Lus10030445 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Lus10030445 pacid=23140692 polypeptide=Lus10030445 locus=Lus10030445.g ID=Lus10030445.BGIv1.0 annot-version=v1.0
ATGTATTCGAGAGGTAATGGGTATGGGCAGCAATCGTATGGTGGGCAGTCTAGCTACGCTCAAAGTTTAGCAAATGCGTATTCTGCTAGCTCTGTCGGAG
GAACGGATTCGAGTTCTCAACATTCGCTGGCTTCTAGGCATCCTTCATTGTTGAGTAGTTCTCAGGAAGCGGATCTTGCTACTGGGTATAGAAGCCACAA
CTCGGTTTCGGGGCACTATGGTGGGCAGTATGCATCTCTATATGGTTCAGCTGGTGGAGTGAGTGCTACTCAACAGGTACCCTCAGCGAGTACCAAGGGA
AGTGGGCTATCGGGTTTGGAAGGCCGTAGAGGCTATTCTTCAGCTGTGTCCGATTCACCTAAATTTGGATCGGGCGACTATGTTCCTTCTTCAACCCATA
CATATGGACACAAAAGTGATCCATTGTACTCAGACAAGATCGCTGATTACACTGGAATAGACCGCAGGCATTATGGCGAGCAGAGTTCATATACCGGCAG
GGACTTGCAAACTGATCCTGCTTCACGGTATACAGATTCTGTTGGCTTGAGTCATCAACGTCAGCAGACTGACAGATATGATCGTCTGGATCAGTCCTCG
ATTCTGCGGCAAGAGCAGTTGTTGAAATCACAATCAATTCACTCTACATCTCTTGATGGAGGTTCCAGACCGCTCGATTATCTAGCAGCAAGAGGGGGCG
CCAGTCGTCATCTCAACCAGGATCTCGCGTCTTATGGCAGACGAGTGGATGCTGATCCTCGTGGCTCCTCAATTCTTAATGCTTCTTCTTACAACACGAC
ACCTGCTTCATCAATATTAGGGGCAGCTCCACGAAGGAACGTTGATGATCTTCTATATTCCCAAAGTTTGACGAATCCAGGCTATGGTGTTAGCCTTCCT
CCCGGTAGAGATTATGGTTCAGGAAAAGCCGCGTTGCAGCAGGGAACATCACTCGATGCGGAATATCGTGCTCTTCATCCGAGGATTGAAGAAAATCGCA
GGGATGAAAGAGCCGGCTATCTTCGGGAGTTTGAGCTGCGAGAGGATGAACGTCGTCGAGAGTTCTTGCGTGAGAGGGAAAAAGAACGGGAGAGGGAACG
AGAAAGGGAGAAGGAAAGGGACCGGGATCGTGAAAGGGAACGAGAACGAGAACGGAGGCGCGACCGGGAGCGCATGTTGGAGCGGCGGGAGAAGGAGCGG
GAGAGGAAGCGCGCAATTGAAATTAGACGAGAACGAACTCCGCAGAGGCCTTCAAGGGATCCTCGTGGATCCTCTGTGACAAAGGAAGGACGATCCTTGC
GACGCGATTCAATAAGCCAAGACGCTTCACATAGGCGTCATTCACCTGTTAAAGATAAACGGAGGGAATATGTCTGCAAGGTACATGGTTCCTGTTTGGT
TGACTTAGAGAGAGACTATTTGTCGCTGGATAAGAGATACCCCAGACTCTTTGTGTCACCGGAGCTTTCTAAGGTTGTTGTCAATTGGCCAAAGGAGAAT
TTCAAGCTTTCGATACATACCCCTGTGAGTTTTGAGCATGATTTTGTGGAAGAGGAAAATGCTTTGGAGTCTAAGGACCTCTCCAGCAACCTCTTGGGGC
ATCATGATCTTAGTCAGTCAGCGAGTGGACGCACAGTATGGAACGCTAAGATAATTTTGATGAGTGGAGTGAGCAAAAGCGCTTTTGAGGTGCTGTCATC
TGCGAAGATCGCTGATGATCGCCCACCTCATATTTGTAACATACTTAGATTTGCGGTGCTTAAGAAGGATCGTGCTTTTATGACAATCGGTGGGCCTTGG
GATCCTATTGATGGCGGTTATCCATCCGTTGATGATTCTCCTTTGATTCAAACAGCACTAAGATATGCAAAGGATGTTACCCAACTTGATCTCGGGAACT
GTCGTCAGTGGAATCGGTTTCTTGAGGTACAGTACAACCGATTTGGAAAGGACGGGTTATTTAGTCATAAGGAGGTCACCGTTCTGTTTGTTCCTGATTT
GTCTGAATGTCTTCCTTCACTTGAATCGTGGAGAGACCAGTGGCTTGCACATAATAAGACTGTTTCCGACAGAGAGCGCGAGCTTTCTCTGAAAAAAGAG
AGATCTGCTGAAAAGAAGGAAGGTTCTAAAGGATCAGACCCTTCAAAAGATCCGAAACAAGGAATCAAACAAGAAAAGACCAAAGCATCTGTCTCTTCTG
GTTCCAACAAAAAGGAGAATGCTTCCAAGGACAAGACTGTATCCCCGAAGGATGCCGAAAGTAAAATGGACGAGGGGAACTTATCCAAGGATGTCTCGAA
AGATGAGCAGGGTGAACCGTCCAGTACTCCGACATCTGCTGGTGCAAAAACAGGGAAAAAAACAGGGAAAAAGAGGATTGTTAAGAGAATTGTAAAGAAA
AAGGCGGTTGCCAAGCCAGGTGGCGGTGTTGCTGCTGCCGGTGCAGCAACGAACAATTCGTCGAGTGGAGAGAAACCTGCTGAAAGTGAAGATAAGGGCT
TAGTAGAACCTGAAGGGAAAGGTGAGGTGACACCTCAACCAGTAGAGAAGAAACCAAAGGATAAATCAGAACCCAACTCCGGAGGGACTGGGTCAGCTGC
TAGTGTTAAAACAGCTGTTAAGAAAAAGATTATCAAGCGGGTGCTTAAGAGAAAGCTCACGCCGAACAAGGAAGCTACTGATGGAACTACTGGCACAGTT
ACGGAAGAGAAGAAGGTTGCTTCGGAAGCCAAGGAAACTGAAAATGCCGGGAATCAAGTTGAAGGCAATAGCGAGAATGCTAACAAAGTAACTCCTGCGG
CATCTTTGAAATCTCCAGAGAAGCAAGTGAGTGTGCCAAAGGCGAGTAAACCAGAAAGTAGCAAGGCCGTCAAGGAAGATAAGGACGAAAAGAAGCCTGA
AGGGAAGAATGGTACCGGAACCGGTTTGGAGGTTAAGGGTGAGAAGCCAAAGGGCGCTGAGAAGGATAATAATAATACTGGCAAGAAAGGGGCATCAGTG
GATGAACAGAAAGCAAAGGTTGAGAAAAAGGATAAGAATGGGAAAGATCAACAACCAAGGAGTAGGTCTCCTAAGGAAATGAAAGAGAAGAAAAAATCTG
AAGAGCCACCGAGACACCCTGGATTGATTTTGCAGACGAAATGCGACAAGAGAACTATGCTGTACTCATTATCTCTTTCTCTGGACTCGCTTTTGGATTA
CACGGAACAGGACACCGACGAATCGACAATTGAGCTTTCTTTATTTGCGGAATCCATGCATGAAATGCTTCAGTATCAGATGGGTAGTCGTCTGCTAGAT
TTCCTTCAGAAACTTCGCGTTAAATTTGTTGCAAAGAGGAATCAAAGGAAACGGCAGCTTGATGTGGAAAAGGAAGCAAAGGACAAAGATGACAAATCTT
CGTCGAAGCGCCAAAAAACCAGCGAGTTTCCTGTGAAGAAAGAATCCGGAACTGAAACGTCGGATGCAACAAACCAACCAGACGAACCGAAAACTACAAA
AGAGGATACTTTAGTTCAAAATGAGGTGAAGGCTCAAAATGACGTAGAAGATGAAGAAGAAGAAGATCCAGACGCAGATCCGGATGAGGATCCAGATGTG
GACCCGGATGAGGATCCAGATGAGGACCCGGATGTGGATCCAGATGAAGATCCGGACGAAGATCCTGAACCTGAAGGAGATGCAGAAATGGAGGATGCTG
AGCAAATTGTAGCCAGCGATCAGAAAGATAAAAAGGAAGCAGCAAAAGCCACTAGTGCCGACGGTGGAAAAACAGTTGTTGCGGCTGTAAATGAAGCTGG
GAAAGATAGTGAAATGAAAGATGCGGAGCCAAAAGTCAAGCAAGAACCAGATTCCTCCGAGAAAGTCAACTTGGAAGCCGAGAAGAAGAAGAAAGAGTCT
TCTTCAGCTGTTAAGGAGCCAGTGGTTAATACGGAGCTTCTGCAGGCGTTCAGGTTTTTCGATCGGAATCTAGCTGGGTTCATCAGGGTTGAGGATATGA
GGTTAATCATCCACACCCTGGGGAAGTTCCTCTCTCACAGAGACGTAAAGGATCTTGTTCAGAGTGCATTGCTGGAGAGCAACACCGGGAGGGACGACCG
GATCCTCTATAACAAGCTTGTGAGGATGAATCTGTAG
|
|||||||||||||||
|
AA sequence
|
>Lus10030445 pacid=23140692 polypeptide=Lus10030445 locus=Lus10030445.g ID=Lus10030445.BGIv1.0 annot-version=v1.0
MYSRGNGYGQQSYGGQSSYAQSLANAYSASSVGGTDSSSQHSLASRHPSLLSSSQEADLATGYRSHNSVSGHYGGQYASLYGSAGGVSATQQVPSASTKG
SGLSGLEGRRGYSSAVSDSPKFGSGDYVPSSTHTYGHKSDPLYSDKIADYTGIDRRHYGEQSSYTGRDLQTDPASRYTDSVGLSHQRQQTDRYDRLDQSS
ILRQEQLLKSQSIHSTSLDGGSRPLDYLAARGGASRHLNQDLASYGRRVDADPRGSSILNASSYNTTPASSILGAAPRRNVDDLLYSQSLTNPGYGVSLP
PGRDYGSGKAALQQGTSLDAEYRALHPRIEENRRDERAGYLREFELREDERRREFLREREKEREREREREKERDRDRERERERERRRDRERMLERREKER
ERKRAIEIRRERTPQRPSRDPRGSSVTKEGRSLRRDSISQDASHRRHSPVKDKRREYVCKVHGSCLVDLERDYLSLDKRYPRLFVSPELSKVVVNWPKEN
FKLSIHTPVSFEHDFVEEENALESKDLSSNLLGHHDLSQSASGRTVWNAKIILMSGVSKSAFEVLSSAKIADDRPPHICNILRFAVLKKDRAFMTIGGPW
DPIDGGYPSVDDSPLIQTALRYAKDVTQLDLGNCRQWNRFLEVQYNRFGKDGLFSHKEVTVLFVPDLSECLPSLESWRDQWLAHNKTVSDRERELSLKKE
RSAEKKEGSKGSDPSKDPKQGIKQEKTKASVSSGSNKKENASKDKTVSPKDAESKMDEGNLSKDVSKDEQGEPSSTPTSAGAKTGKKTGKKRIVKRIVKK
KAVAKPGGGVAAAGAATNNSSSGEKPAESEDKGLVEPEGKGEVTPQPVEKKPKDKSEPNSGGTGSAASVKTAVKKKIIKRVLKRKLTPNKEATDGTTGTV
TEEKKVASEAKETENAGNQVEGNSENANKVTPAASLKSPEKQVSVPKASKPESSKAVKEDKDEKKPEGKNGTGTGLEVKGEKPKGAEKDNNNTGKKGASV
DEQKAKVEKKDKNGKDQQPRSRSPKEMKEKKKSEEPPRHPGLILQTKCDKRTMLYSLSLSLDSLLDYTEQDTDESTIELSLFAESMHEMLQYQMGSRLLD
FLQKLRVKFVAKRNQRKRQLDVEKEAKDKDDKSSSKRQKTSEFPVKKESGTETSDATNQPDEPKTTKEDTLVQNEVKAQNDVEDEEEEDPDADPDEDPDV
DPDEDPDEDPDVDPDEDPDEDPEPEGDAEMEDAEQIVASDQKDKKEAAKATSADGGKTVVAAVNEAGKDSEMKDAEPKVKQEPDSSEKVNLEAEKKKKES
SSAVKEPVVNTELLQAFRFFDRNLAGFIRVEDMRLIIHTLGKFLSHRDVKDLVQSALLESNTGRDDRILYNKLVRMNL
|
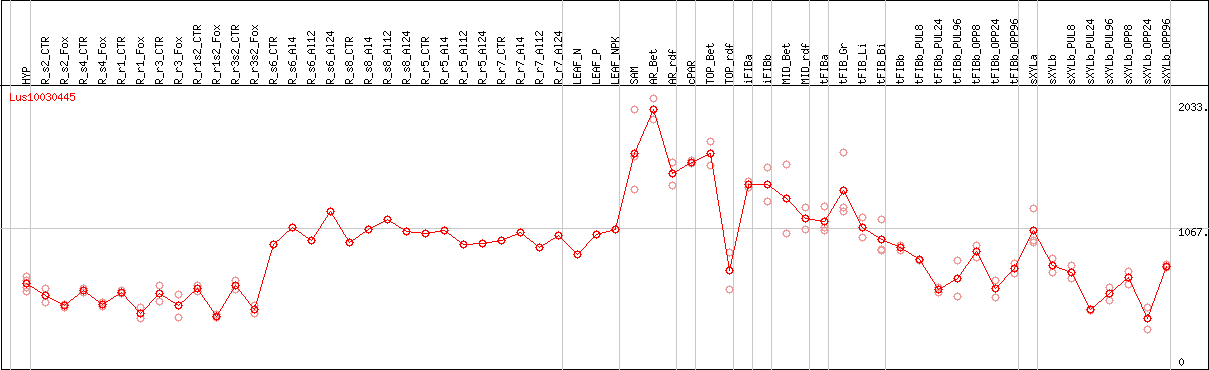
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030445 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
