Lus10030851 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030851 pacid=23155397 polypeptide=Lus10030851 locus=Lus10030851.g ID=Lus10030851.BGIv1.0 annot-version=v1.0
ATGGCATTGCTACACTTCAAGTCCATGATTACGGACGATCCTTTCGGGGCACTGAGTTCGTGGAATGACTCGACCCATTTGTGCCAGTGGTATGGCGTGT
CTTGTTCCGATGGTAGGGTCGCTAACTTAAGTCTCACATCTCAAAGACTCTATGGCTCTATCTCGCCTCACATCGGCAACCTCAGCTTCTTGAAGGGGTT
GCATCTCTACAACAACAGCTTCGTCCGTGAAATCCCTTCCGAAATCGGAAGACTTGGCATGCTGCAAATGTTGGAGCTTCATAACAACACGCTCGGCGGT
GAGATCCCATCCAACATATCGGAATGCTCGGAGCTAACCGTGATACATCTGCAGGACAACAACCTCCAAGGACCACTACCATGGCAGTTTGACCGATTGC
AAAAGCTCGAAATTTTGTTTATATATACAAACAATCTAACAGGTCCCATCCCTCCCTCAATCGGCAACTTGTCGTCTCTCCAACTGTTCGCGCCAGTCCA
GAACAGCTTCAGTGGCACGATTCCCGAGTCTCTAGGTCGGTTGAAGAGCCTCACGGCGCTGTATCTGAACGTCAACGACTTCTCCGGTGAAATCCCCGCT
TCCATTTTCAATCTTTCTTCCCTCACGGACCTGTGGTTCGGAGGGAACTACAAACTCCATGGTGAACTTCCTCCCGACCTAGGGATACTACTTCCGAATC
TTGTGAACTTCGACGTGGGTGGGAACCGTTTCAGCGGTCGTGTCCCGCCTTCCATATCCAATATCTCCAACTTGTTTATGCTCCAAATGTCAGAAAACAA
TTTTACAGGGAGCATGCCTAGCTTGGCAGCCTGTCCCAAGTTCGCGCGCCTTTTGGTACAGAGTAACTCTCTAGGAACCGGGAAAGCTTCCGATTTGGAC
TTCCTGGCATCTTTGGTGAATGCATCGGGTTTGGAAGTAGTGCACATTGAGAAGAACAATTTCGGAGGAATCATACCGGAAAATATTGGCAACTGGTCGA
CCAACCTCTTGCAGCTAACCCTTAACGACAACAAAATATCAGGAAACTTTCCAAACGCGATACAATCACTTGTCAACTTGGAGGTGCTTTGGGCAAACGA
CAATTACCTGACAGGTACAATCCAATCTAGCTTCGGAAACCTGCGATATCTCCGACGGTGTAATTTGTTCAACAACAGAATTTCGGGGAATATTCCGATG
TCTATTGGAAACCTGACGGAATTGATTGAGCTGGATATTAGCGCTAACCGACTGCAGGGAGAGATTCCTGCTTCTATTGGGAATTTGCAGAATTTGTTGT
TGTTCAACCTTTCTTACAATGATTTACATGGTGTCATCCCTCTGCAAGTTATGGGTCTTACTTCCTTGTCGCGAGGCCTTGACTTGTCGTATAATCGATT
CACCGGTCCGATTCCCTCCGAGGTGGGAAATTTGAGGAATTTGGGTTCGTTGGTTCTGTCTCACAACATGTTGTCTGGTATCATTCCTAGTAGTTTGGGT
AGTTGCGTTATGTTGGAGACTCTGAATTTGGGAGAGAATCTCCTACAAGGTTCTATACCTTCTTCTTTGAGTTCGATAAGAGGTATCGAGAAGTTGGATC
TTTCTGGCAACAACTTATCGGGTCGGATACCAATTTTTTTGGAGCGCATGAATATCTCCAAGTTATTGAATTTGTCATATAACAATTTCGATGGCGAGAT
GCTTGCAGAAGGAGTCTTTAGAAATATCACCATCGTTTCAATCGTCGGAAATAACAAGTTATGTGGGGGAATGGCCGAGTTCAAACTTTCACAATGTAAG
AACTTGGTACACAAGTCATTGAGCAGTAAGTGGAAGATTGTTATATCAACAACCTTGAGCCTAATTTTTGTTGCCTTCATGGGCTTCTGCTTCTTGATTT
TTTGGATAAAAATGAGAGGAAAACAAGGTGTTGATTCAGTTGATGATATGCAATTACAACAATTGTCTTACCAAACGCTCCTCAAGGCAACCGATGGCTT
CTCGTCAGCAAACCTAATTGGTGTAGGCAGCTTTGGCTCCGTATATAAGGGAGTCCTTGAGGAAAACGATTTGACTACCAATGTTGCAGTTAAGGTATTC
AACCTTCAACGCCGGGGAGCTTCCAAGAGTTTTATAGCAGAATGTGAAGTTTTGAAGAACATCAAACACATAAATCTTGTCAAGATATTGACTGTGTGTT
CAAGTGTCGATCACAAAGGAGATGATTTCAAAGCTCTCATTTATGAGTTTTTGGCTAATGGAAGCTTGGAGGATTGGTTACATCCTCCGGTTGAAAGAAC
GGATGAGCCTACAAAGAGCTTGAATTTCCTTCAAAGGATAAATCTTGTCATTGATATAGCTTCCGCAATCGATTATCTTCATAACAATTGTGGAACTCCA
ATTGTTCATTGTGATATTAAGCCTAGCAACATACTGCTTGATGAGAATATGAGGGCCAACGTAGGTGATTTTGGGTTGGCAAGATTCATTTCTGCCGTTC
CAAATCCCTCAATGAGTTCCATTGGGATTAAAGGAACAGTTGGCTATGCACCTCCAGAATATGGAATGGGAAACGAGGTGTCAATACAAGGTGACATGTA
CAGTTTCGGCGTCCTCTTACTTGAGTTGTTTACCGGACGAGGGCAATTGACGAGATTTTCACAGAAGGCTCAAACCTTCATAACAATGTTAGAAATGCGT
TATCGAATCAGCAAGCAACAAAAGTTGTGGATCCAATTTTGGTCAACGAACTACGTAGTAGGCTTACAACCCTTCGCAATCATACTAGTGGTAGCACCTT
GGAAGAAACAAGCAATGGCCTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030851 pacid=23155397 polypeptide=Lus10030851 locus=Lus10030851.g ID=Lus10030851.BGIv1.0 annot-version=v1.0
MALLHFKSMITDDPFGALSSWNDSTHLCQWYGVSCSDGRVANLSLTSQRLYGSISPHIGNLSFLKGLHLYNNSFVREIPSEIGRLGMLQMLELHNNTLGG
EIPSNISECSELTVIHLQDNNLQGPLPWQFDRLQKLEILFIYTNNLTGPIPPSIGNLSSLQLFAPVQNSFSGTIPESLGRLKSLTALYLNVNDFSGEIPA
SIFNLSSLTDLWFGGNYKLHGELPPDLGILLPNLVNFDVGGNRFSGRVPPSISNISNLFMLQMSENNFTGSMPSLAACPKFARLLVQSNSLGTGKASDLD
FLASLVNASGLEVVHIEKNNFGGIIPENIGNWSTNLLQLTLNDNKISGNFPNAIQSLVNLEVLWANDNYLTGTIQSSFGNLRYLRRCNLFNNRISGNIPM
SIGNLTELIELDISANRLQGEIPASIGNLQNLLLFNLSYNDLHGVIPLQVMGLTSLSRGLDLSYNRFTGPIPSEVGNLRNLGSLVLSHNMLSGIIPSSLG
SCVMLETLNLGENLLQGSIPSSLSSIRGIEKLDLSGNNLSGRIPIFLERMNISKLLNLSYNNFDGEMLAEGVFRNITIVSIVGNNKLCGGMAEFKLSQCK
NLVHKSLSSKWKIVISTTLSLIFVAFMGFCFLIFWIKMRGKQGVDSVDDMQLQQLSYQTLLKATDGFSSANLIGVGSFGSVYKGVLEENDLTTNVAVKVF
NLQRRGASKSFIAECEVLKNIKHINLVKILTVCSSVDHKGDDFKALIYEFLANGSLEDWLHPPVERTDEPTKSLNFLQRINLVIDIASAIDYLHNNCGTP
IVHCDIKPSNILLDENMRANVGDFGLARFISAVPNPSMSSIGIKGTVGYAPPEYGMGNEVSIQGDMYSFGVLLLELFTGRGQLTRFSQKAQTFITMLEMR
YRISKQQKLWIQFWSTNYVVGLQPFAIILVVAPWKKQAMA
|
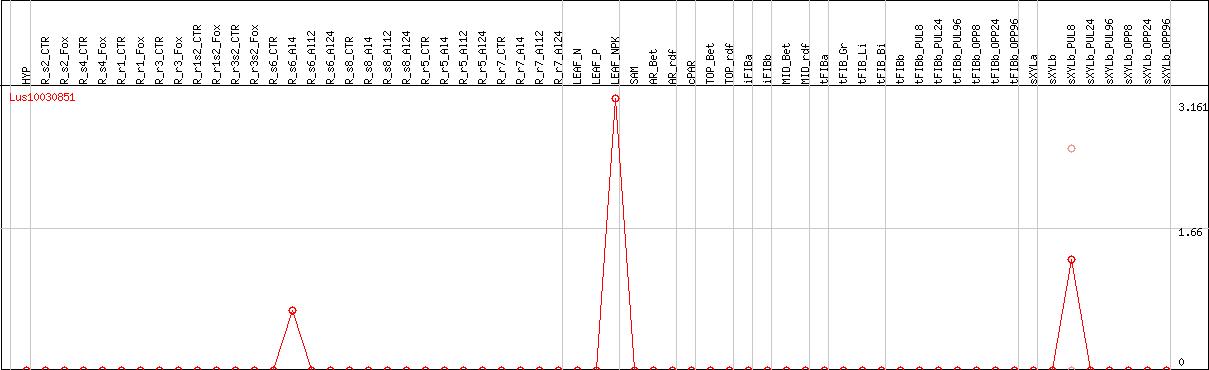
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030851 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
