External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01200 202 / 1e-66
ATRAB-A3, AtRABA3
ARABIDOPSIS RAB GTPASE HOMOLOG A3, RAB GTPase homolog A3 (.1)
AT5G65270 172 / 7e-55
AtRABA4a
RAB GTPase homolog A4A (.1)
AT5G47960 168 / 3e-53
SMG1, AtRABA4c
SMALL MOLECULAR WEIGHT G-PROTEIN 1, RAB GTPase homolog A4C (.1)
AT4G39990 164 / 8e-52
ATGB3, AtRABA4b, AtRab11G
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A 4B, GTP-BINDING PROTEIN 3, RAB GTPase homolog A4B (.1)
AT3G12160 161 / 1e-50
AtRABA4d
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
AT1G09630 149 / 5e-46
ATRAB-A2A, ATRAB11C, ATRABA2A
ARABIDOPSIS RAB GTPASE A2A, RAB GTPase 11C (.1)
AT1G28550 141 / 1e-42
AtRABA1i
RAB GTPase homolog A1I (.1)
AT3G15060 140 / 1e-42
AtRABA1g
RAB GTPase homolog A1G (.1)
AT5G45750 140 / 2e-42
AtRABA1c
RAB GTPase homolog A1C (.1)
AT1G16920 140 / 2e-42
ATRABA4B, RAB11, ATRABA1B
RAB GTPase homolog A1B (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005990
300 / 8e-106
AT1G01200 206 / 1e-67
ARABIDOPSIS RAB GTPASE HOMOLOG A3, RAB GTPase homolog A3 (.1)
Lus10023153
161 / 2e-50
AT5G65270 396 / 3e-142
RAB GTPase homolog A4A (.1)
Lus10000637
160 / 6e-50
AT5G65270 395 / 8e-142
RAB GTPase homolog A4A (.1)
Lus10025738
158 / 3e-49
AT5G65270 404 / 4e-145
RAB GTPase homolog A4A (.1)
Lus10035924
158 / 3e-49
AT5G65270 404 / 3e-145
RAB GTPase homolog A4A (.1)
Lus10023829
162 / 4e-49
AT3G12160 377 / 5e-132
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
Lus10016284
155 / 2e-48
AT3G12160 386 / 2e-138
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
Lus10021023
162 / 9e-48
AT3G12160 384 / 3e-132
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
Lus10012032
151 / 2e-46
AT3G12160 379 / 1e-135
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00071
Ras
Ras family
Representative CDS sequence
>Lus10030221 pacid=23153013 polypeptide=Lus10030221 locus=Lus10030221.g ID=Lus10030221.BGIv1.0 annot-version=v1.0
ATGTATATTATCGAATACAGGGCAGTTACAAGCGCATACTACAGAGGAGCACTAGGAGCAATGCTGGTGTACGACATCACAAAGAGGCCGACCTTTGACC
ACGTGGCGAGGTGGGTGGAGGAGCTCCGGGCCCACGCCGACAGTTCGATCGTGATCATGCTGATCGGCAACAAAGCCGATCTAGCAGGTGACCACAGGGC
AGTCCCCACGGAGGACGCGGTGGAGTTTGCCGAGGAGCAGGGGCTCTTCTTCTCGGAGGCCTCAGCGATGAGCGGGGACAACGTGGACAAGGCCTTCTTC
CAGGTGCTGGAGAAGGTGCACGCCGCGGTGTCGAAGAAGTCGCTGCAGGCTACAGCTGCGGCCTCTTCTGATGTCGCGAAATCGAACGGAGCGTTGCAAG
GGGATAGGATTGATGTGATTAGTGGGGCGGAAATGGAGATAAGTGAGATGAAGAAGTTGCCGTCTTGCTCTTGTTGA
AA sequence
>Lus10030221 pacid=23153013 polypeptide=Lus10030221 locus=Lus10030221.g ID=Lus10030221.BGIv1.0 annot-version=v1.0
MYIIEYRAVTSAYYRGALGAMLVYDITKRPTFDHVARWVEELRAHADSSIVIMLIGNKADLAGDHRAVPTEDAVEFAEEQGLFFSEASAMSGDNVDKAFF
QVLEKVHAAVSKKSLQATAAASSDVAKSNGALQGDRIDVISGAEMEISEMKKLPSCSC
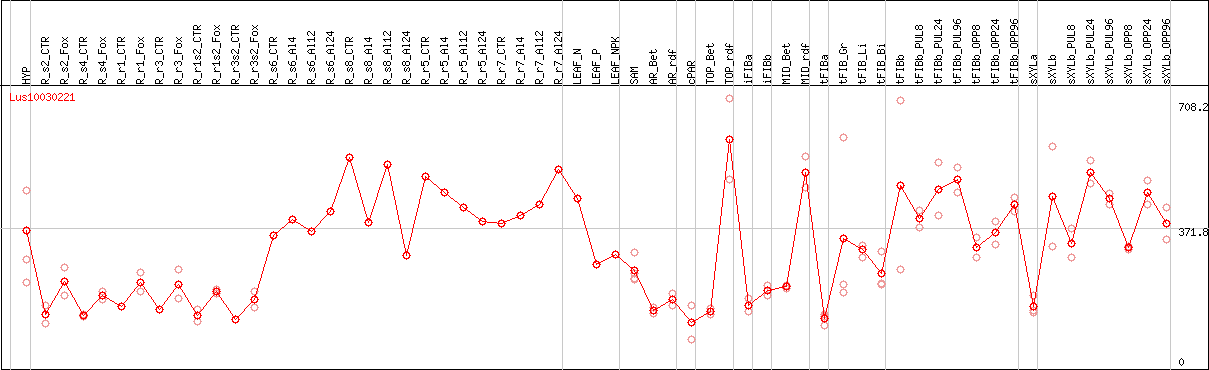
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030221 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.