External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G46500 485 / 2e-170
UBDKGAMMA4, ATPI4KGAMMA4 ,PI4K GAMMA 4
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
AT1G64460 436 / 5e-155
Protein kinase superfamily protein (.1)
AT5G24240 439 / 2e-152
Phosphatidylinositol 3- and 4-kinase ;Ubiquitin family protein (.1)
AT1G13640 206 / 2e-61
Phosphatidylinositol 3- and 4-kinase family protein (.1)
AT2G03890 205 / 6e-61
UBDKGAMMA7, ATPI4KGAMMA7 ,PI4K GAMMA 7
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
AT1G26270 203 / 2e-60
Phosphatidylinositol 3- and 4-kinase family protein (.1)
AT2G40850 200 / 9e-60
ATPI4KGAMMA1 ,PI4K GAMMA 1
phosphoinositide 4-kinase gamma 1 (.1)
AT3G56600 191 / 1e-56
Protein kinase superfamily protein (.1.2)
AT1G27570 102 / 4e-24
phosphatidylinositol 3- and 4-kinase family protein (.1.2)
AT3G43950 57 / 2e-09
Protein kinase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004005
627 / 0
AT2G46500 483 / 1e-169
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
Lus10018104
476 / 4e-167
AT5G24240 689 / 0.0
Phosphatidylinositol 3- and 4-kinase ;Ubiquitin family protein (.1)
Lus10022411
476 / 8e-167
AT5G24240 731 / 0.0
Phosphatidylinositol 3- and 4-kinase ;Ubiquitin family protein (.1)
Lus10003190
460 / 2e-161
AT2G46500 587 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
Lus10001817
459 / 4e-160
AT2G46500 665 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
Lus10019197
210 / 1e-62
AT2G03890 873 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Lus10036834
209 / 3e-62
AT2G03890 874 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Lus10005095
192 / 4e-56
AT2G03890 700 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G099000
511 / 3e-180
AT2G46500 784 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
Potri.012G014600
493 / 2e-173
AT5G24240 767 / 0.0
Phosphatidylinositol 3- and 4-kinase ;Ubiquitin family protein (.1)
Potri.001G090300
486 / 2e-170
AT2G46500 699 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 4, phosphoinositide 4-kinase gamma 4 (.1.2.3)
Potri.015G013600
464 / 2e-161
AT5G24240 747 / 0.0
Phosphatidylinositol 3- and 4-kinase ;Ubiquitin family protein (.1)
Potri.008G112200
214 / 2e-64
AT2G03890 866 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Potri.010G137400
213 / 6e-64
AT2G03890 917 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Potri.006G031600
204 / 3e-61
AT2G40850 489 / 2e-168
phosphoinositide 4-kinase gamma 1 (.1)
Potri.017G101500
180 / 6e-52
AT2G03890 737 / 0.0
UBIQUITIN-LIKE DOMAIN KINASE GAMMA 7, phosphoinositide 4-kinase gamma 7 (.1.2)
Potri.002G171400
85 / 8e-21
AT1G64460 86 / 6e-22
Protein kinase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00454
PI3_PI4_kinase
Phosphatidylinositol 3- and 4-kinase
Representative CDS sequence
>Lus10030251 pacid=23153068 polypeptide=Lus10030251 locus=Lus10030251.g ID=Lus10030251.BGIv1.0 annot-version=v1.0
ATGTCGGCTGCTGGGAATGATCCAATCAAGTCTATGGAAGGTACAGGAGGAGCTTATTTCATGCAGGATTCTCTTGGGCAGAAGTTTGTTTCGGTTTTTA
AACCGATAGACGAGGAGCCTCTAGCTGTCAACAACCCTCAGAAGTTGCCTTCTTCGCCTAACGGTGAGGGACTGAAGAAAGGTACGGTAGTCGGAGAAGG
TGCTCTGAGGGAAGTTGCAGCGTACATTTTGGATCATCCAAAGGGAGGAAAGAGGTCGTTATTCGGGAACGATGAACACGGATTTGCTGGAGTTCCGCCG
ACTGTTATGGTGAAGTGTTTGCACAGAGGATTTCACCATCCGGAGGGTGTGAACGTTAAGATAGGTTCTTTGCAGATGTTTATCAAGAACGAGGGCAGTT
GTGAGGATATCGGTCCTGGTGTTTTTCCTGTCGAGGAAGTTCACAAAATCGCTGTTTTCGATATAAGGCTGGCTAATGCGGATAGGCATGGTGGGAATCT
TCTGTTCCGGAAAGATGCGGAATCTGGCAGGACTGTGTTGATTCCGATCGATCATGGATACTGCTTGCCTGAAAGTTTCGAAGACTGCACGTTCGAGTGG
CTCTACTGGCCGCAAGCTCGTCAGCCTTTCAACGCAGAGACTGTCGACTACATAAGATTGCTGGACGCAGAAGACGATATCGCCCTTCTCCAGTTCCACG
GCTGGGATGTGCCGGCCGGTTGTGCTCGGACGCTTCGAATCTCCACCATGCTTCTGAAGAAAGGAGTCGAGAAGGGGCTGACTCCCTTCGACATCGGGAA
CATCATGTGCAGAGAGAGGTTGAACAAGGAATCGGCAATCGAGGAGATTGTTCGCGAAGCAGAAGACTTGGTTCTTCCAGGGACGAGCGAGGAAGCGTTG
TTCGAGGCCATATCGGAGATCATGGATCGCCGTCTCGATGAGATTGCCGTGACGAAGTAG
AA sequence
>Lus10030251 pacid=23153068 polypeptide=Lus10030251 locus=Lus10030251.g ID=Lus10030251.BGIv1.0 annot-version=v1.0
MSAAGNDPIKSMEGTGGAYFMQDSLGQKFVSVFKPIDEEPLAVNNPQKLPSSPNGEGLKKGTVVGEGALREVAAYILDHPKGGKRSLFGNDEHGFAGVPP
TVMVKCLHRGFHHPEGVNVKIGSLQMFIKNEGSCEDIGPGVFPVEEVHKIAVFDIRLANADRHGGNLLFRKDAESGRTVLIPIDHGYCLPESFEDCTFEW
LYWPQARQPFNAETVDYIRLLDAEDDIALLQFHGWDVPAGCARTLRISTMLLKKGVEKGLTPFDIGNIMCRERLNKESAIEEIVREAEDLVLPGTSEEAL
FEAISEIMDRRLDEIAVTK
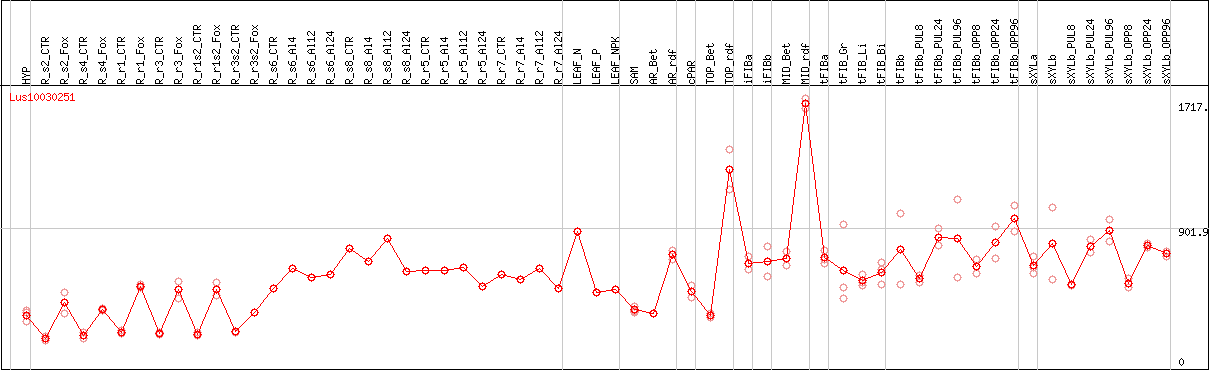
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030251 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.