Lus10029852 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10029852 pacid=23159878 polypeptide=Lus10029852 locus=Lus10029852.g ID=Lus10029852.BGIv1.0 annot-version=v1.0
ATGTTTCCACGAGGGAAAGCTGCAGAAGAAGAGAAAGGACATGGCCGTGTCGATACAGAGGAATTCATCGATGATTTCGCTTCAAAAATAGAACCGACGA
TTCGATCAGTGATCCGTGATGAGGTTAACACAATCATCCGACAGGCTTCTGGACCAACTCTTGAGCCATTGGCTCTAGGGGTTCCTGCGGAGGGATTGAT
TCTGCAGTTTATGGACAAGATTCCATCAACCATCTTCACAGGAAGCAAGATTGTAACTGAAGATGGCAGCCCAGTAAGGATCCAGCTCCTCAATGCTTCA
TCCAACACAGTTGTCAAATCAGGACCTTTCTCTTCCCTAAAAGTTGAAATCACTGTTCTTGATGGCGACTTTGTTTCTGAAGACTGGTCAGACAGAGATT
TCAGTCGCAATGTGATTAGACAAAGAGATGGAAAGAGGCCTTTGGTGACTGGGGAGCTTATCATTGTGCTGTCAGATGGAGTTGGACAGCTCGGAGATTT
GGCTTTCACTGATAATTCCAGCTGGGTTAGATGCCGGCGGTTTAGGCTGGGTGCTAAACCAGTACTGAAAAGAACTTCTGGAGAAGTTATTAGGATTAGA
GAAGGTAGAAGTGATCCTTTTGCTGTCAAGGACCACCGTGGTGAAGGGTACCAAAAGCACTACCCTCCAAGGTTGGGAGATGAAGTTTGGAGGTTAGAGA
AGGTAGCAAAAGGTGGAAAACTGCATAATCGCCTTTCTTCTTTCGGAATCAATACTGTGAAAGAGCTTCTTCAGGCCAATGCCGTCGACCAACCTGCACT
AAGAAATAAAGTTGTTGGAGGTGTAATCTCGAACAGAGTATGGGACATGATCATGGTGCATGCCAATGCATGTGAGGTAGAAGAAAACAAGTTCTATGCC
TATGGTAAACATGGAGAAGGCATCAGACTGCTGTTCGATCCTCTATACAAGCTTGTAGGCGCCAGTTTTGATGATGAGAAAAGCTACATTCCTTTGGATA
AACTAACAAATGCTCAGAAGATCTTGGTGGAGCTGTTTAGAGGGCAGGAAGCTATCCATGACCGGATGGAAATGGTCCCCGTTGATTCTTCTTCAGTTGC
TTTCACCTCATTCTTGCCTTTGGAGGAAGTTGATCATCAGAGGAAATACGTGCCGTCATTTGTGTCAAAGACATATCAGCTGGTGGACGATCCGTCCACT
GACCACATTGTGTCTTGGGTGGGAGACGGATCAACCTTCCTAGTACGGGTCCCGGCTGAGTTCGCACGTGACGTTCTTCCTAAATACTTCAAGCACAATA
ACTTCTCCTCCTTTGTCCGCCAACTCAACACCTTTGGTTTTAGAAAGTTTGCACCGGAGAGATGGGAATTCGTGAACGAGTTCTTCAAGAAAGGGGAAGA
ACATTTGCTCTGCCAAATCAAGCGCAGGAAAACCCAGACATTTCTGAAGAAAGATGAATCCAAACCAAGTCCGGATTTGAAAGATAACAACTCGATAAGC
TCACAATCGTCATGCCTTTACAATGACAGCGGAATTACCAACAGGAAGTGGAAGCAGGAAACTTCCGCGGTCACTTATTCTGGTTCCTGTGCATCAGAAG
TGACGACATGGGGACAGGAGATTCCTCCTTTTCTCACCCTATGTACAGAAGAAGAAGTGGCTTCCCATGTTCCCAATTGGCTATCGTTGCCCAAACCTGA
ATCGAACCTAAAAGCAAAATCTGCTTGGCGCAAGCTTCGGACAGCTTTGAAATTCACTTCTATTCTCACAAGAAGGACACGTCTAATTGAAGCTGCTAAA
GTTGATTTCAGATTGCTGAGCTTCCAAAGAATGCTAAAAGAACAGGGAAATGTGAAGTTAAGTAGTATATCAGATGAGCAAGCGTTTTCCTTCATCAAGG
TGAAGGGAAACATGAAGCCCAGCTGGCTAGTAGAAGCTGCTGCAGCAAGGATTCAGAACAAGTTCCGGAGTTGGAAAGCCAGGATGGAGTTCTTAGCTAT
TCTGCATAAAATTGTTAAAATTCAGGCTCATGTAAGAGGACACCACGTCAGGAAAAACATCAAGGTGATATTGTCATCTGTAGGCATTACGAAGAAGGTG
GTACTTCGAAGGAGAAGGAACAGAAGTATGTTGCGCGGGTTTAGACCTGGGTTGGTCGATGAAAGTCACAGAGGCAGCATCCAGCAGCATTTATCAGCAG
GAGAGGATGAGCAAGCGCTTCCCTTAATCAAGGTGAAGGGAAACAAGAAGCTCGCCGAGCCAGTAGAAGCAGCTGCAGTAAGGATTCAGCGCATGTTCCG
GAGTTGGAAAGCCAGGAAGGAGTTCTTAGTTATTCTGCAAAAAGTTGTTAAAATTCAGGCTCATGTAAGAGGACACCATGCCAGGAAAAACTTCATGGTG
ATGTCATCGTCTCTAAGCATCGTGGAGAAGGTGGTACTTCGATGGAAAAGGAACAGAAGCACGTTGCATGGGTTTAGACCCGAGTTGGTCGATGAAAATC
ACCACAGCAGCATCCAGCCTGATTTATCGGCCAGAGAGGACGATTATGGTTTCTTTAAAGAAGAAAGGAAGCAAACAGAAGAAAGATCTCAGATTGCTCT
TGTGAAGGCGACGTCTGTGATTCAGCTTTCGGAGCCTAAAGTGGAGATAACCCAAATGAAAGATCTAAAATTCGACATGGCTAAAGCATTGCAGCTTCAA
AGTTTCACTTCGAAACAAAATCTTCATAGCCGCAACTCCATGTTTACACAAGAGATTTACATTTCAAAGTACCCCTCGTTCCAAATTTCTGCCCACAACT
TTGTGTCCTTAGTATTCCCATACTGCAACTTTCCTCAGCTATGGAAGGGAGGGGAACAGCTGTTAGTAAACCTGAAGTCTATGGATCTGAACCACTCGAA
TCTGTTTGAGATCCCAAACTTATCAAAGGCTCAGAAATTGGAAAGTGTGAATCTTGAAGGCTGTGGAAACTTGGTTGGGGTGCCATCCATTCAGTATCTT
ACAAATCTTAAGAATCTAAACCTAAAGGGCTGTCATGGTCTGAAGGGGCTACCAAGTTTGATCAGATTGAAGTTCCTCAACAGTCTGGATCTCTCCGACT
GCTCAAGCCTCACCAGGCTTCCTCCATCGCTTGGGTGTCTGCCGAATCTTTGCGAATTGAACCTCAGGAACTGCACAAAACTGGATTACCTCCCGCGTAC
TGTTCTTCACTTAATGAGATCCCTTGAAACTCTGAATGTTTCTGGCTGCTACAGCTTGTGGAAACCTATCAATGGAGACCTCGTCAGCAATTCTACTGAA
GGAACTTGCCTTGAATCGCAATCGGTCAAGCCTGTGGAATTTGATGCTGCTGGAGATTCACCATCGCAAGATCTGTCACCCGAATTTAATGAAATGCACG
GAAAGGAGGCTTTAGAATGGGAGCTGGATGATTATACTTCTACGGCCACTCCATTGATAGATTATCTACCTTCAAAACTCAGGACTAGGCTGTTGCTTCA
TCCTGCAATGGATTTTGAAGATAAAAATGATCATGGAGATACAACGAGGACTGTATCTGTTAGTTGCCAGGTGGAAGATTGTGGAGTTGATCTTAGCCAG
GCAGAGACTTATCACCAACGTCATAGAATCTGCGAGGTTCATTATCAGGCTAGTAATGCATTAGTCAGAGATGTCCTGCAACGGTTTTGCCTGCAATGTT
GCAGGTTTCATATCCTTGAAGATTTCGATGATGGGGAACGAAACTGCAGAAGCCGGTTAGCAGGCTATCATCAAAAGCAGAGGAGGCAGAGTATTGCTAC
AGAGGATGAAAAACCTTCAACTCTCGAAACGGATGAAAACGGGTCACTGTATAAGAGAACAAAGTGGGAAGACTGGGATGAAGAAGCTAATGATGCATTT
GGGAGCATTGAGAACGATGTTCTGGAAATGGTCGAGCAACTTTCGAGCTGGAATTGCGATGGAAAGGAAGTGCCGGAGGATGTGGTGGCTTGGTTGGCAG
AGCTGGAACAGAGTGTTCCAGAGACAGTGATGAGTATGAAGAGTGCCAAAGCTGAAGCAGGTGTTGTTGGGGAGAAGTTGAAGGAGATGGATGCAAGGTT
GGTAGTTGGAAGGGTAGAGCTGAGTTTGCTGGATAAGGAAATGTCAAGGATTTTGAAGGAGGAAGAGGAGATTGAGGCTCAGCTTCAGCTTTTGATGGAT
GAGAAGAATAGAAGAGATTTGTTGATGCAGCAGAAGGGGACTGAACCATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10029852 pacid=23159878 polypeptide=Lus10029852 locus=Lus10029852.g ID=Lus10029852.BGIv1.0 annot-version=v1.0
MFPRGKAAEEEKGHGRVDTEEFIDDFASKIEPTIRSVIRDEVNTIIRQASGPTLEPLALGVPAEGLILQFMDKIPSTIFTGSKIVTEDGSPVRIQLLNAS
SNTVVKSGPFSSLKVEITVLDGDFVSEDWSDRDFSRNVIRQRDGKRPLVTGELIIVLSDGVGQLGDLAFTDNSSWVRCRRFRLGAKPVLKRTSGEVIRIR
EGRSDPFAVKDHRGEGYQKHYPPRLGDEVWRLEKVAKGGKLHNRLSSFGINTVKELLQANAVDQPALRNKVVGGVISNRVWDMIMVHANACEVEENKFYA
YGKHGEGIRLLFDPLYKLVGASFDDEKSYIPLDKLTNAQKILVELFRGQEAIHDRMEMVPVDSSSVAFTSFLPLEEVDHQRKYVPSFVSKTYQLVDDPST
DHIVSWVGDGSTFLVRVPAEFARDVLPKYFKHNNFSSFVRQLNTFGFRKFAPERWEFVNEFFKKGEEHLLCQIKRRKTQTFLKKDESKPSPDLKDNNSIS
SQSSCLYNDSGITNRKWKQETSAVTYSGSCASEVTTWGQEIPPFLTLCTEEEVASHVPNWLSLPKPESNLKAKSAWRKLRTALKFTSILTRRTRLIEAAK
VDFRLLSFQRMLKEQGNVKLSSISDEQAFSFIKVKGNMKPSWLVEAAAARIQNKFRSWKARMEFLAILHKIVKIQAHVRGHHVRKNIKVILSSVGITKKV
VLRRRRNRSMLRGFRPGLVDESHRGSIQQHLSAGEDEQALPLIKVKGNKKLAEPVEAAAVRIQRMFRSWKARKEFLVILQKVVKIQAHVRGHHARKNFMV
MSSSLSIVEKVVLRWKRNRSTLHGFRPELVDENHHSSIQPDLSAREDDYGFFKEERKQTEERSQIALVKATSVIQLSEPKVEITQMKDLKFDMAKALQLQ
SFTSKQNLHSRNSMFTQEIYISKYPSFQISAHNFVSLVFPYCNFPQLWKGGEQLLVNLKSMDLNHSNLFEIPNLSKAQKLESVNLEGCGNLVGVPSIQYL
TNLKNLNLKGCHGLKGLPSLIRLKFLNSLDLSDCSSLTRLPPSLGCLPNLCELNLRNCTKLDYLPRTVLHLMRSLETLNVSGCYSLWKPINGDLVSNSTE
GTCLESQSVKPVEFDAAGDSPSQDLSPEFNEMHGKEALEWELDDYTSTATPLIDYLPSKLRTRLLLHPAMDFEDKNDHGDTTRTVSVSCQVEDCGVDLSQ
AETYHQRHRICEVHYQASNALVRDVLQRFCLQCCRFHILEDFDDGERNCRSRLAGYHQKQRRQSIATEDEKPSTLETDENGSLYKRTKWEDWDEEANDAF
GSIENDVLEMVEQLSSWNCDGKEVPEDVVAWLAELEQSVPETVMSMKSAKAEAGVVGEKLKEMDARLVVGRVELSLLDKEMSRILKEEEEIEAQLQLLMD
EKNRRDLLMQQKGTEP
|
DESeq2's median of ratios [FLAX]
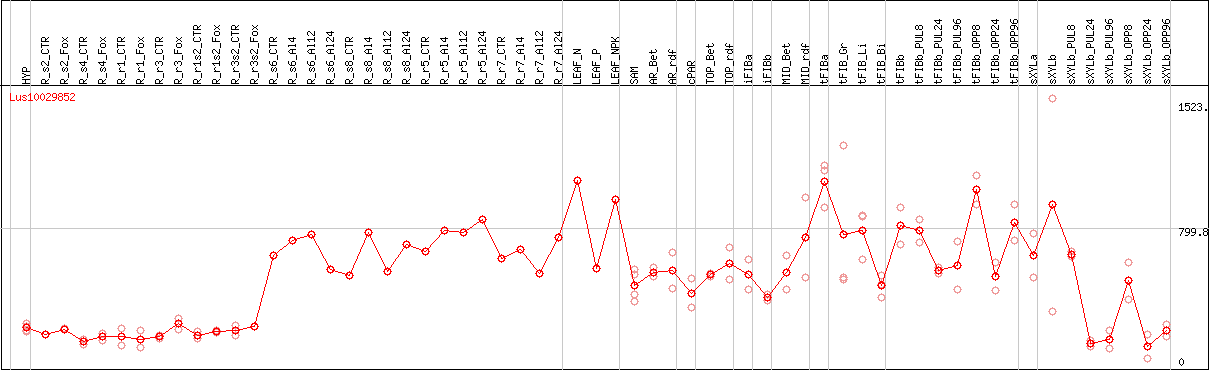
Coexpressed genes
Lus10029852 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
