External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G55330 268 / 1e-90
PPL1
PsbP-like protein 1 (.1)
AT2G39470 137 / 2e-39
PnsL1, PPL2
Photosynthetic NDH subcomplex L 1, PsbP-like protein 2 (.1.2.3)
AT3G05410 64 / 1e-11
Photosystem II reaction center PsbP family protein (.1.2)
AT5G11450 58 / 1e-09
PPD5
PsbP domain protein 5, Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
AT4G15510 57 / 2e-09
Photosystem II reaction center PsbP family protein (.1.2.3)
AT1G06680 54 / 2e-08
PSII-P, OEE2, PSBP-1, OE23
OXYGEN-EVOLVING ENHANCER PROTEIN 2, OXYGEN EVOLVING COMPLEX SUBUNIT 23 KDA, photosystem II subunit P-1 (.1.2)
AT1G77090 41 / 0.0005
Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003289
450 / 4e-162
AT3G55330 267 / 7e-91
PsbP-like protein 1 (.1)
Lus10023436
158 / 2e-47
AT2G39470 255 / 3e-86
Photosynthetic NDH subcomplex L 1, PsbP-like protein 2 (.1.2.3)
Lus10040314
157 / 3e-47
AT2G39470 254 / 5e-86
Photosynthetic NDH subcomplex L 1, PsbP-like protein 2 (.1.2.3)
Lus10000352
59 / 4e-10
AT4G15510 309 / 4e-106
Photosystem II reaction center PsbP family protein (.1.2.3)
Lus10000614
55 / 1e-08
AT4G15510 308 / 9e-106
Photosystem II reaction center PsbP family protein (.1.2.3)
Lus10029903
54 / 2e-08
AT3G05410 286 / 2e-97
Photosystem II reaction center PsbP family protein (.1.2)
Lus10022110
52 / 2e-07
AT5G11450 340 / 2e-117
PsbP domain protein 5, Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
Lus10001861
52 / 2e-07
AT5G11450 342 / 7e-117
PsbP domain protein 5, Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
Lus10020717
48 / 2e-06
AT1G06680 380 / 2e-134
OXYGEN-EVOLVING ENHANCER PROTEIN 2, OXYGEN EVOLVING COMPLEX SUBUNIT 23 KDA, photosystem II subunit P-1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G210200
325 / 4e-113
AT3G55330 299 / 2e-103
PsbP-like protein 1 (.1)
Potri.008G050300
193 / 1e-61
AT3G55330 164 / 8e-51
PsbP-like protein 1 (.1)
Potri.010G210000
149 / 5e-44
AT2G39470 278 / 3e-95
Photosynthetic NDH subcomplex L 1, PsbP-like protein 2 (.1.2.3)
Potri.005G010700
62 / 3e-11
AT4G15510 319 / 4e-110
Photosystem II reaction center PsbP family protein (.1.2.3)
Potri.013G006500
62 / 4e-11
AT4G15510 306 / 5e-105
Photosystem II reaction center PsbP family protein (.1.2.3)
Potri.005G206700
47 / 5e-06
AT1G06680 400 / 2e-142
OXYGEN-EVOLVING ENHANCER PROTEIN 2, OXYGEN EVOLVING COMPLEX SUBUNIT 23 KDA, photosystem II subunit P-1 (.1.2)
Potri.009G062500
47 / 5e-06
AT3G63540 240 / 1e-81
Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
Potri.002G055700
44 / 7e-05
AT1G06680 382 / 3e-135
OXYGEN-EVOLVING ENHANCER PROTEIN 2, OXYGEN EVOLVING COMPLEX SUBUNIT 23 KDA, photosystem II subunit P-1 (.1.2)
Potri.002G072400
42 / 0.0003
AT1G77090 345 / 2e-120
Mog1/PsbP/DUF1795-like photosystem II reaction center PsbP family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0619
Mog1p_PsbP
PF01789
PsbP
PsbP
Representative CDS sequence
>Lus10030326 pacid=23153139 polypeptide=Lus10030326 locus=Lus10030326.g ID=Lus10030326.BGIv1.0 annot-version=v1.0
ATGGCTTCCCTCCAAAACTCACCTTCCCTCTATCGCACATTGTCTTTCTTCCCCCACCAGCAGCAGGTTGGAGGATTACAGAAGCAGCAGCAAACAACGA
TTCCCAACTGTGGGAAAGGAATTTCGTTTCTTATTCGAGCCCAGCAACCATCTCTTAGAGGATTACAGAAGCAGCAGCAAACAACGATTCCCAACTGTGG
GAAAGGAATTTCGTTTCTTATTCGAGCCCAGCAACCATCTCTTAGTGAGTTCTCTATTTCAATCCCTTTGGGTAGACGCCAATTGGTGGGTGTTGGGATC
ATTGCTCCTCTAGTTTCTCTGGTGGAACAGACTTCCACTTCATTTGCTGCAGAAAACAAGAAAGGGTTCCTCTCTGTAACCGACCAGAAGGATGGATACT
CATTCGCTTACCCCTTTGGTTGGCAAGAAGTGGAGATTCAAGGACAAGATAAAGTGTTCAAAGATGTGATTGAGCCACTAGAAAGTGTCAGTGTAACCAT
GTTCCCGACTAGCAAGCTAGATATTCGAGATTTTGGCCCGCCGGACCAGGTTGCAGGAACTCTGATAAAAAAGGTTCTGGCTTCTCCTACGCAGAAGACA
AAGCTCATCCAGGCTAAAGAGCAAGATGTAGAAGGAAGAGCTTACTACACATTTGAGTTCGTTGCTCAGGCTCCCAACTTCACCCGTCACGCTCTCAGCG
CAATTGTTATCGGCAATGGAAAGCTATACACCTTGACAACTGGAGCGAAAGAAAGAAGATGGGAGAAGATGAAGGATAAATTGCAGACGATCGTGGACTC
CTTCAAAGTTGTCAATGTCTGA
AA sequence
>Lus10030326 pacid=23153139 polypeptide=Lus10030326 locus=Lus10030326.g ID=Lus10030326.BGIv1.0 annot-version=v1.0
MASLQNSPSLYRTLSFFPHQQQVGGLQKQQQTTIPNCGKGISFLIRAQQPSLRGLQKQQQTTIPNCGKGISFLIRAQQPSLSEFSISIPLGRRQLVGVGI
IAPLVSLVEQTSTSFAAENKKGFLSVTDQKDGYSFAYPFGWQEVEIQGQDKVFKDVIEPLESVSVTMFPTSKLDIRDFGPPDQVAGTLIKKVLASPTQKT
KLIQAKEQDVEGRAYYTFEFVAQAPNFTRHALSAIVIGNGKLYTLTTGAKERRWEKMKDKLQTIVDSFKVVNV
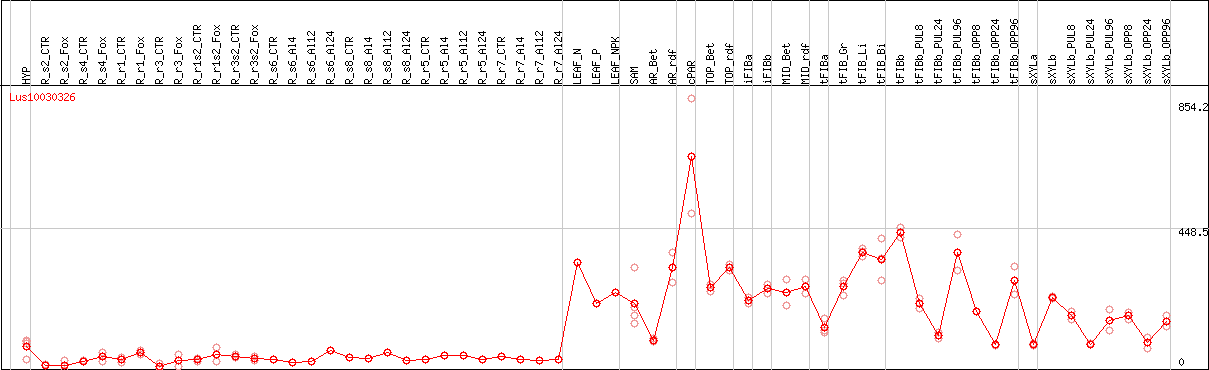
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030326 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.