Lus10030864 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030864 pacid=23155281 polypeptide=Lus10030864 locus=Lus10030864.g ID=Lus10030864.BGIv1.0 annot-version=v1.0
ATGGAGCTGTCGAGAAGTAATTGCAACGTCGAGGAGGAGGTGGCAGAGGGAAGGAGCCGTGGCGCCGGGGCCGTGCCGTTCGAGATCGATCTGAACGAGG
CGCCTGTCGACTGTTTGCCTGTCGAGGATGGCGACGGCTCACGGCCTGATGCGGCGGCGGAGAGCTGCAGTAGGAGAGGGAAACGGCCGCTATTTGACAT
TAATGAGCTGCCGTGCGATGGGGAGAGAGACGGAGAAGCGACCGGCGGTGATTCGGTTACCGATTGGAGATTGTCCGATGAGCGTAGAGCTTCTTCTAAT
TATGCTCAGGCGACGAGTGGTAAGTTATTGTTCGGCCAGAATATCTTGGATAACGTAACAACACCTGCCCGTTTGCCAAGTGTTCTCAGCCCGGGAGGTT
TAGGTAATGTACATTTAGGAATAAACACTTGTGGAAGTCCGCAGAAGATAGATACACCATTTATGGTTATAGAGAAACAATGGCCAAGCCCTGATTTGGT
TCATACAGAAGCAGATGGGAGTCGAAGTTGTGCATCCACTCTTCGAGCATGTCAAGAACCTGGGACAGCGACGATCCAAATGAATTCTTTGGAAATCCCC
TCTCCTCCTATCACGCCATTGGCTTCAAGCATTCCTGTAGATTTAGTTGGTGATGCTCTTCAGTCATGGGAAGTACTGTGGCGGTTCTCAGAAGTTTTGG
AAATGGAAAAATCGATATCTTTTCCGGAATTTGAAGGGGCACTTGCAAATGTTTATCCCATGACTCGTATGAATTCTGGTTCGACAGCTAACAGGGGCAG
TATTACCTCAACGTCTGAAACTATGACAGCGGCTACCGATATGGGACAGTTGTGTAATTCACATATGTGTGCCACTGGGAGTTTGACTCTGGCTCACTGC
TCACTGATGAGAGTTGTATTGGCTGATCTACATGGAAAATTGTGTACTCTCACAGATCCTACTTTTGATAGAAGGAGGAAGAAGAAAGAGGCAGACACTT
TGAATTTGTTTAAGGCAACTATGCTTCATTTCCTGCCAATAAATGAACTTACTTGGCCAGAACTAGCTCGAAGATATTTATTGACTCTCTCATCCATGGA
TGGAAACCTTGAGTCTGTGGAGACAGTTAGTCGAGAGAGCCGCAAAGTTTTTCACTATTTACGAGGTGCTACAGGTGTGCTTGCCAGTTCAATTCCCGGA
TCCGAGGAAGTGGAAGATGAGGCACTGTTGGATGCTGAGGCTGCTAAGAAAATCTTGTGCTCCAATGACTCTGATGAGCTGCCTTGCGAAAAAGTTAACC
CGAGTAGTGAGGAAATTCCTGATTGGGCACAAGTGCTAATACCTGCCAAAACTCTAGCAACGAATGTGGGGGCCAGGATTCGGAAGCTTGTCCACGAGGC
TTTGGCTAGAAATCCTCCGGAATGGGCTAAGAGAGAATTGGAACGGTCCATCAGTAAGATGATTTACAAAGGAAATGCTGCAGGGCCCACTAAGGCCATT
GTTAAATCGGTATTGAGGAGGGTTACTGGTGAAGAAGAGCCAGATAAACCAAATAGAAAGAGCAGAATCAAGTGCACTACTACGATACCTGATTTGATCA
TGAGCCAATGCCAAAAAGTTCTTCACCTTGCCCTTGCGGCTGATAAGGAGAAAGTCTTCTGTGATTTGGTGGGAAGAACCTTTCTCAATGCATCTGATCG
TGATAATGAGGGATTACATGGACATGCTGCTACGGCATCTCATTCTTTGGACTGGAAAAAGATCGATTTTAAATTAACTTCTGGAGCCTATGGGGAATCA
TATGAAGCTTTCACAGAGGATGTTCGAGGGGTTTGGAATCAAATAATTGCTGCTCAGGCTGATCAGTCCCATCGGGTCCAATTGGCCAAGACATTGTCTC
AAAATTTTGAAGCACTTTATGCGCAACAGGTTCTCCCTCTTGTGCACAAGCTGAAGGGGAAAAATAAAATAAAAACCTCAGGAAGCAAGAAAGAATTGGG
AGATTTTCTTGATCATGGAAGTTCGATTACAGAAGGTCATGTCAATGAAGGCATATGCAGAGCATGCGGTATCGACAGGGATGACTGTAATGTTCTTCTG
TGTGATAGATGTGATTCTGGTTACCATACATACTGCCTCAATCCTCCACTTATGCGAATCCCTGCTGGTAATTGGTACTGTCCTCTATGTACTCCTAGTC
AACCCATGGAAGATGATGGTACACAAATTCCTCAATTGGTTAGCAAGGGGATGAAAAAGACCAGTCATGGAGAGTTCATCCATGCATTCATAGAGGAACT
TGGTGATCTGAATACAACCATGGGAGTGAACGATTACTGGGACTACAGCACAAAGGAGAGAGTCCAGCTTCTGAAATTATTGACTGAAGAAGTCGCGAGC
TACAGCACCATCAGGCCCCACCTTGACCAATGTACCATAATCTCTGCGGATCTGCAGCAGAACGTACGATCAGTTTCTGTAGAACTGGGAGAGTTGAAGT
GCAGAGAAAAAATCCTAGCAAACAAGGAGGCAAAAGCAAATACCAATATGTTCAACGGTTTTGGCTGGCCTAGAGCAGGTGGGGAGACTAACTGGCCTGC
ATCTTTTGTGAAAATGATGGGATGCAATGGAAGCACTTCATATTCATCTGACTATTCGGAGAATGTTGCCCTGAGGCACGGAATGGACACATTCAAACAA
CCAGCATTGCATCCGGCTCTCCTTTCAGAGCAATGTCATCCGAGCAGTGGAAGGACATTCCTGCAGATGCCGATTGCTGACACTCTGAACAGCTATATAC
TCGGGCATTCCACTTCCTCATCCTCTCAGAGTGATGCAGAGGAGAATGGTCACTTTAAGGTCGGGGAAAGAGAGCCGAAACCTAAATGGTTCTCTTGTCA
GTCACAAGCAGAGGTGAATGAGCTGATTAGATGGTTGAGATGTTTTGAGCCTATGGACAATGACTTCATAGAGTCCTTACTGCGGCTTCAAAAATTTGAA
TTCGTGGAACCTGCCAATGTTGGGGACATTGGTCAGGGCACATCTCAGTTAACTTTGACGCCCCAAACCACTGAAATGGGTGGCAAGTTTTGTTCTCTTG
CAACGAGAGCTTTGGTCTCATTGGAGAGAAAATATGGTCCTTGCTCAAAGACTGACGGGACCATCATCCCGGTGGACAAGTTGCTGGATGAGGAAGAAAA
GAAGCAGGTTGTAATGTACCGTTGTGGATGCTTAGAGCTGGTATGGACGTCCAGGCATCATTGTCGACTCTGTCACATGACGTATTCCTCCAGCTATGGA
ATCACGGCACATAACAGCGCTGAGTGTAGTTCTGGGACCAGTGCTTCTGAGAGTGCCCAGCTGAAAGGGAAATCTGCAGACGCCAAAGCTGTGCTGGGAG
AACAAGAGGAGTGCTCTGAGGAAGCAACTCCAAACAATTTAGGCACAGGCCCTAGCACTGGCTCTGAGCTAGACGGGGTGAAAAAAGGTTGCACATCCCC
TTACGAACTTAAAGCAGTAAGTGCAAAGTTCGTCACTCAGAGCTCGAACAAGGTGCTGGTGAAGCAGATTGGGCTTCTTGGTTCAGATGGGCTTCCATCA
TTAGTAGCAGCTGCAGCATCACCTTGTCTCAGCGACCCGAGCTTAAAGCTTGCCAGAGGTACGGAGGGCTCAGGGGAATCAAGCTGTTTGAAATCAGGAA
ACCCTCTGTCTCTGACCCATGATTCGTCGCTGAGGCCGGTAACGGGAAAAGAGGCCCAATTTCTCAGACAGCTCAAAATCTGCCTACTGGACATTGATGC
TGGCCTGCACGATGCAGCCCTGAAGCCTTCCAATGCTGAATGTGGAAGAAGGCGTGCCTGGCGGGAGGATATCAAGTCTGCCAAGTCAATAAATGAGATG
GTTCAAGCGACACTGGCACTCGAGAATGCGATCAAAGCCAATTACCTGAGAAGGGAATGGTGGTTCTGGTCGCCACCTTCCGCCTTATTGAGGATCGCTA
CAATCTCCGCTCTGGCACTCAGAATATACACTTTGGACGCAGCAATCCTCTACGAGAAGCCTACTCCTCCAGCGATAAAACCTGCAGTCAACATCGACAG
TCAGAACCCAGATGGAAGCTCATCAGACGACCGTACAATTTCTGCAACTGATCCAGTTCCAAGCAACAGCAACATACCAGCAGCTCAGAGTTCGAAACGC
CAGAGAGTTTGGAAATTGGCAAAACAAAAAGGTGAAGCCAGCACTGCAAAGGGAGCAAACAATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030864 pacid=23155281 polypeptide=Lus10030864 locus=Lus10030864.g ID=Lus10030864.BGIv1.0 annot-version=v1.0
MELSRSNCNVEEEVAEGRSRGAGAVPFEIDLNEAPVDCLPVEDGDGSRPDAAAESCSRRGKRPLFDINELPCDGERDGEATGGDSVTDWRLSDERRASSN
YAQATSGKLLFGQNILDNVTTPARLPSVLSPGGLGNVHLGINTCGSPQKIDTPFMVIEKQWPSPDLVHTEADGSRSCASTLRACQEPGTATIQMNSLEIP
SPPITPLASSIPVDLVGDALQSWEVLWRFSEVLEMEKSISFPEFEGALANVYPMTRMNSGSTANRGSITSTSETMTAATDMGQLCNSHMCATGSLTLAHC
SLMRVVLADLHGKLCTLTDPTFDRRRKKKEADTLNLFKATMLHFLPINELTWPELARRYLLTLSSMDGNLESVETVSRESRKVFHYLRGATGVLASSIPG
SEEVEDEALLDAEAAKKILCSNDSDELPCEKVNPSSEEIPDWAQVLIPAKTLATNVGARIRKLVHEALARNPPEWAKRELERSISKMIYKGNAAGPTKAI
VKSVLRRVTGEEEPDKPNRKSRIKCTTTIPDLIMSQCQKVLHLALAADKEKVFCDLVGRTFLNASDRDNEGLHGHAATASHSLDWKKIDFKLTSGAYGES
YEAFTEDVRGVWNQIIAAQADQSHRVQLAKTLSQNFEALYAQQVLPLVHKLKGKNKIKTSGSKKELGDFLDHGSSITEGHVNEGICRACGIDRDDCNVLL
CDRCDSGYHTYCLNPPLMRIPAGNWYCPLCTPSQPMEDDGTQIPQLVSKGMKKTSHGEFIHAFIEELGDLNTTMGVNDYWDYSTKERVQLLKLLTEEVAS
YSTIRPHLDQCTIISADLQQNVRSVSVELGELKCREKILANKEAKANTNMFNGFGWPRAGGETNWPASFVKMMGCNGSTSYSSDYSENVALRHGMDTFKQ
PALHPALLSEQCHPSSGRTFLQMPIADTLNSYILGHSTSSSSQSDAEENGHFKVGEREPKPKWFSCQSQAEVNELIRWLRCFEPMDNDFIESLLRLQKFE
FVEPANVGDIGQGTSQLTLTPQTTEMGGKFCSLATRALVSLERKYGPCSKTDGTIIPVDKLLDEEEKKQVVMYRCGCLELVWTSRHHCRLCHMTYSSSYG
ITAHNSAECSSGTSASESAQLKGKSADAKAVLGEQEECSEEATPNNLGTGPSTGSELDGVKKGCTSPYELKAVSAKFVTQSSNKVLVKQIGLLGSDGLPS
LVAAAASPCLSDPSLKLARGTEGSGESSCLKSGNPLSLTHDSSLRPVTGKEAQFLRQLKICLLDIDAGLHDAALKPSNAECGRRRAWREDIKSAKSINEM
VQATLALENAIKANYLRREWWFWSPPSALLRIATISALALRIYTLDAAILYEKPTPPAIKPAVNIDSQNPDGSSSDDRTISATDPVPSNSNIPAAQSSKR
QRVWKLAKQKGEASTAKGANN
|
DESeq2's median of ratios [FLAX]
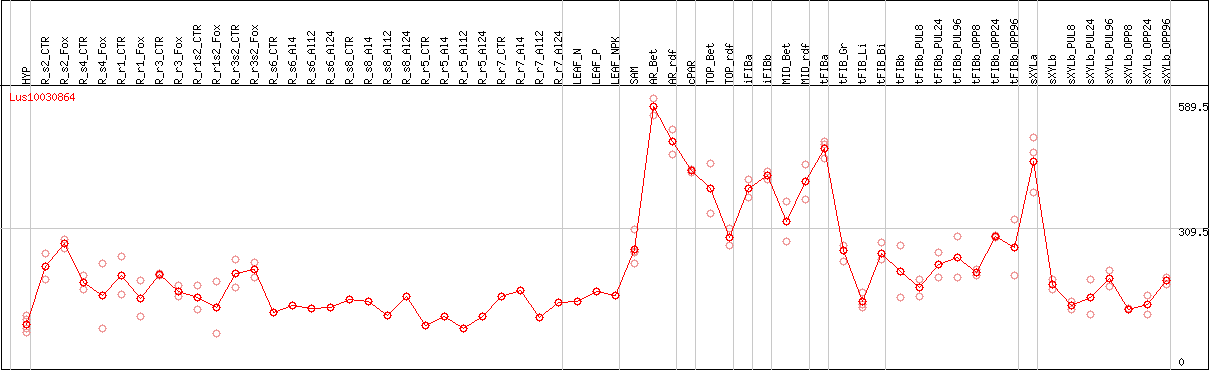
Coexpressed genes
Lus10030864 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
