAGATCGTTTGTTCCTAAG
CACGGTTTCCGTGCACGGAAAGGGTATGGTTTCTCTCGGGTACCCGATCCAAGTCGTCGCCGATCAGTTCAGCGCGCTGAATAATCAGCGGCTGCATTTG
GGGATGGCTACGGTGGAGGAGAGCCAAGTGATTCTTCCGCCAACGGTTGTCGAAATCGAGAAGTACTACACTGGCGGGGTATTGACGGTTAGGGGGATTG
GAGGAGGTGGAAGGAGTGTTAATTTGACATGTGACTCCGCCGGCTCCGCTAGGAGGTCCCCTCCGGAGACGGTCGACGGAACAGGCTGCAGGTTTTACTG
CTCCACCCTCAAAATTGGTGGAGTAGATTTGGTTTACATAGTCGAATTCCATGGCATTACAGTGATGAACTACCAGGAATCGAATGTATTAGATGTGGTC
TTGTACGGTTTACTGGTGCTCCCCTTTACCGCATTGGTCTGGAAGAAGAAAGAATCAACACACAGGGAGCAACATCTATGCGCTTCACTGATCAAGCAGA
CGGAAGCAACGCAGCAAGCCGAGCGCAAGAGCATGAACAAGAACAGGATTTTCGCGGGCATCAGCCACGACGTTCGTAACGCAATCGCTCAACTCGTCGC
ATTAACCGAGTCCTGTAAAGATGATGATGCCAGCAGTCTGCAGAAAATCGAGCATTGCGCTTACGATGCTCTAGGCATTTTGGACTCGGTGCTCGACATC
AACAGGATCGAATCAGGGAAACTACTGCTGAAGAATGAAGACTTCAACTTAGCCAAGCTAGTGGAAGAAGTGGTGGACAGATTCTATGACTTGGGGAACA
AGAAAGGTGTGGACGTGGTGTTCGATCCGTCTCATGACTTTATGATGAAGTCGAGGCTTGTACGGGGCGATCCAACGAGGTTGACACAGATACTGTCGAA
CTTGCTAAGCAATGCCATCAAGTTTACATCGGAAGGGCATGTAACGGTTCGTGCTATGGTCAAGAAGAGGAACTTCGGCAGGGATATCATTGCTTATAAC
AGGAGGTTTACTTGCTGGAAGGCATTTTGGAGATTGTGTGAGAAGAGCAATGGGGATAGTGACAACGATGTTATTAATACCCTTCTTGAAGGGGTTGAGG
AGAATCCGGATGAAGTGGAGTTCGAGTTCGAGGTTGATGATACGGGGAAAGGTATCCCGAAAGACGAGCAGAAGTCAGTGTTCGAAGACTACATCCAAGC
TAGAAACAATGTTGCTTGTTCCGACTCGGGAGGAGGGTACGGCTTGGGACTTGGCAATGTTCAGTCGTTGGTGCGGTTGATGAAGGGAGAGCTACAGATT
GTTGAGAAAGAACAAGAGGAGAAAGGGACTTGCTTTAGATTCAATGTTTTTCTCTCCATGTCCAATACTACTGCAGTGGAAGCAGAAGAAGGCCAGCAGC
AACAGAAACCCACTTTCCGACCATCTTTCAAATCGGAAGATTCTCAAGTAATGATCTTGATCGAAGGGGAAGAGAGGAGGAAGGTGTTGAAGAAGTTCAT
CGAGAGCATGAACGTAAAAGCGAGAATCATCGACCAGTCAAGAAGCTTGAAGCAAGAGCTAGAGAGGATCAAGACAAAGATCCAAAATCCTCCCTCAACT
CTCTCCCAACTACTGATCAGCAGCAGCCACTCTGTTTGTTCTATAAGTAGTACTGATCATGCACCGAAAGTCGTTTTGTTGATCATAGATGTCAATTACT
ACAGCGAAATTGTTCGTGATTGCTTGCACGAGTACTTCAAGACAGGGACCGACAATGGCTCCTCGTGGAAGGTCGTTTGGTTACACGAGATACAGCAGGC
TGGAGTAGACCATGGATTGCGAGAAACCAAGTATGATTGCTTGGTGCACAAGCCATTCCATGGTTCCCGTGTCATGCAAGTTTTTTCACTGCTTCCCGGG
AAGGGCACCACGTTACCAAGCACTGCTCCCGATGCTACTACTACTCCGGTGGTGAGATTTCGAAGGGAATTCTGTATTAGCTCCAACTGTTATACGATTG
ACCTGCGTGACACTGACAATGACTCGGTTTCAAGCCATCGGGAGGAGGTTGTGTCCGTGAATGGTCATGATATGTTGCTGCCGTTGAAGGGGAAACGGGT
TTTGATTGTGGACGATAATGAGCTGATGAGGGAAGTGAATGCTCGGAGGATGAGTAAGTTAGGAGCCGAAAGTGTCGAAGTCTGTGGAAACGGGAAGGAG
GCATTGGATAGAGTGTCAACCGCGTTGAGAGGGTCTCAGAGCGATCTTCTCCCCTATGACTGCATTTTCATGGATGGCGAGATGCCAGAAATGGATGGGT
ACGAGGCGACGAGAATGATAAGGGAGGAGGAAAGGAGGAATGGAGAAGGGAAGATGATGGGAATAGTAGCCGTAACAGGGCACTCAGATGCAGACGAAAT
GACGAAGAGCTTGGAAGCTGGGATGGACTTCCATTTGACGAAGCCATTAAAAGAGAATGAGGTGGTTGATGCTGTAAGAAAGCTTGATGCTAAGCATGCT
ATTTGTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029849 pacid=23159818 polypeptide=Lus10029849 locus=Lus10029849.g ID=Lus10029849.BGIv1.0 annot-version=v1.0
METETIAPATLVDQQQKKRVIFLVAVDESEESLHALSWFLNNTLPSASSSDHPPTLVLLYVKLTPSPPLFFTPHGYMFSPEVMATVNQYKNEVAECVMEK
AKRVCKEAVNDKVKVETVIERGDARDVICQVADKLSVDLLVMGSHGYGPIKRAFLGSVSNHCAQNVKCPVLIIKRPKIQSNSLGVVGTIVLIKSCLLTNT
GAVNANSKMTGESYTFYSAIENSGSHIFPGKETTANLATLLASSFVKSNEDSLPSFANVMKNVSPKLFLAMSIIPNLSQATFATRDCLFFSYFRKDYGTF
AVYSNTSFSPVWYTQPVNEDTGELYGVALESNSTLSSNSSWFLNGLETNDVVSSVRTSWNRDRLFLSTVSVHGKGMVSLGYPIQVVADQFSALNNQRLHL
GMATVEESQVILPPTVVEIEKYYTGGVLTVRGIGGGGRSVNLTCDSAGSARRSPPETVDGTGCRFYCSTLKIGGVDLVYIVEFHGITVMNYQESNVLDVV
LYGLLVLPFTALVWKKKESTHREQHLCASLIKQTEATQQAERKSMNKNRIFAGISHDVRNAIAQLVALTESCKDDDASSLQKIEHCAYDALGILDSVLDI
NRIESGKLLLKNEDFNLAKLVEEVVDRFYDLGNKKGVDVVFDPSHDFMMKSRLVRGDPTRLTQILSNLLSNAIKFTSEGHVTVRAMVKKRNFGRDIIAYN
RRFTCWKAFWRLCEKSNGDSDNDVINTLLEGVEENPDEVEFEFEVDDTGKGIPKDEQKSVFEDYIQARNNVACSDSGGGYGLGLGNVQSLVRLMKGELQI
VEKEQEEKGTCFRFNVFLSMSNTTAVEAEEGQQQQKPTFRPSFKSEDSQVMILIEGEERRKVLKKFIESMNVKARIIDQSRSLKQELERIKTKIQNPPST
LSQLLISSSHSVCSISSTDHAPKVVLLIIDVNYYSEIVRDCLHEYFKTGTDNGSSWKVVWLHEIQQAGVDHGLRETKYDCLVHKPFHGSRVMQVFSLLPG
KGTTLPSTAPDATTTPVVRFRREFCISSNCYTIDLRDTDNDSVSSHREEVVSVNGHDMLLPLKGKRVLIVDDNELMREVNARRMSKLGAESVEVCGNGKE
ALDRVSTALRGSQSDLLPYDCIFMDGEMPEMDGYEATRMIREEERRNGEGKMMGIVAVTGHSDADEMTKSLEAGMDFHLTKPLKENEVVDAVRKLDAKHA
IC
|
DESeq2's median of ratios [FLAX]
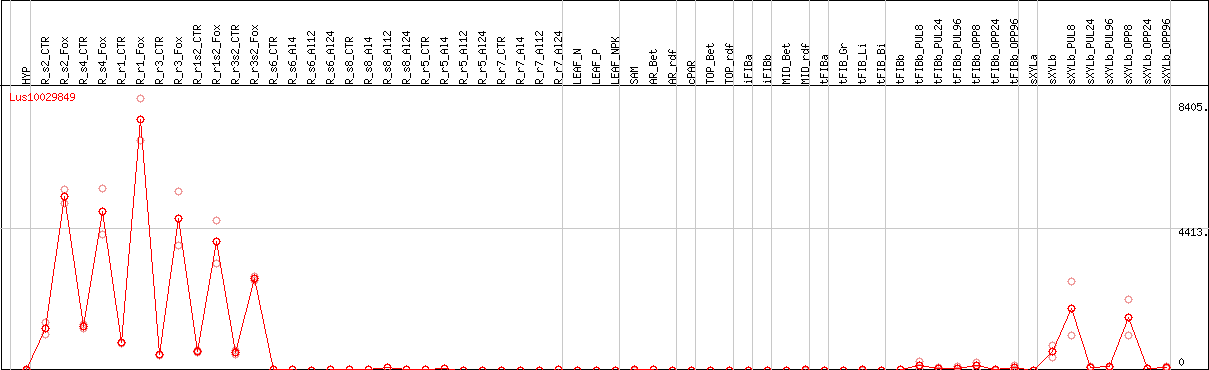
Coexpressed genes
Lus10029849 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
