External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G59900 546 / 0
AT-E1 ALPHA, AT-E1ALPHA
pyruvate dehydrogenase complex E1 alpha subunit (.1)
AT1G24180 526 / 0
IAR4
IAA-CONJUGATE-RESISTANT 4, Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
AT1G01090 219 / 2e-68
PDH-E1 ALPHA, PDH-E1ALPHA
pyruvate dehydrogenase E1 alpha (.1)
AT5G09300 144 / 2e-40
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1), Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.2)
AT1G21400 141 / 1e-38
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
AT5G34780 96 / 2e-22
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013295
591 / 0
AT1G59900 687 / 0.0
pyruvate dehydrogenase complex E1 alpha subunit (.1)
Lus10010728
555 / 0
AT1G59900 687 / 0.0
pyruvate dehydrogenase complex E1 alpha subunit (.1)
Lus10029216
541 / 0
AT1G59900 695 / 0.0
pyruvate dehydrogenase complex E1 alpha subunit (.1)
Lus10002678
219 / 1e-68
AT1G01090 688 / 0.0
pyruvate dehydrogenase E1 alpha (.1)
Lus10018945
149 / 4e-40
AT1G21400 644 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
Lus10028648
146 / 9e-40
AT1G21400 645 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
Lus10020895
132 / 2e-35
AT5G09300 629 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1), Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.2)
Lus10033480
122 / 6e-32
AT5G09300 558 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1), Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.2)
Lus10030196
87 / 7e-21
AT1G01090 271 / 4e-91
pyruvate dehydrogenase E1 alpha (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G192500
529 / 0
AT1G59900 673 / 0.0
pyruvate dehydrogenase complex E1 alpha subunit (.1)
Potri.010G038400
528 / 0
AT1G59900 651 / 0.0
pyruvate dehydrogenase complex E1 alpha subunit (.1)
Potri.002G179500
222 / 1e-69
AT1G01090 672 / 0.0
pyruvate dehydrogenase E1 alpha (.1)
Potri.005G064000
147 / 8e-41
AT5G09300 654 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1), Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.2)
Potri.005G185400
137 / 6e-37
AT5G09300 619 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1), Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.2)
Potri.002G074900
132 / 4e-35
AT1G21400 586 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0254
THDP-binding
PF00676
E1_dh
Dehydrogenase E1 component
Representative CDS sequence
>Lus10030820 pacid=23155180 polypeptide=Lus10030820 locus=Lus10030820.g ID=Lus10030820.BGIv1.0 annot-version=v1.0
ATGGAAGCCGCCATCACCAAGAAGGATTGCATCATCACGGCGTACCGCGACCACTGCACGTTCTTAGGCCGAGGGGGGACGCTCCTCGAGGTGTTCTCCG
AGCTCATGGGGAGGAAAGACGGGTGCTCCAAGGGGAAAGGTGGGTCGATGCATTTTTATAAGAAGGAGGCTGGATTCTACGGCGGTCATGGGATCGTCGG
AGCTCAGGTGCCGTTGGGGTGTGGATTGGCTTTTGCGCAGAAGTATAAGAAGGAGGAGACTGCGACTTTTGCGCTTTATGGAGATGGTGCTGCGAATCAA
GGGCAGTTGTTTGAGGCTCTTAACATCTCGGCTCTGTGGGATCTGCCCATCATCTTGGTTTGTGAGAACAACCATTATGGTATGGGTACTGCTGAATGGC
GGGCAGCTAAGAGCCCTTCGTATTACAAGCGTGGCGATTATGTTCCTGGATTGAAGGTTGATGGTATGGATGTACTTGCTGTTAAACAAGCGTGCAAATT
TGCAAAGGAGCATGTGTTGAAGAATGGACCAATTATTCTCGAAATGGACACGTACAGGTACCACGGTCACTCCATGTCCGACCCCGGTAGCACCTACCGT
ACCCGTGATGAAATCTCTGGCGTCAGACAGGAACGTGATCCAATTGAAAGAGTAAGGAAGTTGATTCTAGCCCATGATCTAGCTACTGAGAAAGAGCTAA
AGGATATCGAGAAGGAAGCAAGGAAAGAAGTAGACGAAGCCATTGCTAAAGCCAAGGAGAGCCCAATGCCAGAGCCATCTGATCTCTTCACAAACGTATA
CTCCAAAGGTCTCGGAACTGAGTCGTTCGGACCTGATAGGAAAGAAGTGAGAGCCACTCTTCCATAG
AA sequence
>Lus10030820 pacid=23155180 polypeptide=Lus10030820 locus=Lus10030820.g ID=Lus10030820.BGIv1.0 annot-version=v1.0
MEAAITKKDCIITAYRDHCTFLGRGGTLLEVFSELMGRKDGCSKGKGGSMHFYKKEAGFYGGHGIVGAQVPLGCGLAFAQKYKKEETATFALYGDGAANQ
GQLFEALNISALWDLPIILVCENNHYGMGTAEWRAAKSPSYYKRGDYVPGLKVDGMDVLAVKQACKFAKEHVLKNGPIILEMDTYRYHGHSMSDPGSTYR
TRDEISGVRQERDPIERVRKLILAHDLATEKELKDIEKEARKEVDEAIAKAKESPMPEPSDLFTNVYSKGLGTESFGPDRKEVRATLP
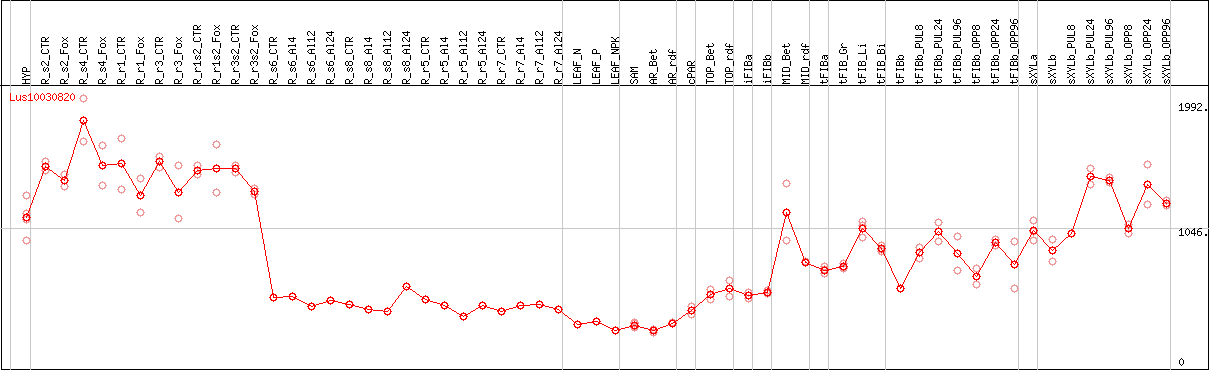
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030820 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.