Lus10030265 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030265 pacid=23153095 polypeptide=Lus10030265 locus=Lus10030265.g ID=Lus10030265.BGIv1.0 annot-version=v1.0
ATGGGCGAGTATGAAACTGCGGAGTCACAGCCACCAAGTGTGCTCTTGCCCAACGGGTTACTGCCTGATGAAGCAGCCTCCGTGATGCAAGTGCTTGACT
CGGCCAGATGGTCGAAAGCTGAAGAGAGAACTGCTGAACTCATAGAATGCATTAAGCCAAACGAACCCTCTGAGAACCAACGTAAGGCCGTGGCATCTTA
TGTGCAGCAGCTTATTGCCAAGTGCTTTCCTTGCCAGGTCTTCACTTTCGGGTCCGTTCCGCTTAAAACTTATTTGCCAGATGGAGATATTGACTTGAGC
GCCTTCAGTAAGAATCAGACTTTGAAAGAGACTTGGGCTCATCAGGTTAGGGATATGTTGGAGAACGAGGAGAAGAATGAGAATGCCGAATTTCGTGTAA
AGGAAGTGCAGTACATTCAGGCCGAGGTGAAACTGGTAAAGTGCCTTGTGGAGAACATCGTTGTGGATATATCGTTTAACCAGCTTGGTGGGCTATGCAC
ACTCTGTTTCCTTGAAGAGGTTGATCATAAGATAGGCAATGATCATTTACTCAAGCGGAGCATTATTCTTATAAAGGCATGGTGTTATTATGAAAGCCGT
ATATTGGGTGCTCACCATGGTCTGATATCCACTTATGCACTGGAAACTCTGGTTCTCTATATCTTTCATGTTTTCAACAGTTCCTTCGCTGGACCACTTG
AGGTCCTCTATCGTTTCCTTGAGTTCTTCAGTAAATTTGATTGGGACAACTTCTGTGTTAGCCTCTGGGGCCCTGTACCAACCAGTTCACTCCCAAACAT
GACAGCTGAAGCACCCCGAAAGGATGATGGAGATTTACTACTTGAAAAATCATTTCTTGATGCCTGTGGCTTGGCATATGCAGTTTTTCCCAATGGTCAA
GAAAATAATGGCTCCCCGTTCATCTCCAAGCACTTCAATGTCATTGATCCTTTGCGTGTTAACAACAACCTGGGTCGCAGTGTTAGCAAAGGTAACTTCT
TCAGGATTCGTAGTGCATTTGCATTTGGGGCAAAGAAGCTTGCTAGGTTACTTGAATGTGACAGCGACAACATACACACTGAAGTGAATCAGTTTTTCCA
GAATACCTGGGAAAGACATGGAAGTGGTCACCGTCCTGATGCTCCAATTAATGCTTTGGGGCGGCTGATTGGGTCAAGGAATGAGAACCTCATAAATACT
TCAAATCGTGGAAGTCAAAATGATATATCACAGCATGGTAGTCGTCCATCAGAAAGCACGTCACAGAGTAGTGATATTGCCACAGCAGTTCGAACACAAA
TTCAAATGATTAACAACTCCAGTTTGAGGACATCTGATGAGTCTAGACCGGAAATTAATTCAGATAATGGCGTTCACGCTGAAAATGATCCTAGAACTTC
TAAACCTGACAGTGCGTTTAGTGGTTTACAAGGAAGGTATATATTTGCTAGGACACGCTCAAGTCCTGAGCTTACTGATACCTATAATGAAGCTTCCTCA
CAAACTAGACAGAATAAAGCACCTGAAGCAAAGGGTCAGGCCCCGGCAGTGATGTTGGACAGCATCAAGAGACAGAACACTGAACATGAAAAGTCATCTA
GGCATGGTACTGGATCTTTAGAAGATGATGCTTCTGTTACACGCAGCTCAATGCACCAGAATCTTGATGCTGCTGAATCAAGTAACTATAGTGATGATTC
AGCCCTTTTTGCTGCAAGTGAGGAATTCACTTCTATCCTGGGTACACAAGGGATGTATCAGGATGCCGTGAACATGATGGCTTCTTCGAGTGGACATGGC
TTTAATGGACAAGTCCATCTTCCCCTGAATTTAGTTTCTGGTCAGATGCCCTTTCCCATACCACCTGTTGCTCTAGCTTCTATGGGATATTTTCAACGGA
ATTTAGGTGGAATGGTACCACCAAATACTTCATTGTTGGAGACCCATTGGGGTCCAAGCATGCAATTTCATCAGGGTTTGTTTCCTTCTCAGTTGAATCA
CTTTTTTCCCACTGTAGGATTGGGATCTAGTCCAGAAGGTTCAGTTGAACATTCAAATGAGAACTTAAGTTCTTTGGAATCAAATCTTGTGGTGGCTGGC
AACGATTTCTTGCATGAACATGAAAGGGCTTTTAGTGATGGTCTTGAACTTGATAATGGCAGTTCCCTGATGCATCAATCTGACGACAAACATCAATCAA
GTAGTAATCACTTTGTTCCTTTGTCTAGAACTGGTGGCTCTAGGAACTCTTCTAGAGTTCAACCTTGGGAGTCTCAACATTCAACTAGGCGGGATCATAT
GGAAATTGATCCTTCGGAAAACAGAGGTGCTGAAGTGCTCATTGAGGAAAACCTTGCTAGCTCAAGGCCTTCGTCAAGTAATTCTTGGAGAAGTAAGCCT
GCATCTGAGAGCTCATCTGATGGATTATCTGCAAAGACAAAGACAGTGAGGGAAAAGAGAGGGAGGAAGACTTCGGCCGCCGTTGCTTATGGGAAAGGTA
GGGGAATCAATGAGAATGGCTCCCTTCACATGGATGATGACACGCAGCAGGTCTCCTCAAGTCCTGAGACTGTTGAAAGGGGCTTCGGGCCTCCAGCTCC
TAGTGCTTTGCATGTTTCAAGGCATAATAATGTGCCTCGATTTGATCTGGCACAGACAGGTGAATCTGATTCATTAGCATCTATTCCTCCGGTGCAATTA
GGTCATGGTGCACGGTTTTATCCTACGGGGCCACCTGTTCCATTTGTTACGATGCTACCCATTTATAACTTCCCAACAGAGAGGGGAATTTCTGAAGCAT
CAACTAGCCAGCATAGCTTTGAGGAGATCATAGATAACAATGTTCCACATCAGAATGGTAATTCAATGAGAGTTTCGGTCATTGACGAGCCATTTGAACA
CAAGTTTGACATCCTCAATAGCGATTTTGCTAGTCATTGGCAGAACTTGCAATTCGGAAGGTTTTGTCAAGATCCTCCGTCGCCCTCACCGGTGACCGGT
CCTTCACCGGTTGTTGTACCGCGGGTGTATTTACAGGGCCGCTTCCCAGGTCCTGGAAGGCCCATTTCACCTAATACGAATCTTTTCTCTCAGCTGATGG
GTTTCGGGCCTCGTATCCTTCCAGTTGCTCCAGTGCCAGCTGTATCAAACAGACCTGCTGCCGTCTACCACCAACGTTTAGTTGATGAGGTGCCAAGATA
CCGCGGTGGAACTGGAACCTACCTTCCAAATCCTCAGAACGTGGCTGTCCGAGATAGGCATTCTCCAAACTCGAGAAAAACAAACTACAACTTTGATAGA
AGTGACTACCATGGTGATCGGGAGGGGAACTGGAACAATTCGAAGTCGAGAGCCCCCGGGCGCAGTCAGAGTCGTAACCAGGCGGATAAGTCGAGATCTG
ACCGGCCATGGGTTTCACACAGACACGACACTACCGTTTCTTATCAGTCTCAAAACGGGCCAAGCCGACCAAATGCCTCTCTAGTCGGTTCTGCGACCAA
TACCACTTATGGCATGTATCCAATAACGTCTATGAATTCGAATGGACAGGTTGTCATGATGTATCCGTTTGACCAAAACGCTGCCGGTTATGTCTCTCCA
GCGGACCAGATCGAGTTTGGCTCGTTTGGCCCTCCGGGATACTCTGGTGTAAATGATCTAAATGAGGGAAGTAGACCCGGCGAGGGATTCGAGGACCAAA
GATTTCACAGTAGTGCAGCAGTTCAACAGTCTTCCCCGGACCCGCCTTCTTCTCCCCGCTTTCAGAGAGACCTTAAACCCTGCTCAGTCCTCTCTACTGC
ACTCAATTTTTCAGCTGCCGGTCGAGCTATGCCTATGCCCAATATCAAGCTGCTTTCCCAGCTCCGCCTTCTCAACCCGTCTGTTCCAATTACCTTTATC
CCTCCTACCTCCTCCCTCGCCGCCGCCCGTTCCATCAACCTTATCTCCTTATCTGCTGGGGGTTTCAATTTCGCCCCCGTCCGCCACTTCTCCGCCCGTG
CCTTTCGCTCCAGGACGGCCTTCTCCCATGATTTAGGCCGCTATCGCTCCAATGAATTCGGCCGCAAGGCTTCTTCTTCCAAGAGCTTGATTGACGACGA
GGCCGAGCTGAGCGACTGGGTCGGGGTTTTGCGTACCAGCTCGATTCGCGGCCGCCTCACCAGCGATGAGGACTCTGATTCCGATCGGCGATCCGGCGGC
GGCGGTAGAGGAGGCAGAGGAAGGGGAGACGGAGGGAGAAGAGAAGGTGTTGGTGATGAAGGGAGATACAGAGGGCCGCCGCCGAGGAGAACCGGCGGTA
GAAGTGGAGATTCGGAGAGGGAGTTTGATCGGACGGGGGTTCGGAGTGGTGGTAGTCAAGGGGAGTCGTTTGGGAGGAATTCAGCAGTGAGGCAGTTCAG
AGAAGGTGCGCCGCCGCCACCACAGAGGAGAAGTAACGGTAGAAGTGGAGGTTTTGAGAATGGGAGGGGAAGAGGGAGAGGAGTTGATTCAGGGTTTAGG
ACGGATGCTGTTATTGTTAAGAAGAGTCCTATAGGGAGAAGAAGCTTGTTGGTTTCAGATGAGGATGAAGACGACGAGGAAGAGGACAATGTGGCATCAA
TTGGAGGCTTTGATGATCTGTTGGAGGATGACAGTGATGATGATGACACTGAAGAAGACGGGGATGAGGACGATGATGATAGTTTGGATAAGAATAGCGC
GTCTTCTTTGTTGGGATTGGAAAAGAAGTACCGGGCGAAAGTGAGTTCTACTGGAAGTTCGGAATCTTATCTAAGTGGAACTAGATTTGACCAGTGCTCA
GTATCACCGCTGTCGTTGAAAGCAATCAAAGATGCTGGATACGAGAACATGACTGTGGTTCAGGAAGCAACGCTGCCTGTTATACTGAAAGGAAAGGATG
TTCTAGCTAAAGCTAGAACAGGTACTGGAAAAACAGTCGCATTTCTTCTTCCATCCATTGAAGTCATTGTGAAAACACCTCCTGCTGATCATGATAACAG
AAGGCCACCGATTCTCGTGCTTGTCATTTGCCCAACAAGGGAGCTCGCTATACAAGCTGCAGCGGAAGCAAACAAGCTGCTGAAGTACCATTCTTCTCTT
GGTGTTCAGTGTGTCATTGGAGGTACAAGGCTTGCCCTGGAACAGAAAAAACTGCAAGCTAATCCCTGCCAGATCCTTGTTGCTACGCCTGGGAGGCTCA
GAGATCATCTTGAGAATACAGCCGGATTCGCAACAAAAGTGAAAGGCGTGAAAGTCCTCGTTCTTGATGAAGCCGATCGGTTACTCGACATGGGATTCCG
GAAAGACATTGACAAAATAATAGCTGCTGTTTCGAAACAACGTCAAACACTTTTGTTCTCCGCTACAGTTCCTGACGAGGTTCGTCAAATGTGTCATGTT
GCGTTGAAAAGAGACCACGAATTCATTAACACGGTTGAAGAAGGATCCGAGGAGACTCATGCAAAGGTGAGCCAGATGCATATGGTTGCTCCACTTGAGA
AACATTTTCATCTTCTCTATGCACTCCTAAGAGATCATATTGCAGACAATGTTGATTACAAGGTTCTCGTCTTTTGTACCACTGCCATGGTCACAAGACT
TGTTGCAGACTGTCTTGGCGAACTCAAACTAAACATCAGAGAGATCCATTCTCGGAAGTCTCAATCTTACAGAACCAAGGTTTCTGACGAATTCCGAAAA
TCCAAGGGTCTCATACTCGTTACTTCCGACGTATCAGCACGCGGAGTCGATTATCCTGACGTTACCCTTGTAGTACAAGTCGGTATACCATCGGACAGAG
AACAATACATCCATCGACTTGGCCGAACAGGGCGCAAGGGGAAAGAAGGAGAAGGAGTACTTCTGCTTGCACCTTGGGAAGAGTTCTTTCTATCTACTGT
GAAAGATTTGCCATTAACCAAGGCTGATGTACCCACAATTGACCCTGACACAACCAAGAAGGTTGAGAGAGCACTGTCCAATGTGGAGATGAAGAGCAAA
GAAGCAGCCTATCAAGCTTGGCTGGGCTACTACAACTCCGACAAGAAAATCGGCAAAGACAAGTACAGGCTAGTGGAGCTTGCTAACGAGTTCAGCAGAA
GCATGGGACTCGACAACCCACCTGCTATTCCGAAGATCACCCTTGGCAAGATGGGTCTCAAGAACATCCCCGGGTTGCGCTCCAAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030265 pacid=23153095 polypeptide=Lus10030265 locus=Lus10030265.g ID=Lus10030265.BGIv1.0 annot-version=v1.0
MGEYETAESQPPSVLLPNGLLPDEAASVMQVLDSARWSKAEERTAELIECIKPNEPSENQRKAVASYVQQLIAKCFPCQVFTFGSVPLKTYLPDGDIDLS
AFSKNQTLKETWAHQVRDMLENEEKNENAEFRVKEVQYIQAEVKLVKCLVENIVVDISFNQLGGLCTLCFLEEVDHKIGNDHLLKRSIILIKAWCYYESR
ILGAHHGLISTYALETLVLYIFHVFNSSFAGPLEVLYRFLEFFSKFDWDNFCVSLWGPVPTSSLPNMTAEAPRKDDGDLLLEKSFLDACGLAYAVFPNGQ
ENNGSPFISKHFNVIDPLRVNNNLGRSVSKGNFFRIRSAFAFGAKKLARLLECDSDNIHTEVNQFFQNTWERHGSGHRPDAPINALGRLIGSRNENLINT
SNRGSQNDISQHGSRPSESTSQSSDIATAVRTQIQMINNSSLRTSDESRPEINSDNGVHAENDPRTSKPDSAFSGLQGRYIFARTRSSPELTDTYNEASS
QTRQNKAPEAKGQAPAVMLDSIKRQNTEHEKSSRHGTGSLEDDASVTRSSMHQNLDAAESSNYSDDSALFAASEEFTSILGTQGMYQDAVNMMASSSGHG
FNGQVHLPLNLVSGQMPFPIPPVALASMGYFQRNLGGMVPPNTSLLETHWGPSMQFHQGLFPSQLNHFFPTVGLGSSPEGSVEHSNENLSSLESNLVVAG
NDFLHEHERAFSDGLELDNGSSLMHQSDDKHQSSSNHFVPLSRTGGSRNSSRVQPWESQHSTRRDHMEIDPSENRGAEVLIEENLASSRPSSSNSWRSKP
ASESSSDGLSAKTKTVREKRGRKTSAAVAYGKGRGINENGSLHMDDDTQQVSSSPETVERGFGPPAPSALHVSRHNNVPRFDLAQTGESDSLASIPPVQL
GHGARFYPTGPPVPFVTMLPIYNFPTERGISEASTSQHSFEEIIDNNVPHQNGNSMRVSVIDEPFEHKFDILNSDFASHWQNLQFGRFCQDPPSPSPVTG
PSPVVVPRVYLQGRFPGPGRPISPNTNLFSQLMGFGPRILPVAPVPAVSNRPAAVYHQRLVDEVPRYRGGTGTYLPNPQNVAVRDRHSPNSRKTNYNFDR
SDYHGDREGNWNNSKSRAPGRSQSRNQADKSRSDRPWVSHRHDTTVSYQSQNGPSRPNASLVGSATNTTYGMYPITSMNSNGQVVMMYPFDQNAAGYVSP
ADQIEFGSFGPPGYSGVNDLNEGSRPGEGFEDQRFHSSAAVQQSSPDPPSSPRFQRDLKPCSVLSTALNFSAAGRAMPMPNIKLLSQLRLLNPSVPITFI
PPTSSLAAARSINLISLSAGGFNFAPVRHFSARAFRSRTAFSHDLGRYRSNEFGRKASSSKSLIDDEAELSDWVGVLRTSSIRGRLTSDEDSDSDRRSGG
GGRGGRGRGDGGRREGVGDEGRYRGPPPRRTGGRSGDSEREFDRTGVRSGGSQGESFGRNSAVRQFREGAPPPPQRRSNGRSGGFENGRGRGRGVDSGFR
TDAVIVKKSPIGRRSLLVSDEDEDDEEEDNVASIGGFDDLLEDDSDDDDTEEDGDEDDDDSLDKNSASSLLGLEKKYRAKVSSTGSSESYLSGTRFDQCS
VSPLSLKAIKDAGYENMTVVQEATLPVILKGKDVLAKARTGTGKTVAFLLPSIEVIVKTPPADHDNRRPPILVLVICPTRELAIQAAAEANKLLKYHSSL
GVQCVIGGTRLALEQKKLQANPCQILVATPGRLRDHLENTAGFATKVKGVKVLVLDEADRLLDMGFRKDIDKIIAAVSKQRQTLLFSATVPDEVRQMCHV
ALKRDHEFINTVEEGSEETHAKVSQMHMVAPLEKHFHLLYALLRDHIADNVDYKVLVFCTTAMVTRLVADCLGELKLNIREIHSRKSQSYRTKVSDEFRK
SKGLILVTSDVSARGVDYPDVTLVVQVGIPSDREQYIHRLGRTGRKGKEGEGVLLLAPWEEFFLSTVKDLPLTKADVPTIDPDTTKKVERALSNVEMKSK
EAAYQAWLGYYNSDKKIGKDKYRLVELANEFSRSMGLDNPPAIPKITLGKMGLKNIPGLRSK
|
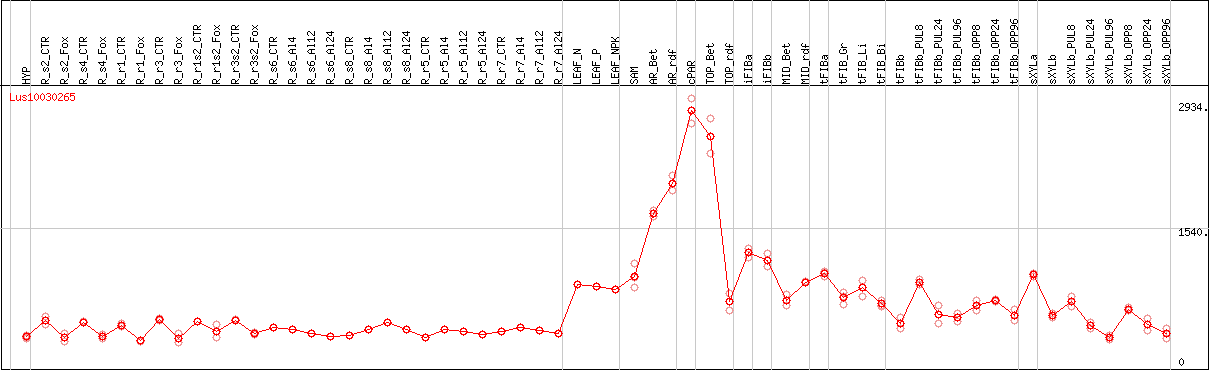
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030265 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
