External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14740 113 / 6e-30
Protein of unknown function (DUF1423) (.1)
AT3G63500 74 / 4e-16
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
AT5G48160 55 / 1e-09
OBE2
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
AT3G07780 48 / 5e-07
OBE1
OBERON1, Protein of unknown function (DUF1423) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013248
202 / 7e-65
AT1G14740 466 / 7e-160
Protein of unknown function (DUF1423) (.1)
Lus10019718
98 / 1e-24
AT3G63500 665 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10016399
95 / 2e-23
AT3G63500 717 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10012327
60 / 4e-11
AT5G48160 878 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10006372
58 / 2e-10
AT5G48160 871 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G138100
143 / 2e-40
AT1G14740 729 / 0.0
Protein of unknown function (DUF1423) (.1)
Potri.010G102600
135 / 9e-38
AT1G14740 773 / 0.0
Protein of unknown function (DUF1423) (.1)
Potri.001G264100
96 / 9e-24
AT3G63500 660 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.009G059000
91 / 4e-22
AT3G63500 693 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.002G221500
61 / 1e-11
AT5G48160 811 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.014G164800
59 / 4e-11
AT5G48160 859 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0390
zf-FYVE-PHD
PF07227
PHD_Oberon
PHD - plant homeodomain finger protein
Representative CDS sequence
>Lus10030769 pacid=23155280 polypeptide=Lus10030769 locus=Lus10030769.g ID=Lus10030769.BGIv1.0 annot-version=v1.0
ATGGAAGACGAGTACCAACACTCTCTAGCAGCAAGGAAAGAAGGATTGGAGAGCATTGAAATCATCGTGCGGATCAAACAAGCCGAGGCAAGGATGTTCC
AGAACAAGGCAGACGAGGCGAGGAGGGAAGCTGAAGGGCTGAAGCGGATGATCCAGGTGAGGACCGATAAGTACGAGGAAGAGTACTCGCAGAAGTTGTC
GAAGCTGTGCTTGCAGGAGACTGAGGAAAGGAGGAGGAAGAAGCTGGAGGAGCTGAAGAAGTTCGAGACCTCTCACTGCGATTACTTCAACATGAAGATG
AGGATGCAGGGTGAGATTGCTGGATTGCTAGAGAGAATGGAGGCCACTAAACAACAGAGGGGTTTAGCTGCAGTGAAAAGGGTTCATTCTCATGTGGGTT
CTTTATTGTTTCTTGGTTGA
AA sequence
>Lus10030769 pacid=23155280 polypeptide=Lus10030769 locus=Lus10030769.g ID=Lus10030769.BGIv1.0 annot-version=v1.0
MEDEYQHSLAARKEGLESIEIIVRIKQAEARMFQNKADEARREAEGLKRMIQVRTDKYEEEYSQKLSKLCLQETEERRRKKLEELKKFETSHCDYFNMKM
RMQGEIAGLLERMEATKQQRGLAAVKRVHSHVGSLLFLG
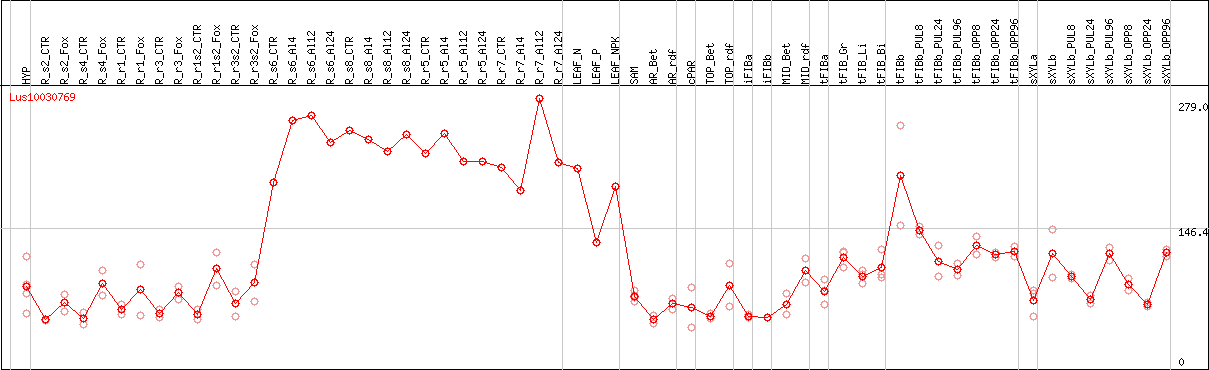
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030769 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.