External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G76680 258 / 1e-84
OPR1, ATOPR1
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 1, 12-oxophytodienoate reductase 1 (.1.2)
AT1G76690 255 / 9e-84
OPR2, ATOPR2
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
AT1G18020 230 / 4e-75
FMN-linked oxidoreductases superfamily protein (.1)
AT1G17990 230 / 4e-75
FMN-linked oxidoreductases superfamily protein (.1.2)
AT1G09400 224 / 3e-72
FMN-linked oxidoreductases superfamily protein (.1)
AT2G06050 194 / 2e-59
AtOPR3, DDE1, OPR3, OPDA-REDUCTASE
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013218
347 / 7e-120
AT1G76690 553 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Lus10001276
254 / 3e-83
AT1G76680 597 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 1, 12-oxophytodienoate reductase 1 (.1.2)
Lus10009473
247 / 6e-78
AT1G76690 568 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Lus10001275
233 / 8e-75
AT1G76680 544 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 1, 12-oxophytodienoate reductase 1 (.1.2)
Lus10001274
201 / 6e-63
AT1G76690 526 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Lus10028663
197 / 1e-60
AT2G06050 576 / 0.0
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
Lus10000966
192 / 8e-59
AT2G06050 576 / 0.0
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G102700
269 / 6e-89
AT1G76690 629 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Potri.013G102750
251 / 2e-82
AT1G76690 580 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Potri.003G004200
251 / 3e-82
AT1G76680 558 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 1, 12-oxophytodienoate reductase 1 (.1.2)
Potri.013G102800
250 / 9e-82
AT1G76690 576 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Potri.013G103000
249 / 3e-81
AT1G76690 568 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Potri.003G004600
247 / 1e-80
AT1G76680 559 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 1, 12-oxophytodienoate reductase 1 (.1.2)
Potri.013G102900
245 / 6e-80
AT1G76690 571 / 0.0
ARABIDOPSIS 12-OXOPHYTODIENOATE REDUCTASE 2, 12-oxophytodienoate reductase 2 (.1)
Potri.006G142800
189 / 2e-57
AT2G06050 603 / 0.0
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
Potri.004G212100
186 / 1e-56
AT2G06050 536 / 0.0
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
Potri.018G065600
185 / 4e-56
AT2G06050 599 / 0.0
DELAYED DEHISCENCE 1, oxophytodienoate-reductase 3 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0036
TIM_barrel
PF00724
Oxidored_FMN
NADH:flavin oxidoreductase / NADH oxidase family
Representative CDS sequence
>Lus10030737 pacid=23155406 polypeptide=Lus10030737 locus=Lus10030737.g ID=Lus10030737.BGIv1.0 annot-version=v1.0
ATGGCGGAATCAGCTCAGCCTCTGCTGACTCCTTACAAAATGGGCAACTTCAGTCTCTCTCACAGGGTAGTTTTGGCACCGCTGACCAGGATGAGATCGT
ACGGGAAGATCCCTCAGCCACACGCCGTCTTGTACTACTCCCAAAGAACGCCACCTGGTGGTCTCGTGATTGCTGAAGCCACCGCCATTTCCAACACTTC
TATCGGTTACTCAAATGTTCCTGGAATCTGGACTGAAGAGCAAACCAATTGTGCTGTGCATGAGAAAGGAGGCATCTTCTTTTGCCAGATTTGGCATTCT
GGAAGAGTCTCTGATTACTGCTTTCAACCAAATGGACAGCCTCCCGTCTCCTGTACGGATAAGCCGATTACAGCTGAGCCCCAGATCGATGGCAGTAAAG
CTCCTGCATATCCTGCTCCCTGTAGACTCGAAACAGAGGAGATTCCTCACATTGTAAACGACTTTGCGGTTGCAGCAAGGAATGCCATGGAAGCTGGTTT
TGATGGCGTGGAGATCCACGGAGCCAATGGATATATCGTTGACCAGTTTGATGCGGTGGAGAGCGAGCACAGCTTGTTGCCCACGAGGAAGGCCTTTGGA
GGTACGTTTATGATCGCCGGAGGATACACTCAATGCAACGGGAATGAAGTGGTTGCCGATGGAGGTGCGGATTTAGTTGCATTCGGGCGTTTGTTCTTGG
CGAATCCAGACATGCCCAGAAGATTCGAGCTGAATAAGTATAACAGGGGCACTTTCTATACCGATGACCCTGTTGTTGGATGCTCCGACTATCCATTTCT
CGACCGAGTAAAAGTCTAG
AA sequence
>Lus10030737 pacid=23155406 polypeptide=Lus10030737 locus=Lus10030737.g ID=Lus10030737.BGIv1.0 annot-version=v1.0
MAESAQPLLTPYKMGNFSLSHRVVLAPLTRMRSYGKIPQPHAVLYYSQRTPPGGLVIAEATAISNTSIGYSNVPGIWTEEQTNCAVHEKGGIFFCQIWHS
GRVSDYCFQPNGQPPVSCTDKPITAEPQIDGSKAPAYPAPCRLETEEIPHIVNDFAVAARNAMEAGFDGVEIHGANGYIVDQFDAVESEHSLLPTRKAFG
GTFMIAGGYTQCNGNEVVADGGADLVAFGRLFLANPDMPRRFELNKYNRGTFYTDDPVVGCSDYPFLDRVKV
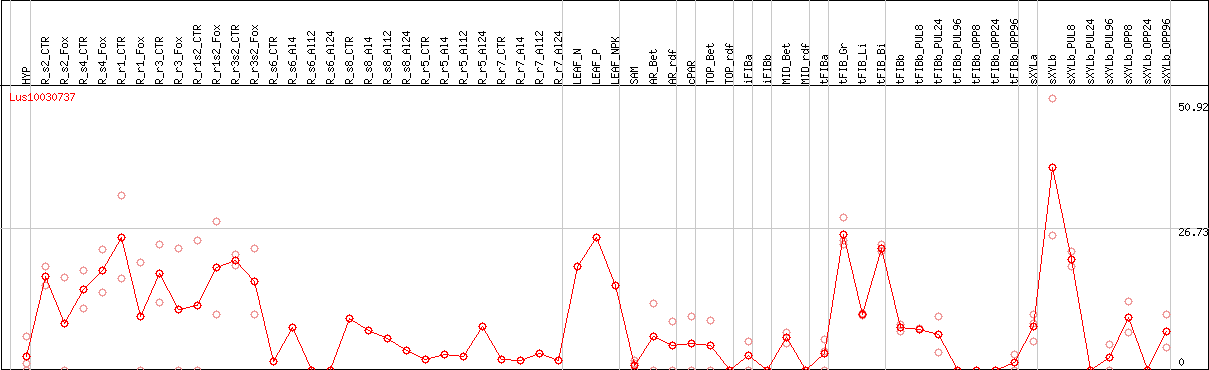
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030737 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.