Lus10030817 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030817 pacid=23155421 polypeptide=Lus10030817 locus=Lus10030817.g ID=Lus10030817.BGIv1.0 annot-version=v1.0
ATGGTGGGAGGAGGTGCTGCCACTGGAGGGGGAGGAGGAGCTGGAGCAGGAATTAGTGCCCAAAAGCTGACTACAAATGATGCATTGGCTTATCTCAAGG
CTGTCAAGGACATATTCCAGGACAAACGAGAAAAATACGATGAATTTCTCGAGGTTATGAAAGATTTCAAAGCTCAAAGAATCGACACTGCCGGTGTCAT
AGGGAGGGTAAAGGACTTGTTTAAAGGGCATCGTGACCTGATTTTGGGTTTTAATACCTTCTTGCCAAAGGGCTATGAGATCACTCTACCGCTGGAGGAT
GAACAGCCTCCGCAGAAGAAACCAGTTGAATTTGAAGAAGCCATTAATTTTGTGAACAAAATTAAGAAAAGGTTTCAGGGTGATGATCATGTTTACAAAG
CATTTTTAGATATTTTGAACATGTACAGACAGGAGAATAAATCCATCACAGAGGTTTATCAGGAGGTGGCTGCTCTGTTCAAAGACCACCATGATTTGCT
TGGAGAGTTTACCCACTTTTTACCTGATAATTCAGCTACAGCTTCTACTATGTTTGCTTCTTCAAATAGGAATTCTATTCTCCGTGATAGGAGCTCCCCG
ATGCCTTCAAATCGTCAAATGCATGTTGACAAGAAAGAACGGATTCATCCAGATCGTGATTTCAGTGTTGATCGTCCTGATCCAGACCATGATAGAGCTC
TGATGAGAGCTGATAAAGAGCAAAGGAGACGAGTCGAAAAAGAAAAGGAGAGAAGAGAAGATAGAGATAGGAGGGAACGTGATGATAGAGACTATGAGCA
TGATGCTAGTCATGACTATCGGTTTCCACTCAAGGGGAAAGTTGGTCGTAAGTTGGAAGATTCCAGTGCTGAGTGTCAAGTTGTTGGGGGGGATGATAAC
TATGGAATGCATCCTGTTTCATCCAGCTATGATGATAAAAGTGCAGTTAAAAGTGCATTAAGTCTTGAACTCGGTTACTGTGATAAAGTGAAGGAGAAGT
TGGTGGAGAAGACTGGAGCTTACCAGGAATTTTTGAAGTGTCTCAATCTCTACACAAAAGACATAATCGATCGAAAAGAATTACAAAATTTGGTGGCTCA
ATTACTTGCGGGATATCCTGGTCTTATAGATGGCTTCAATGACTTTTTGGCACGTTGTGAAAAGAATGAGAGCTTACTTGCTTGTCTTGTGAGTAAAAGT
AATTTTCCCAGACCAGTGAAGGCGGAGGACAGGGATAGAGAGCGCGAGAGGAATGATGTCGCTAAAGAAAGGGATCATGACATGCGAGAGAGAGATAGAC
TTGACAAAAATGTGGCCTTTACTAATAAAGACACTGGAAATCAAAAGATGTCCTTGTATTCTAGCAAGGACAAGCTTTTGTGGAAACCAGTCAATGAACT
TGACCTTTCCAATTGTGACAGCTGCACTCCAAGTTATCGGCTTCTTCCAAAGAATTATCCCATTCCCACAGCGAGCTACAGGACAGAGATTGTTGCTGAA
GTTCTGAATGACCATTGGGTATCCGTCACTTCAGGAAGTGAAGATTACTCTTTTAAGCACATGCGCAAGAACCAGTATGAAGAAAGCCTATTCCGATGCG
AGGATGACAGGTTTGAGTTGGATATGCTGCTGGAATCTGTAAAGGTAACAACTAAGCGTGTAGAGGAGTTGATGGAGAATATCAACAACACTAAGAAAGG
AGACAATCCCGTTCATTATGAGGACTACCTAACAGCTCTAAACCTGAGGTGCATTGAACGGTTATATGGGGATCACGGGCTTGATGTGATGGACGTGTTA
AGGAGGAACACATCTCTTGCTTTGCCCGTGATATTGACTCGCTTGAAGCAGAAGCAAGAAGAGTGGGCTAGGTGCCGCTCAGATTTTAATAAAGTCTGGG
CTGAAATCTATTCCAAGAACTACCACAAATCACTTGACCATCGTAGCTTCTACTTCAAGCAGCAAGATACAAAGAGCCTGAGCACCAAAGCTCTTTTGGC
AGAGATTAAGGAAATTAGTGAGAAGAAGCGAAAAGAAGATGATATGCTTCTAGCATTTGCTGCTGGAAACAGGCATCCAATTATACCAAATTTGGAGTTT
GAGTATCCAGATCCTGATGTCCATGAGGATTTGTATCAGCTCATCAAGTATTCTTGTGGGGAAGTTTGTACAACGGACCAGTTAGACAAGGCCATGAAAA
TTTGGACAACTTTCTTAGAACCAATGCTCGGCGTTCCTTCTCGGCCTAACGGTACAGAGGACACAGAAGATGTTGTGAACGCAAAGAACTATTCTTCCAA
GCATGGAGATAGCGAGGGAAGTCCAAGCAACTCGAAGCCAGGAAATCCTTCAAGAAATGGAGAAGAAAGCATCCAACCTCAGCAGTCAAGTTCAACCAGG
GTATGGTTGGATAATGGGGTCAAAGAAAATGGCTCTCCGAGCACCAATCATACTGCTCGTAAACTTGATACTTGCTCCAGTGCCCTTAAAAACGATAAGC
CAGACAATGAAGCCAAACAAGCTACTTCCAATGAACGGTTGGCGAATTCGAGTGCGACACAACTTGCTACAGGGGCGGAGATAAGTAATGGAAGAAACAT
CCCTGAATCAGGTGCTGCGGCAGCTTCATCGATGCCTAATAGTGGCACCACTGATGGTGGGTTCGCAATAGGATCCAGTAAAGAAGTATTGCCTTCAACA
GAGGCTGGTGAATTTTCAAGGCCTGTGATATCGACAAATGGGGTGGTTGCAGAGGGTATTAATAAAAGTCACAAGTACGAGGAACCGAATGCGCAGTTTA
AGGTTGAAAGAGAAGAGGGTGAGTTGTCTCCGAATGGAGATTTCGAGGAGGATAATTTTGCAGTTTATGGAGAAGCTGGTTCAGAACCTGCTGCTGCAAA
TAAACCGAAAGACAGTGCCCCAAGCAGACACTATCAAGGAGGAAGAAATGGGGAAGACGAAACCCGTCGAGAGGCGGCAGGAGAAAACGATGCGGACGAC
GAAAACGATGCTGAGGAGAGTGCACAAAGAACGTCGGAAGACAACAGTGAGAACGTCTCTGAGAATGCTGATGTCTCCGGGAGCGAGTCCGGTGATGGTG
AAGAGGAGTGCTCTAGGGAAGAGCATGAAGAAGATGGCGATCATGCTGATAATGATGATAAGGCCGAGAGCGAAGGCGAGGCTGAAGGGATGGCCGATGC
TCACGACGTTGAGGGAGATGGAATGGTTTTACCGTTTTCTGAAAGATTCCTTCTTAACGTGAAACCTCTAGCGAAGCATGTCCCGCCATCGCTGCATGAA
CGAGGAAAGGCTTCGCGCATTTTCTACGGAAATGATTCGTTCTATGTGCTCTTCAGGCTTCACCAAACTCTGTACGAGAGAATAAAATCTGCGAAGATCA
ACTCATCGTCTGCTGACAGGAAATGGAGAACTTCAAACGATACAAGTTCTACTGATCTGTATGCTAGGTTTATGAGTGCACTGTACAACCTACTTGATGG
ATCTTCGGATAACACAAAATTCGAGGATGACTGCCGAGCTATAATCGGGACACAGTCTTATGTCTTATTTACTTTGGATAAGCTTATATATAAGCTAGTC
AAACAGCTCCAAACAGTGGCAACCGACGAGATGGACAGCAAGCTACTTCAGTTATACGCTTACGAAAAATCAAGAAAAGCTGGAAGATTCGTCGACATTG
TGTATCATGAGAACACCCGCGTTCTCCTCCACGACGAGCATATATACAGAATCGAATGTTCCTCGAGCCCAACAAGCTTGTCGATACAGCTTATGGACTT
TGTACATGACAAGCCTGAGGTGACTGCTGTTTCCATGGACCCCAACTTTGCGACGTACCTACACAACGACTTCCTCTCCATCGTTCCCAAGAAGAAAGAG
AAGCCCGGTGTTTTCCTGAAGAGGAACAAACGCAGGTATGCGGTGGGCGATGAATTCCAAACCATGGAAGGATTCCATATTTTCAACGGGTTAGAATGCA
AAATTGCCTGCAGTTCTTCCAAGGGTGGGAAGTCTCAGGTGCCTTCAAGTCCAGAAAGGATCATAGGAAGCAGAGGGAATTGGAGGAAGCGCGTAAAGCT
GGTCTTGCTCCTGCCGAAGTTGATGAAGATTCATATGCCTCGGTATGTGTCCTCAGTGCCTTGGTATGTTAATCCTGAGCCTAGGAAGCCTAGTTTGAAA
CATCAAAGGAATTGGAAGCCGGATCCGATCAATACGAACTCGGGGTACAAAAGAGTTACCAAAGTGTTTCGTGCCGAAAAATACATGAAAGGGGCTTGTC
AGAATTGTGGAGCCACGACACATGATTTGAAATCTTGCATGCAGAGACCTAGGAAAATCGGGGCGAAGTGGACTAACAAGCACATTGCCCCTGACGAGAA
GCAGCTTAATGTCCATGCATGGGAAGCTTTTGGTAAGGGTCAAAATATTTACGTGCAAGCTGCTCCTTCTCAAGCGGAGTTGCTGTATAAGAATTTTAAG
GTGATAAAGGAGAAACTTGAGACGCAGACAAAGGATCAAATGCTTGAGAAGTATGGGAATGCTGCCAGCCTAGACCCGATTCCAGAAGAGCTTTTGCTTG
GTCAGAGTGAACGTCAAGTCGAGTATGATCGTGCTGGTGGACTTATCAAGGGACAGGAAGTTGCGCTTCCGAAAAGCAAACAGAGCGTTCGAAACAGTTA
CTGCACCGGTGCCGCCGGAATCAAGGCAGCCGAGGCTGCATCTGATCTGATGAAAGCAAATATCGCGCGGATGGAAGCTGACGATTATGAGGTACCGGAA
CCTGTGGAGGAGAAGAGGCTTGCGACGTGGGGTTCTGAGGTGCCTGATGATCTGGTGTTGGATGAGAAGCTGCTAGCCGAATCACTGAAGAAGGAAGACC
AGAGGAAGAAAGAAGAGAAGGACGAGAGGAAACGAAAGTATAATGTGAAGTGGAATGCTCAGGTTACTGTTGAGGATATGGAGGCGTATCGGATGAAGAA
AGTGCATCATAATGATCCAATGAAGGATTTCCTTAACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030817 pacid=23155421 polypeptide=Lus10030817 locus=Lus10030817.g ID=Lus10030817.BGIv1.0 annot-version=v1.0
MVGGGAATGGGGGAGAGISAQKLTTNDALAYLKAVKDIFQDKREKYDEFLEVMKDFKAQRIDTAGVIGRVKDLFKGHRDLILGFNTFLPKGYEITLPLED
EQPPQKKPVEFEEAINFVNKIKKRFQGDDHVYKAFLDILNMYRQENKSITEVYQEVAALFKDHHDLLGEFTHFLPDNSATASTMFASSNRNSILRDRSSP
MPSNRQMHVDKKERIHPDRDFSVDRPDPDHDRALMRADKEQRRRVEKEKERREDRDRRERDDRDYEHDASHDYRFPLKGKVGRKLEDSSAECQVVGGDDN
YGMHPVSSSYDDKSAVKSALSLELGYCDKVKEKLVEKTGAYQEFLKCLNLYTKDIIDRKELQNLVAQLLAGYPGLIDGFNDFLARCEKNESLLACLVSKS
NFPRPVKAEDRDRERERNDVAKERDHDMRERDRLDKNVAFTNKDTGNQKMSLYSSKDKLLWKPVNELDLSNCDSCTPSYRLLPKNYPIPTASYRTEIVAE
VLNDHWVSVTSGSEDYSFKHMRKNQYEESLFRCEDDRFELDMLLESVKVTTKRVEELMENINNTKKGDNPVHYEDYLTALNLRCIERLYGDHGLDVMDVL
RRNTSLALPVILTRLKQKQEEWARCRSDFNKVWAEIYSKNYHKSLDHRSFYFKQQDTKSLSTKALLAEIKEISEKKRKEDDMLLAFAAGNRHPIIPNLEF
EYPDPDVHEDLYQLIKYSCGEVCTTDQLDKAMKIWTTFLEPMLGVPSRPNGTEDTEDVVNAKNYSSKHGDSEGSPSNSKPGNPSRNGEESIQPQQSSSTR
VWLDNGVKENGSPSTNHTARKLDTCSSALKNDKPDNEAKQATSNERLANSSATQLATGAEISNGRNIPESGAAAASSMPNSGTTDGGFAIGSSKEVLPST
EAGEFSRPVISTNGVVAEGINKSHKYEEPNAQFKVEREEGELSPNGDFEEDNFAVYGEAGSEPAAANKPKDSAPSRHYQGGRNGEDETRREAAGENDADD
ENDAEESAQRTSEDNSENVSENADVSGSESGDGEEECSREEHEEDGDHADNDDKAESEGEAEGMADAHDVEGDGMVLPFSERFLLNVKPLAKHVPPSLHE
RGKASRIFYGNDSFYVLFRLHQTLYERIKSAKINSSSADRKWRTSNDTSSTDLYARFMSALYNLLDGSSDNTKFEDDCRAIIGTQSYVLFTLDKLIYKLV
KQLQTVATDEMDSKLLQLYAYEKSRKAGRFVDIVYHENTRVLLHDEHIYRIECSSSPTSLSIQLMDFVHDKPEVTAVSMDPNFATYLHNDFLSIVPKKKE
KPGVFLKRNKRRYAVGDEFQTMEGFHIFNGLECKIACSSSKGGKSQVPSSPERIIGSRGNWRKRVKLVLLLPKLMKIHMPRYVSSVPWYVNPEPRKPSLK
HQRNWKPDPINTNSGYKRVTKVFRAEKYMKGACQNCGATTHDLKSCMQRPRKIGAKWTNKHIAPDEKQLNVHAWEAFGKGQNIYVQAAPSQAELLYKNFK
VIKEKLETQTKDQMLEKYGNAASLDPIPEELLLGQSERQVEYDRAGGLIKGQEVALPKSKQSVRNSYCTGAAGIKAAEAASDLMKANIARMEADDYEVPE
PVEEKRLATWGSEVPDDLVLDEKLLAESLKKEDQRKKEEKDERKRKYNVKWNAQVTVEDMEAYRMKKVHHNDPMKDFLN
|
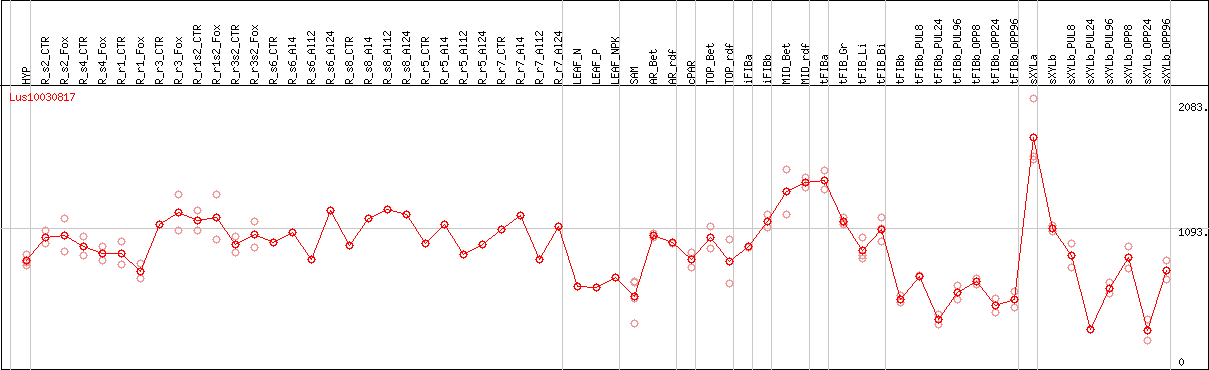
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030817 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
