External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G00870 117 / 8e-30
bHLH
bHLH014
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G32640 117 / 2e-29
bHLH
JIN1, JAI1, ZBF1, RD22BP1, ATMYC2
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT4G17880 112 / 3e-27
bHLH
bHLH004, MYC4
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT5G46760 100 / 3e-23
bHLH
bHLH005, ATR2, MYC3
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT5G46830 94 / 2e-21
bHLH
ATNIG1, bHLH028
NACL-inducible gene 1 (.1)
AT4G09820 76 / 3e-15
bHLH
BHLH42, TT8, bHLH042
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G46510 73 / 3e-14
bHLH
ATAIB, bHLH107, bHLH017, INU1, JAM1, MYL1
ABA-inducible BHLH-type transcription factor (.1)
AT1G63650 71 / 2e-13
bHLH
EGL1, ATMYC-2, EGL3
ENHANCER OF GLABRA 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT5G41315 71 / 2e-13
bHLH
MYC6.2, GL3
GLABROUS 3, GLABRA 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G01260 65 / 2e-11
bHLH
bHLH013, INU4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005999
374 / 4e-128
AT1G32640 268 / 3e-85
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10004574
115 / 3e-28
AT1G32640 618 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10035366
109 / 1e-26
AT1G32640 376 / 9e-125
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10030970
110 / 2e-26
AT1G32640 630 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10000484
108 / 3e-26
AT4G17880 562 / 0.0
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10032267
81 / 9e-17
AT5G41315 487 / 2e-165
GLABROUS 3, GLABRA 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10024631
81 / 1e-16
AT1G63650 470 / 3e-159
ENHANCER OF GLABRA 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10004575
80 / 2e-16
AT4G17880 445 / 9e-152
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10030243
63 / 1e-10
AT1G01260 505 / 2e-173
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G176900
227 / 1e-70
AT4G17880 277 / 2e-86
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.014G103700
213 / 2e-65
AT4G00870 252 / 1e-78
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G083500
157 / 2e-44
AT4G00870 218 / 5e-66
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.003G147300
145 / 1e-39
AT4G00870 209 / 1e-62
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G142200
118 / 2e-29
AT1G32640 604 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.003G092200
113 / 1e-27
AT1G32640 559 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.002G054100
89 / 2e-19
AT4G09820 317 / 6e-100
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G208600
86 / 3e-18
AT4G09820 296 / 2e-92
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G083900
67 / 3e-12
AT5G41315 464 / 5e-156
GLABROUS 3, GLABRA 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G099700
65 / 2e-11
AT1G01260 556 / 0.0
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
PFAM info
Representative CDS sequence
>Lus10030212 pacid=23153066 polypeptide=Lus10030212 locus=Lus10030212.g ID=Lus10030212.BGIv1.0 annot-version=v1.0
ATGGATGAGCTCATAATCTCGCCGTCAACTTCCTCCTCCGTCCTAACCGCTCCTCCTCCGCCGCCTCCTACTCTCCAGCAACGTCTCCAGTTCCTCCTCC
AGACCCAAATCCCAGACTGGTGGGCCTACGCCATCTTCTGGCAGACCTCCAACGATGCCGAGTCGGGTCGGGTCTTCCTCTCCTGGGCAGACGGCCATTT
CCAAGGCTCCCCTGACTTCCCTCCGAAGCTCACCTCCTCCTCCCCCAACAATGACGTTTCCGCCCCCGGAAACGATGCCGTATCGGAGGCGGAGTGGTTC
TACGTCATGTCACTGACCCGCTCCTTCTCCGCCGGGGACGGCGTCCCGGGCAAGGCTCTGACCACTGGCTCCCTAATGTGGCTGACCGGATCACACGAGC
TCCAGTTCCACAACTGCGATCGGGCCAAGGAGTCCCAGCTCCACGGTGTCAGGACTTTAGTCTGCATCCCTGCCCACGGCGGGGTAGTAGAGCTCGCGTC
ATGTGACGTCATCAAGGAGAACTGGGGGCTCGTCCAGCAGGTCCAGTCCCTGTTCGGGTCGGATTCATCTTCTTCTTCTTCGATCCCCAAAAATAACCCG
CTTCCGTTTAACCTCGACATGAACATCTCGTTTGCCGACATCGGGATCATCGCCGGAATCCAAGAGGACGATTATGATGACGACAACCTTAATAACAATA
TGGATAACAGCAACAATAATAAGTCGTCATTGTTGCTGCCACTGGACTCGGAGCATTATTCTGACTCGGATAGTCCGGTGAACAAAACGTCTCCGTTGAT
GAAGAGGGTGCCGAAGAAGAGAGGGAGGAAGCCGAGGATGCCGGGGAAATCGGGGGAAGGGGGCCCCACTGAACCGCGTGGAGGCAGAGCGGCAGAGGCG
GGAAAAGCTCAACCACAGATTCTACGCTCTACGCGCGGTGGTACCTAA
AA sequence
>Lus10030212 pacid=23153066 polypeptide=Lus10030212 locus=Lus10030212.g ID=Lus10030212.BGIv1.0 annot-version=v1.0
MDELIISPSTSSSVLTAPPPPPPTLQQRLQFLLQTQIPDWWAYAIFWQTSNDAESGRVFLSWADGHFQGSPDFPPKLTSSSPNNDVSAPGNDAVSEAEWF
YVMSLTRSFSAGDGVPGKALTTGSLMWLTGSHELQFHNCDRAKESQLHGVRTLVCIPAHGGVVELASCDVIKENWGLVQQVQSLFGSDSSSSSSIPKNNP
LPFNLDMNISFADIGIIAGIQEDDYDDDNLNNNMDNSNNNKSSLLLPLDSEHYSDSDSPVNKTSPLMKRVPKKRGRKPRMPGKSGEGGPTEPRGGRAAEA
GKAQPQILRSTRGGT
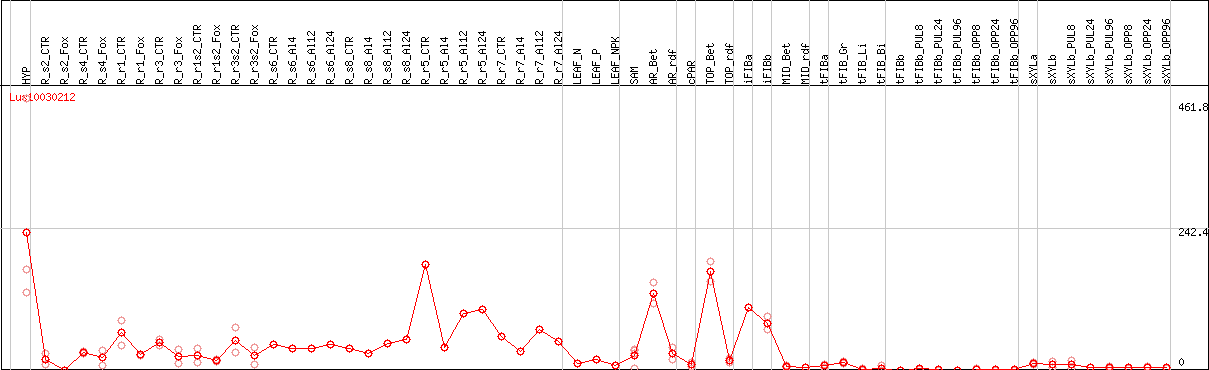
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030212 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.