External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01250 67 / 1e-14
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT5G25810 50 / 5e-08
AP2_ERF
TNY, TINY
TINY, Integrase-type DNA-binding superfamily protein (.1)
AT3G60490 47 / 4e-07
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT5G11590 47 / 5e-07
AP2_ERF
DREB3, TINY2
TINY2, Integrase-type DNA-binding superfamily protein (.1)
AT2G44940 47 / 7e-07
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT3G16280 45 / 2e-06
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G32800 45 / 3e-06
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT4G16750 44 / 4e-06
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT2G35700 44 / 7e-06
AP2_ERF
ATERF38
ERF family protein 38 (.1)
AT2G25820 43 / 1e-05
AP2_ERF
ESE2
ethylene and salt inducible 2, Integrase-type DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005967
145 / 3e-45
AT1G01250 141 / 1e-42
Integrase-type DNA-binding superfamily protein (.1)
Lus10007799
50 / 2e-08
AT2G44940 136 / 7e-42
Integrase-type DNA-binding superfamily protein (.1)
Lus10023873
49 / 4e-08
AT2G36450 121 / 3e-35
HARDY, Integrase-type DNA-binding superfamily protein (.1)
Lus10014376
50 / 5e-08
AT2G36450 167 / 2e-52
HARDY, Integrase-type DNA-binding superfamily protein (.1)
Lus10004738
50 / 6e-08
AT2G35700 144 / 7e-44
ERF family protein 38 (.1)
Lus10002801
49 / 1e-07
AT5G11590 209 / 4e-68
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Lus10034949
44 / 6e-06
AT4G32800 169 / 3e-51
Integrase-type DNA-binding superfamily protein (.1)
Lus10038270
43 / 1e-05
AT5G11590 168 / 4e-52
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Lus10025834
42 / 5e-05
AT5G11590 164 / 2e-50
TINY2, Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10030242 pacid=23152995 polypeptide=Lus10030242 locus=Lus10030242.g ID=Lus10030242.BGIv1.0 annot-version=v1.0
ATGGCCGCCAAGGCGTACAACGTCGCAGCGCACTGCCTCAAAGGACGCAGCGCCCTCCTCAATTTCCCCGACCAGGTCAACGACTTACCCACTCCTGCCA
CGTGTCGCTCTCCCAGGGACATCCAGGCCGCCGCCGCCGTAGCCGCACGGTCCGTTATTATCCGTCATCATTCCTCCTCTTCCTCCTCGCCCAGCTCCTC
GCCGTTAAGTGACGGCGCTGGGGGTAACGGAGTTGAGAGGAGTGATAATGAGGATGAGGATGGTGACGGCGGGTTTTGGGAGGAGATTGAGTTGCCGGAA
CTGCTCATTACCAGCAGTAGTAGCTGTGATGAGCTGTGGAGTTTGGCTGCCGGCGGGAAGGATTCTGCGGCGGATTGGTAG
AA sequence
>Lus10030242 pacid=23152995 polypeptide=Lus10030242 locus=Lus10030242.g ID=Lus10030242.BGIv1.0 annot-version=v1.0
MAAKAYNVAAHCLKGRSALLNFPDQVNDLPTPATCRSPRDIQAAAAVAARSVIIRHHSSSSSSPSSSPLSDGAGGNGVERSDNEDEDGDGGFWEEIELPE
LLITSSSSCDELWSLAAGGKDSAADW
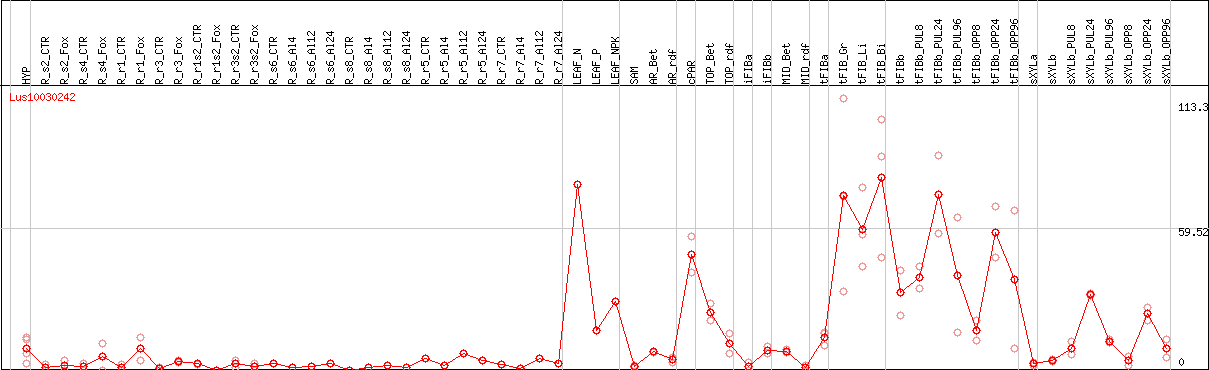
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030242 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.