External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G79580 253 / 8e-82
NAC
ANAC033, SMB, NST6
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT2G46770 219 / 9e-69
NAC
NST1, ANAC043, EMB2301
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G32770 216 / 1e-67
NAC
NST3, ANAC012, SND1
SECONDARY WALL-ASSOCIATED NAC DOMAIN 1, NAC SECONDARY WALL THICKENING PROMOTING 3, NAC domain containing protein 12 (.1)
AT4G10350 215 / 2e-67
NAC
BRN2, NST4, ANAC070
BEARSKIN 2, NAC domain containing protein 70 (.1)
AT3G61910 211 / 7e-66
NAC
NST2, ANAC066
NAC SECONDARY WALL THICKENING PROMOTING FACTOR2, NAC domain protein 66 (.1)
AT1G33280 210 / 8e-66
NAC
BRN1, ANAC015, NST5
BEARSKIN 1, NAC domain containing protein 15 (.1)
AT4G36160 201 / 1e-61
NAC
ANAC076, VND2
VASCULAR-RELATED NAC-DOMAIN 2, NAC domain containing protein 76 (.1)
AT2G18060 201 / 2e-61
NAC
ANAC037, VND1
Arabidopsis NAC domain containing protein 37, vascular related NAC-domain protein 1 (.1)
AT3G24503 202 / 2e-60
ALDH1A, REF1, ALDH2C4
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
AT5G66300 193 / 2e-59
NAC
ANAC105, VND3
VASCULAR-RELATED NAC-DOMAIN 3, Arabidopsis NAC domain containing protein 105, NAC domain containing protein 105 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009939
285 / 5e-96
AT1G79580 314 / 4e-107
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Lus10019387
264 / 2e-82
AT3G24503 548 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10033239
229 / 7e-72
AT1G32770 318 / 2e-106
SECONDARY WALL-ASSOCIATED NAC DOMAIN 1, NAC SECONDARY WALL THICKENING PROMOTING 3, NAC domain containing protein 12 (.1)
Lus10008271
221 / 1e-71
AT1G32770 277 / 8e-94
SECONDARY WALL-ASSOCIATED NAC DOMAIN 1, NAC SECONDARY WALL THICKENING PROMOTING 3, NAC domain containing protein 12 (.1)
Lus10002687
228 / 3e-71
AT2G46770 358 / 1e-121
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10030205
227 / 7e-71
AT2G46770 357 / 5e-121
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10023626
227 / 1e-69
AT3G24503 753 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10017340
222 / 2e-69
AT2G46770 347 / 7e-118
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10001664
222 / 3e-69
AT1G32770 343 / 2e-116
SECONDARY WALL-ASSOCIATED NAC DOMAIN 1, NAC SECONDARY WALL THICKENING PROMOTING 3, NAC domain containing protein 12 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G176600
278 / 4e-92
AT1G79580 344 / 8e-118
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Potri.008G080000
272 / 7e-90
AT1G79580 333 / 2e-113
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Potri.014G104800
231 / 3e-73
AT2G46770 380 / 3e-131
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.002G178700
230 / 1e-72
AT2G46770 366 / 1e-125
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.011G153300
226 / 9e-71
AT2G46770 297 / 4e-98
NAC SECONDARY WALL THICKENING PROMOTING FACTOR1, Arabidopsis NAC domain containing protein 43, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.001G448400
225 / 2e-70
AT1G32770 295 / 4e-97
SECONDARY WALL-ASSOCIATED NAC DOMAIN 1, NAC SECONDARY WALL THICKENING PROMOTING 3, NAC domain containing protein 12 (.1)
Potri.018G075000
217 / 2e-66
AT3G24503 751 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Potri.013G092400
209 / 5e-65
AT4G10350 436 / 5e-154
BEARSKIN 2, NAC domain containing protein 70 (.1)
Potri.019G066000
209 / 1e-64
AT4G10350 451 / 8e-160
BEARSKIN 2, NAC domain containing protein 70 (.1)
Potri.007G014400
209 / 1e-64
AT2G18060 427 / 1e-149
Arabidopsis NAC domain containing protein 37, vascular related NAC-domain protein 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0099
ALDH-like
PF00171
Aldedh
Aldehyde dehydrogenase family
CL0099
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
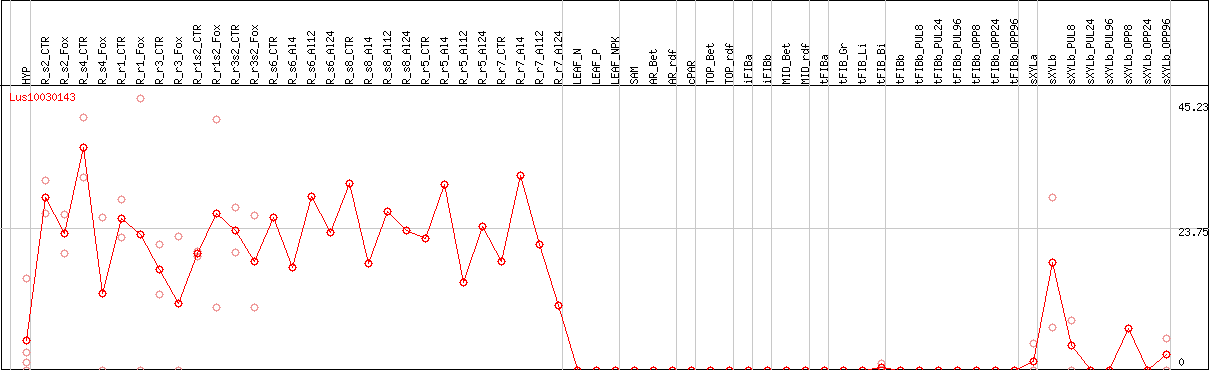
>Lus10030143 pacid=23172547 polypeptide=Lus10030143 locus=Lus10030143.g ID=Lus10030143.BGIv1.0 annot-version=v1.0
ATGGATGAAGTTTTGATCCAACACGATGAGTTAAAATGTGATCGCATTGCGCTTTGGACGTTGATTGCTGATACATGGTTAGATACAAAGGAAGGAATAT
ACGATGAATTCGTCGGCAAGTTGGCTGTGAAAGCCAAAGACTGGGTCGTTGGTGATCCTTTCCACCCGGCTTCTCGTCTTGGTCCACAGGTCGATAAGAA
ACAGTACGAGAAGATACTGCACATTGAGCAAGGAAAGAAAGAAGGAGATACCTTATTAACCAGCGGTAAAGCTGCTGCCAATAAGGGATATTACATCCAT
CCTACTGTCTTCACCGATGTTACGGATATGACCATAGTGAAAGACGAGATCTTCGGTCCGGTGATGTCACTCATGAAGTTCAAAACGACGGAAGAAGCGA
TCAAGTTGGCGAACAATACAAGGTATGGGTTAGCAGTGGGGATAGTGACCAACAATTTGGACATTGCAAACACAATTTCAAGATTGATCAGAGCGGGAGT
CATCTGGATTAACTTTACTTTGTGTTCGATTATGATTGCCCTTACGGAGGGTACAAATCTATTGCAGCTCTATATATTGATAAGATTCATGGCGGGGAGT
GGGCAGCTGACGGTGCCTCCAGGTTTCAGGTTCCACCCAACGGATGAGGAACTTCTGTACTATTATCTCAGGAAGAAAGTTTCTTATGAGGCTATTGACC
TCGACGTTATCCGGGAAGTCGACCTCAACAAGCTCGAGCCTTGGGACCTCAAAGAGAAGTGCAGGATAGGAACAGGGCCTCAAAATGAGTGGTATTTCTT
CAGCCACAAGGACAAGAAGTACCCGACTGGGACTCGAACCAACCGAGCAACTACGGCCGGGTTTTGGAAGGCGACGGGGAGGGACAAAGCCATACACCTC
GGAAATTCACAGAGGATAGGGATGCGTAAAACTCTCGTGTTCTATACCGGACGTGCACCTCATGGCCATAAAACCGACTGGATTATGCATGAATATCGTC
TCGACGACGACAACTCTGACGTTCAGGTACTTCTAAATCATGCGTTGTAA
AA sequence
>Lus10030143 pacid=23172547 polypeptide=Lus10030143 locus=Lus10030143.g ID=Lus10030143.BGIv1.0 annot-version=v1.0
MDEVLIQHDELKCDRIALWTLIADTWLDTKEGIYDEFVGKLAVKAKDWVVGDPFHPASRLGPQVDKKQYEKILHIEQGKKEGDTLLTSGKAAANKGYYIH
PTVFTDVTDMTIVKDEIFGPVMSLMKFKTTEEAIKLANNTRYGLAVGIVTNNLDIANTISRLIRAGVIWINFTLCSIMIALTEGTNLLQLYILIRFMAGS
GQLTVPPGFRFHPTDEELLYYYLRKKVSYEAIDLDVIREVDLNKLEPWDLKEKCRIGTGPQNEWYFFSHKDKKYPTGTRTNRATTAGFWKATGRDKAIHL
GNSQRIGMRKTLVFYTGRAPHGHKTDWIMHEYRLDDDNSDVQVLLNHAL