External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G03050 360 / 1e-116
RHD7, ATCSLD3, KJK, CSLD3
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
AT5G16910 352 / 9e-114
ATCSLD2
cellulose-synthase like D2 (.1)
AT4G38190 278 / 4e-86
ATCSLD4
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
AT1G02730 174 / 4e-49
SOS6, ATCSLD5
SALT OVERLY SENSITIVE 6, CELLULOSE SYNTHASE LIKE D5, cellulose synthase-like D5 (.1)
AT2G33100 78 / 4e-16
ATCSLD1
CELLULOSE-SYNTHASE LIKE D1, cellulose synthase-like D1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026610
456 / 5e-154
AT3G03050 1028 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10026609
436 / 1e-146
AT3G03050 1326 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10038008
403 / 2e-133
AT3G03050 1818 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10030455
363 / 3e-119
AT3G03050 1258 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10009248
320 / 1e-101
AT3G03050 1929 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10013851
280 / 6e-87
AT4G38190 1876 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Lus10026568
280 / 7e-87
AT4G38190 1868 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Lus10022982
250 / 6e-76
AT4G38190 1823 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Lus10001619
248 / 1e-75
AT4G38190 961 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G082200
440 / 4e-147
AT3G03050 1987 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Potri.019G049700
425 / 3e-141
AT3G03050 2020 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Potri.009G170000
281 / 2e-87
AT4G38190 1873 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Potri.004G208800
273 / 2e-84
AT4G38190 1835 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Potri.003G097100
224 / 9e-67
AT3G03050 1610 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Potri.001G136200
214 / 4e-63
AT3G03050 1640 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Potri.014G125100
181 / 1e-51
AT1G02730 1820 / 0.0
SALT OVERLY SENSITIVE 6, CELLULOSE SYNTHASE LIKE D5, cellulose synthase-like D5 (.1)
Potri.002G200300
168 / 3e-47
AT1G02730 1813 / 0.0
SALT OVERLY SENSITIVE 6, CELLULOSE SYNTHASE LIKE D5, cellulose synthase-like D5 (.1)
Potri.003G177800
77 / 6e-16
AT2G33100 1569 / 0.0
CELLULOSE-SYNTHASE LIKE D1, cellulose synthase-like D1 (.1)
Potri.001G050200
75 / 6e-15
AT2G33100 1576 / 0.0
CELLULOSE-SYNTHASE LIKE D1, cellulose synthase-like D1 (.1)
PFAM info
Representative CDS sequence
>Lus10030454 pacid=23140614 polypeptide=Lus10030454 locus=Lus10030454.g ID=Lus10030454.BGIv1.0 annot-version=v1.0
ATGGCTTCAAAGTCGTTCAAAGCCAGTAGATCAAACACGTCGATGAGCTCCGACGCGGCGGACGCGCAGCAGCAGCAGCAGCGGGGGACAGTCCCCCCGA
CGGTGACGTTCGGCCGGAGGACATCTTCGGGGAGGTACGTGAGCTACTCGAGGGAGGATCTAGACAGTGAGATAGGTAGCAGCGACTTCATGAACTACAC
AGTCCACATCCCGCCGACTCCGGACAACCAGCCGATGGATCCTTCGATTTCTCAGAAGGTCGAGGAGCAGTATGTTTCGAACTCGCTCTTCACTGGCGGG
TTCAACAGTGTGACTCGAGCTCACTTGATGGATAAGGTTATTGAGTCGGAGGCGAATCATCCTCAGATGGCTGGTTCTAAGGGGTCTTGCTGTGCGATCC
CTGGATGTGATGCTAACGTGATGAGTGACGGGCGTGGATTGGATATTCTGCCATGTGAGTGTGATTTTAAGATTTGTAGGGATTGTTATATTGATGCTGT
GAAGACTGGGGGTGGAATCTGTCCCGGGTGTAAGGAGAGTTATAAGAACACTGATCTTGATGAGGTTGCTGTTGATAAGTCGAGTCCTCTTCCTCTGCCT
GCTCCTGGTGGTGGTACTATGTCGAAGATGGAGAGGCGGCTTTCGTTGATGAAGTCTACGAAGTCTGTTCTTATGAGGAGTCATACTGGTGAATTTGATC
ACAATAGGTGGTTGTTTGAGACTAAGGGGACTTATGGTTATGGTAATGCCATTTGGACGAATGATGGTGGGTTTGGTGATGGTAAGGGTATTACATTTCC
ATGA
AA sequence
>Lus10030454 pacid=23140614 polypeptide=Lus10030454 locus=Lus10030454.g ID=Lus10030454.BGIv1.0 annot-version=v1.0
MASKSFKASRSNTSMSSDAADAQQQQQRGTVPPTVTFGRRTSSGRYVSYSREDLDSEIGSSDFMNYTVHIPPTPDNQPMDPSISQKVEEQYVSNSLFTGG
FNSVTRAHLMDKVIESEANHPQMAGSKGSCCAIPGCDANVMSDGRGLDILPCECDFKICRDCYIDAVKTGGGICPGCKESYKNTDLDEVAVDKSSPLPLP
APGGGTMSKMERRLSLMKSTKSVLMRSHTGEFDHNRWLFETKGTYGYGNAIWTNDGGFGDGKGITFP
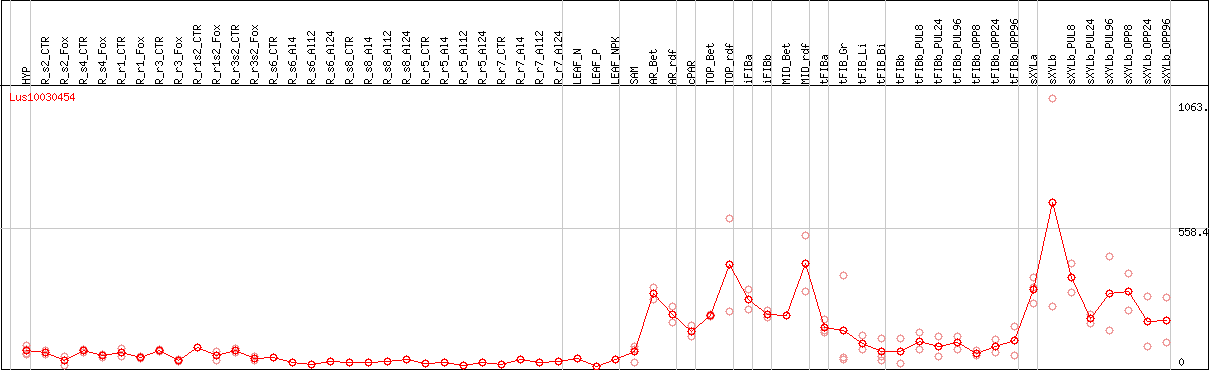
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030454 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.