External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14740 393 / 8e-133
Protein of unknown function (DUF1423) (.1)
AT3G63500 260 / 2e-80
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
AT5G48160 226 / 7e-70
OBE2
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
AT3G07780 225 / 1e-69
OBE1
OBERON1, Protein of unknown function (DUF1423) (.1)
AT5G57380 52 / 2e-07
VIN3
VERNALIZATION INSENSITIVE 3, Fibronectin type III domain-containing protein (.1)
AT3G24440 41 / 0.0006
VRN5, VIL1
VERNALIZATION 5, VIN3-LIKE 1, Fibronectin type III domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013248
477 / 2e-170
AT1G14740 466 / 7e-160
Protein of unknown function (DUF1423) (.1)
Lus10016399
266 / 2e-81
AT3G63500 717 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10019718
265 / 2e-80
AT3G63500 665 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10006372
231 / 1e-71
AT5G48160 871 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10012327
230 / 3e-71
AT5G48160 878 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Lus10028170
47 / 1e-05
AT5G57380 239 / 8e-72
VERNALIZATION INSENSITIVE 3, Fibronectin type III domain-containing protein (.1)
Lus10042869
45 / 3e-05
AT5G57380 237 / 5e-71
VERNALIZATION INSENSITIVE 3, Fibronectin type III domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G138100
471 / 3e-162
AT1G14740 729 / 0.0
Protein of unknown function (DUF1423) (.1)
Potri.010G102600
464 / 6e-159
AT1G14740 773 / 0.0
Protein of unknown function (DUF1423) (.1)
Potri.001G264100
274 / 4e-84
AT3G63500 660 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.009G059000
273 / 9e-84
AT3G63500 693 / 0.0
Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.014G164800
229 / 5e-71
AT5G48160 859 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.002G221500
228 / 1e-70
AT5G48160 811 / 0.0
OBERON2, Protein of unknown function (DUF1423) (.1), Protein of unknown function (DUF1423) (.2)
Potri.018G076500
42 / 0.0004
AT3G24440 531 / 0.0
VERNALIZATION 5, VIN3-LIKE 1, Fibronectin type III domain-containing protein (.1)
Potri.018G091500
42 / 0.0005
AT4G30200 612 / 0.0
VIN3-Like 2, vernalization5/VIN3-like 1, vernalization5/VIN3-like (.1.2.3.4)
Potri.006G158676
41 / 0.0008
AT3G24440 523 / 9e-179
VERNALIZATION 5, VIN3-LIKE 1, Fibronectin type III domain-containing protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0390
zf-FYVE-PHD
PF07227
PHD_Oberon
PHD - plant homeodomain finger protein
Representative CDS sequence
>Lus10030770 pacid=23155222 polypeptide=Lus10030770 locus=Lus10030770.g ID=Lus10030770.BGIv1.0 annot-version=v1.0
ATGGCGCAGATTGTTCAGGAACTTACTCCCGAGATGTCGAATTCGATCAAGGAGTACGTAAATCGTCTGGTTTTTACTGCTGATAATCGAGACGAGCTGG
TAGGGCTTCAGAGCCGATTGGAGAGGAGGTCTGATTTAATAAAGGAGTCTCTTTTAAAGTGTCACAGAGACCAGCTGGAGATTCTCGTTGCGGTGAAGAT
GGGGCTGGCGAGTTTTGTCTCCGGGAAAGCTCGTCTCCCTACCGAGGAGCTCATCGAGGTGTTTTTGTTCACGAGGTGTAGGAACGTGAACTGCAAGAGT
ATATTACCCGTTGAGGACTGCGACTGTAAATTCTGCTCCGCAAATAATGGGTTTTGTAGCTCGTGCATGTGTCCCGTATGTATGAAGTTCGATTGCGCTA
GCAATACGTGCAGTTGGGTCGGATGTGATGTTTGTTCGCATTGGTGTCATGCTGCTTGCGGGATACAGAAGAATCTCATCAGGCCGGGACCTACTTTGAA
AGGTCCCCGGGGGACAAGCGAGATGCAGTTTCATTGTATCGCCTGCGATCATGCTTCTGAGATGTTTGGGTTCGTTAAAGATGTGTATACTTGCTGTGCT
AAAGATTGGGGTTATGAGACTCTGTTGAAGGAGCTTGCCTGTGTGAGGAAGATTTTCAAAGGGAGTGATGATTTTAAAGGGAAAGAGCTTCATACCAAGG
CTGATGGGTTGATCTCCAAGCTTGAAGATAAACTTATATCTCCTTCAGATGCTTGCCACATCATTCTCATGTTCTTTGACTATGCAGATCGCCTACCGGA
TTTCCCTACCGGCCCTTCTGCCAAAGAAATGATGCCTCGTAACACCGAACCCATCACTCGAAAAGAAGCAACTCCCCTTCCATAA
AA sequence
>Lus10030770 pacid=23155222 polypeptide=Lus10030770 locus=Lus10030770.g ID=Lus10030770.BGIv1.0 annot-version=v1.0
MAQIVQELTPEMSNSIKEYVNRLVFTADNRDELVGLQSRLERRSDLIKESLLKCHRDQLEILVAVKMGLASFVSGKARLPTEELIEVFLFTRCRNVNCKS
ILPVEDCDCKFCSANNGFCSSCMCPVCMKFDCASNTCSWVGCDVCSHWCHAACGIQKNLIRPGPTLKGPRGTSEMQFHCIACDHASEMFGFVKDVYTCCA
KDWGYETLLKELACVRKIFKGSDDFKGKELHTKADGLISKLEDKLISPSDACHIILMFFDYADRLPDFPTGPSAKEMMPRNTEPITRKEATPLP
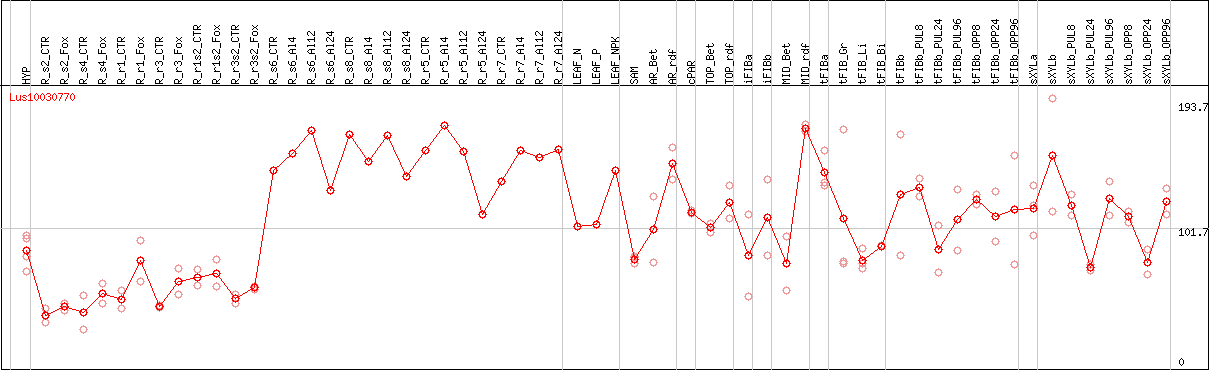
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030770 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.