External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G21200 369 / 5e-128
ATGA2OX8
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
AT1G50960 276 / 2e-91
ATGA2OX7
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 7, gibberellin 2-oxidase 7 (.1)
AT5G58660 170 / 4e-50
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G05600 168 / 4e-49
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G19010 164 / 6e-48
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.3)
AT4G10490 157 / 6e-45
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10500 153 / 2e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G55970 153 / 2e-43
ATJRG21
jasmonate-regulated gene 21 (.1)
AT1G77330 151 / 2e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G11180 154 / 3e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012912
632 / 0
AT4G21200 368 / 2e-127
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10029081
561 / 0
AT4G21200 365 / 3e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10013069
540 / 0
AT4G21200 366 / 5e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10007640
353 / 1e-121
AT4G21200 429 / 1e-151
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10006759
350 / 3e-120
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10018372
346 / 9e-119
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10004808
176 / 2e-52
AT5G05600 457 / 5e-162
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005037
172 / 1e-50
AT3G11180 471 / 3e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10030934
169 / 1e-49
AT1G17020 337 / 2e-114
senescence-related gene 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G096800
510 / 0
AT4G21200 362 / 8e-125
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.008G145300
489 / 6e-175
AT4G21200 354 / 1e-121
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.004G022800
372 / 4e-129
AT4G21200 441 / 7e-156
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.011G026700
361 / 9e-125
AT4G21200 444 / 1e-157
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.001G418200
339 / 3e-116
AT4G21200 369 / 8e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.011G134000
302 / 1e-101
AT4G21200 340 / 1e-116
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.011G134100
291 / 2e-97
AT4G21200 324 / 2e-110
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.001G278200
214 / 2e-67
AT5G58660 312 / 2e-105
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.004G146000
179 / 2e-53
AT3G19000 357 / 3e-122
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.016G117100
178 / 6e-53
AT5G05600 491 / 4e-175
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10030567 pacid=23140785 polypeptide=Lus10030567 locus=Lus10030567.g ID=Lus10030567.BGIv1.0 annot-version=v1.0
ATGATGCTTACGACCAACCCTCCTCTCATTCACGGTTACGAGAAGGTCTCGCGGCTACCTCCATTAGAACACCAGCACGAGCTTCTCCACTCATCCTCTG
TAATGGAGGAATGCCAGCTACCGCTGATAGACCTCAGCGGACTCAGCAGCTGCGACGAAAGGGAGCGAGCTAATTGCGCTGCTGCTATATGCAGGGCTTC
GTCAGAATGGGGATTCTTTCAGGTGGTTAACCATGGGATCGACCTCGATCTTGTTAGAAAGATGAGGAAAGAACAGGTGGATTTGTTCCAACTGCCATTT
GATAAGAAAGCGAGTAGCGGGTTGCTGAACAACTCGTACCGTTGGGGGACTCCAACGGCTACTTGTCCGAAGCAGTTCTCGTGGTCCGAAGCTTTCCATG
TTCCTTTGTCGATGATTTCGCAGGAGGACTGCTACCAAGAGTTCACTTCTTTGAGGGAAGTGATGGTGGAATTTGCAGCAGCAATGTCGAAGCTGGCGGG
ATCGCTTGCGGGGGTCTTGGTAGAGAACTTGGGGCATTCAAGTGGGGATTTTGAGGACATGTGTGAAGAAAGAAGTTGCTTTCTTAGGCTTAATAGATAT
CCAGCTTGTCCGGTGTCCCCGGAGATTTTCGGATTAGTGCCACACACTGACAGTGATTTCCTCACGATTCTGTCGCAAGATGAAGTAGGGGGACTTCAGC
TCATGAAAGATTCCAGATGGGTTGCTGTAAAACCTAATCCAGATGCATTCATAGTGAACATTGGTGATCTTTTTCAGGCATGGAGCAACGACATCTATAA
AAGTGTGGAACACAAGGTTATGGCGAATGCTAAGACAGAACGATACTCGATCGCCTATTTCTTGTGCCCTTCGTACGAGACATCAATAGGGAGTTGCGGA
GAACCATCCAAGTACAGGAGATTCACTTTCGGAGAGTTCAGGAGCCAAATTCAAGCAGAGGTCAAGAACATTGGCTATAAAATCGGCCTGCCAAGCTTTC
AGTTATAA
AA sequence
>Lus10030567 pacid=23140785 polypeptide=Lus10030567 locus=Lus10030567.g ID=Lus10030567.BGIv1.0 annot-version=v1.0
MMLTTNPPLIHGYEKVSRLPPLEHQHELLHSSSVMEECQLPLIDLSGLSSCDERERANCAAAICRASSEWGFFQVVNHGIDLDLVRKMRKEQVDLFQLPF
DKKASSGLLNNSYRWGTPTATCPKQFSWSEAFHVPLSMISQEDCYQEFTSLREVMVEFAAAMSKLAGSLAGVLVENLGHSSGDFEDMCEERSCFLRLNRY
PACPVSPEIFGLVPHTDSDFLTILSQDEVGGLQLMKDSRWVAVKPNPDAFIVNIGDLFQAWSNDIYKSVEHKVMANAKTERYSIAYFLCPSYETSIGSCG
EPSKYRRFTFGEFRSQIQAEVKNIGYKIGLPSFQL
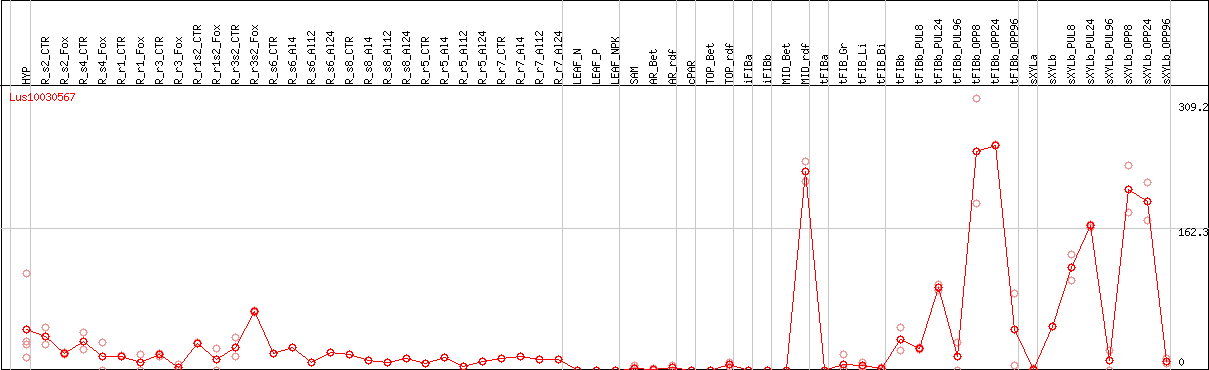
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030567 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.