External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G25810 394 / 2e-139
XTH23, XTR6
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
AT5G57550 392 / 2e-138
XTR3, XTH25, EXGT-A5
xyloglucan endotransglycosylase 3, xyloglucan endotransglucosylase/hydrolase 25 (.1)
AT5G57560 385 / 1e-135
XTH22, TCH4
xyloglucan endotransglucosylase/hydrolase 22, Touch 4, Xyloglucan endotransglucosylase/hydrolase family protein (.1)
AT4G30270 358 / 4e-125
XTH24, SEN4, BRU1, MERI-5, MERI5B
SENESCENCE 4, meristem-5, MERISTEM 5, xyloglucan endotransglucosylase/hydrolase 24 (.1)
AT5G57530 348 / 5e-121
AtXTH12, XTH12
xyloglucan endotransglucosylase/hydrolase 12 (.1)
AT5G57540 347 / 2e-120
AtXTH13, XTH13
xyloglucan endotransglucosylase/hydrolase 13 (.1)
AT2G18800 347 / 2e-120
ATXTH21, XTH21, XTR17
xyloglucan endotransglucosylase/hydrolase 21 (.1)
AT4G25820 345 / 9e-120
ATXTH14, XTR9, XTH14
xyloglucan endotransglycosylase 9, xyloglucan endotransglucosylase/hydrolase 14 (.1)
AT3G23730 341 / 2e-118
XTH16
xyloglucan endotransglucosylase/hydrolase 16 (.1)
AT4G14130 339 / 1e-117
XTR7, XTH15
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012834
400 / 1e-141
AT5G57550 292 / 5e-99
xyloglucan endotransglycosylase 3, xyloglucan endotransglucosylase/hydrolase 25 (.1)
Lus10028947
344 / 1e-119
AT4G14130 437 / 2e-156
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Lus10000678
344 / 2e-119
AT3G23730 410 / 6e-146
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10010938
342 / 6e-119
AT4G14130 386 / 3e-136
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Lus10021638
342 / 8e-119
AT3G23730 413 / 7e-147
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10031393
341 / 3e-118
AT3G23730 388 / 8e-137
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10010939
340 / 9e-118
AT3G23730 384 / 2e-135
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10031392
338 / 5e-117
AT3G23730 384 / 3e-135
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10007349
325 / 5e-112
AT3G23730 394 / 4e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G095100
394 / 2e-139
AT4G25810 420 / 1e-149
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.013G005700
393 / 5e-138
AT4G25810 431 / 5e-153
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G095200
390 / 1e-137
AT4G25810 414 / 3e-147
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G094800
389 / 3e-137
AT4G25810 413 / 1e-146
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.005G007200
380 / 1e-133
AT4G25810 394 / 4e-139
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G094900
367 / 1e-128
AT4G25810 410 / 8e-146
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.006G071200
367 / 2e-128
AT4G25810 379 / 3e-133
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.014G146100
352 / 1e-122
AT3G23730 451 / 1e-161
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.005G201200
349 / 3e-121
AT4G14130 405 / 2e-143
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Potri.005G201250
348 / 6e-121
AT4G14130 395 / 2e-139
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0004
Concanavalin
PF00722
Glyco_hydro_16
Glycosyl hydrolases family 16
CL0004
Concanavalin
PF06955
XET_C
Xyloglucan endo-transglycosylase (XET) C-terminus
Representative CDS sequence
>Lus10030484 pacid=23140639 polypeptide=Lus10030484 locus=Lus10030484.g ID=Lus10030484.BGIv1.0 annot-version=v1.0
ATGGCGACACCCACCTCCAACACCCCTTTCCTACTCCTCTCAATCACCCTCATCATCACCACCACCATCGCGGCTCAGACCAATCTGGACCGCGAATTCG
ACCTCACGTGGGGAGAAGGCCGCGGCAAGATCGTTGCCGACGGCCAGTTGCTCACCCTGACCCTCGATCACACTTCGGGGTCAGGGTTCCAGTCCAAGAA
CAAGTACCTGTACGGGAAGCTCGACATGCAGCTGAAGCTAGTCCCCGGCAACTCTGCTGGCACCGTTACTGCTTACTTCTTGAAGTCTGATGGAGATGCG
TGGGACGAGATTGATTATGAGTTCTTGGGGAACTTGAGTGGCGATCCTTACACGGTCCATACGAACGTGTTCACGAACGGGAAAGGCGATCGGGAGCAGC
AGTTTCGTCTTTGGTTCGATCCGACGACTGATTTTCATACTTACTCGATCCTTTGGAACCCCAAGAGCATCGTTTTCACCATCGACGGCACACCGATCAG
AGAATTCAAGAACCTGGAATCCGTCGGAGTCCCCTTCCCAACATCACAACCCATGAGGCTCTACTCGAGCCTATGGAGCGCCGACGACTGGGCCACCAGG
GGAGGGACCCTGAAGACCGACTGGTCCCAGGCTCCGTTCTCGGCCTCCTACAAAGACTTCAAAGCAGGGGAGGCCTGCATAGTGAATAACGATGGAGCGA
ACCCAACTTGCTCGCCTGCCGGTAGCACGTCTTGGCTTAACGAGCAGCTGAACTCGGTGAACGAGCAGCGGATGAGGTCGGTCCAGAGCCATTACATGAT
CTACAACTACTGTACTGATTCGGAGAGGTTTCCTCAGGGGTTCCCTAAAGAGTGTACTGCAGAATGA
AA sequence
>Lus10030484 pacid=23140639 polypeptide=Lus10030484 locus=Lus10030484.g ID=Lus10030484.BGIv1.0 annot-version=v1.0
MATPTSNTPFLLLSITLIITTTIAAQTNLDREFDLTWGEGRGKIVADGQLLTLTLDHTSGSGFQSKNKYLYGKLDMQLKLVPGNSAGTVTAYFLKSDGDA
WDEIDYEFLGNLSGDPYTVHTNVFTNGKGDREQQFRLWFDPTTDFHTYSILWNPKSIVFTIDGTPIREFKNLESVGVPFPTSQPMRLYSSLWSADDWATR
GGTLKTDWSQAPFSASYKDFKAGEACIVNNDGANPTCSPAGSTSWLNEQLNSVNEQRMRSVQSHYMIYNYCTDSERFPQGFPKECTAE
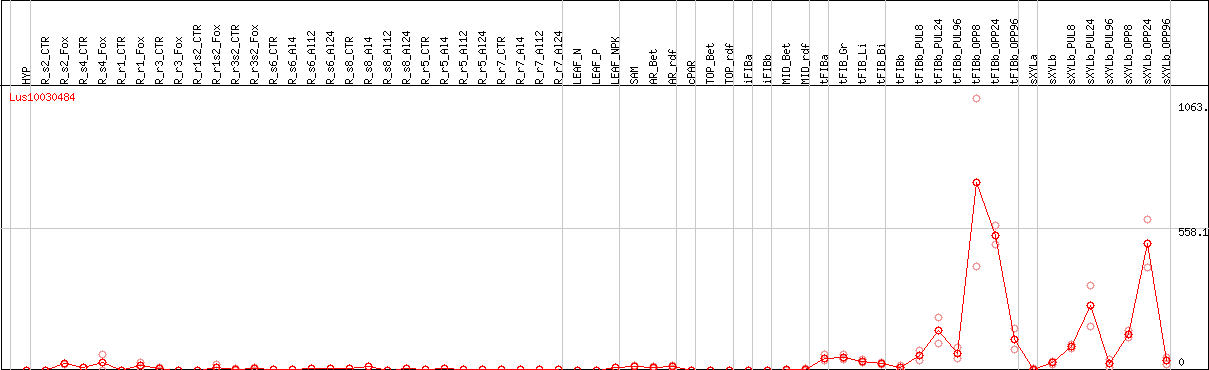
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030484 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.