External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69490 300 / 2e-102
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT1G61110 249 / 4e-82
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G04070 243 / 3e-79
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT3G15510 233 / 4e-75
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G52890 230 / 1e-74
NAC
ANAC019
NAC domain containing protein 19 (.1)
AT1G77450 226 / 1e-73
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT3G15500 227 / 3e-73
NAC
ATNAC3, ANAC055
NAC domain containing protein 55, NAC domain containing protein 3 (.1)
AT4G27410 224 / 3e-72
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT1G01720 220 / 6e-71
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G08790 219 / 7e-71
NAC
ATAF2, ANAC081
Arabidopsis NAC domain containing protein 81, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026617
523 / 0
AT1G69490 298 / 1e-101
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10037156
357 / 2e-124
AT1G69490 318 / 2e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10036773
351 / 3e-122
AT1G69490 318 / 1e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10007410
256 / 5e-84
AT3G15510 241 / 1e-78
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10036117
253 / 7e-83
AT3G15510 241 / 2e-78
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10003269
249 / 4e-81
AT3G04070 324 / 3e-109
NAC domain containing protein 47 (.1.2)
Lus10006547
248 / 1e-80
AT3G04070 337 / 4e-114
NAC domain containing protein 47 (.1.2)
Lus10043095
248 / 2e-80
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10032657
247 / 3e-80
AT3G15510 353 / 1e-120
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G089000
317 / 3e-109
AT1G69490 331 / 6e-115
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Potri.010G166200
303 / 1e-103
AT1G69490 305 / 1e-104
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Potri.004G038000
248 / 4e-81
AT1G61110 332 / 3e-113
NAC domain containing protein 25 (.1)
Potri.019G031400
248 / 6e-81
AT3G04070 332 / 1e-112
NAC domain containing protein 47 (.1.2)
Potri.011G046700
246 / 2e-80
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Potri.011G123500
245 / 7e-80
AT3G15510 323 / 4e-109
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Potri.001G404400
245 / 8e-80
AT3G15510 345 / 1e-117
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Potri.013G054000
243 / 5e-79
AT3G04070 338 / 9e-115
NAC domain containing protein 47 (.1.2)
Potri.006G129400
236 / 2e-76
AT1G61110 247 / 3e-80
NAC domain containing protein 25 (.1)
Potri.002G081000
229 / 1e-74
AT1G01720 416 / 5e-148
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10030446 pacid=23140762 polypeptide=Lus10030446 locus=Lus10030446.g ID=Lus10030446.BGIv1.0 annot-version=v1.0
ATGATGATGATTGCAGGAGAAGGAACAACAACAAGAACAAAGAACATGAACATGAACAATGATTATGTTCTTCCGCCGGGGTTCAGGTTCCATCCTACGG
ACGAGGAGCTGATCATGTGTTACCTCAGGAACCAGGCCACTTCCAGGCCTTGCCCTGTTTCCATCATCCCCGAAGTCGATATCTACAAGTTCGATCCTTG
GCAATTGCCTGAAAAGGCGGGGTTTGGTGAGAAAGAATGGTACTTTTTCAGTCCAAGAGACAGAAAGTACCCAAATGGAGTCAGACCCAACAGAGCCACT
GGCTCAGGCTACTGGAAGGCCACAGGAACTGACAAATCCATCTACAGTGGATCCCAATACGTCGGTGTCAAGAAAGCTCTCGTCTTTTACAAAGGGAAGC
CACCGAAGGGCATCAAAACCGACTGGATCATGCACGAATACCGCCTCTGCGATTCCAGGACCCGAACCGTCAAGGCCAGCGGGTCCATGAGACTGGATGA
ATGGGTCCTATGCAGGATATACAAGAAGAAGCCTAGTGTTAGCAAGCAATATCTGGAACAGGACATCAATGGAACAGAAGCTTTCAAAGAACAACAACAA
CAGATGTTGAAGCTTTCAAAGACATGTTCTTTGAGTGACTTGGATTCCTTGGGACCAATCTCACAGCTACTGGATGATCACAGTAACCCGAATTACCAGA
CAAACACGGGATTTCTTGTTCCCGAAAACACGGTGGGGAGTGCTCAGAACCTCGGTCAGCATCAAGTGATGAATGATCAGCAGCAGCCATTTCTTGTGAA
CCCACAAATGATGCATCAGTTTCAGTGA
AA sequence
>Lus10030446 pacid=23140762 polypeptide=Lus10030446 locus=Lus10030446.g ID=Lus10030446.BGIv1.0 annot-version=v1.0
MMMIAGEGTTTRTKNMNMNNDYVLPPGFRFHPTDEELIMCYLRNQATSRPCPVSIIPEVDIYKFDPWQLPEKAGFGEKEWYFFSPRDRKYPNGVRPNRAT
GSGYWKATGTDKSIYSGSQYVGVKKALVFYKGKPPKGIKTDWIMHEYRLCDSRTRTVKASGSMRLDEWVLCRIYKKKPSVSKQYLEQDINGTEAFKEQQQ
QMLKLSKTCSLSDLDSLGPISQLLDDHSNPNYQTNTGFLVPENTVGSAQNLGQHQVMNDQQQPFLVNPQMMHQFQ
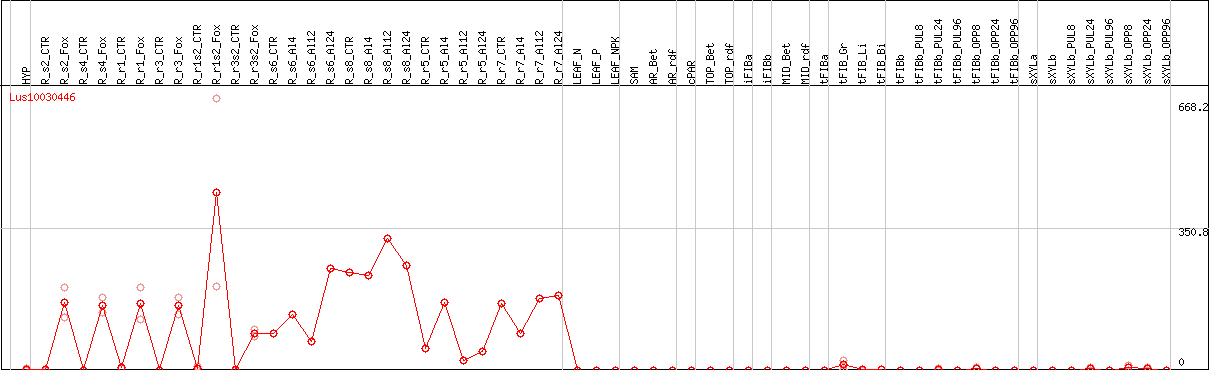
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030446 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.