External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G60820 82 / 2e-21
PBF1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015867
90 / 3e-24
AT3G60820 379 / 2e-135
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10004024
87 / 4e-23
AT3G60820 190 / 4e-61
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10009294
69 / 1e-15
AT1G48050 835 / 0.0
ARABIDOPSIS THALIANA KU80 HOMOLOG, Ku80 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G069800
86 / 9e-23
AT3G60820 362 / 6e-129
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.002G148300
82 / 3e-21
AT3G60820 387 / 2e-138
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF00227
Proteasome
Proteasome subunit
Representative CDS sequence
>Lus10030271 pacid=23152972 polypeptide=Lus10030271 locus=Lus10030271.g ID=Lus10030271.BGIv1.0 annot-version=v1.0
ATGTCGATGAATTTTTCTCCCTACGAAAACAATGGCGGGTCTTGCGTTGCGATTGCCGGAGCTGATTATTGTGTGATTGCTTCCGATACTCGGATGTCCA
CTGGCTACAATATTCTCACCAGAAACCACTCCAAAGTTTGCAAACTGATCAGAAATGATGATCCAGTAAGTCAACAGTTGCAGTAG
AA sequence
>Lus10030271 pacid=23152972 polypeptide=Lus10030271 locus=Lus10030271.g ID=Lus10030271.BGIv1.0 annot-version=v1.0
MSMNFSPYENNGGSCVAIAGADYCVIASDTRMSTGYNILTRNHSKVCKLIRNDDPVSQQLQ
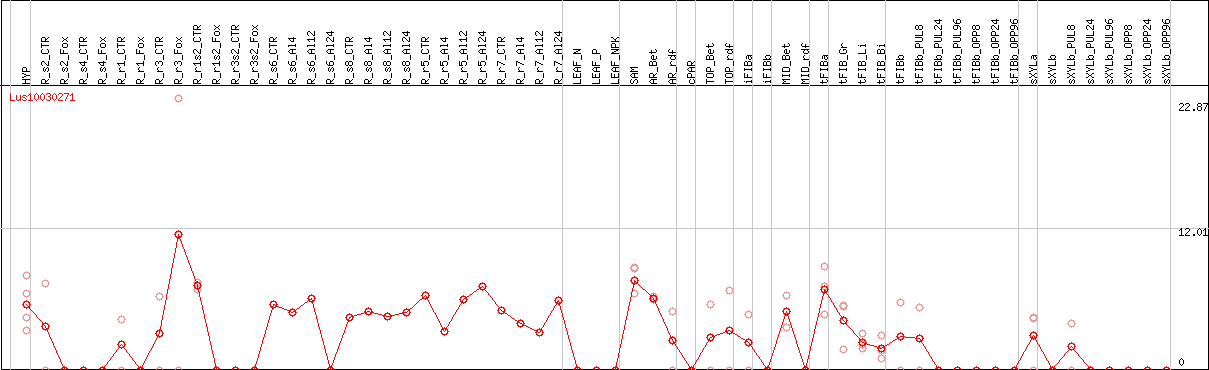
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030271 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.