Lus10030915 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030915 pacid=23155418 polypeptide=Lus10030915 locus=Lus10030915.g ID=Lus10030915.BGIv1.0 annot-version=v1.0
ATGGAGGCGGGGCTTCAGAGTATTAAGGACGTTGTGGAGGAGTTAAAGCGGAATCCCCCTTCTTCTAGCAGTGGAGTTTTGCGGTTTCAGGTTGCTCTGC
CTCCTAGCCCAAAAGCTTTGAATTGGTTTTGTGCCTATCCACCTTTGTCTGGGGTGTTCCCTCGATTTTTCATTTCCAAGGAAACTGCTGATCCGTCGTG
TAAAACTCTCAACCTCAATAGAACACATGGCGTTCTCGGAATTGGGGCCGCAATTTACTTTACTCATTCCTCTTGTAGTTCTCCTGGAGACCGAAGCTTA
ATTAGAAGATATATCTCAAGTGATGCAGTTGGTATGACTGCTTATGGTTTTGAGGACGTCAATTTTCAATCTTCATTCAAGCTTGAAACTGGTTCTGTAT
ACTTTTGCATTCCAGAGATTGAATTAGATGAGCTTGGGGATACTTCTATTCTATCAGCAACACTGGCTTGGAACGACTCTACTTCCAATATAATTAGGCA
GGCAATTCAATCATTGGAGTTATCCTTACGTGAGGCCAATTATCATTTTTGGACAGCCACATGCCAAAGTATAACATCAGCCCTTGGAAAAGCAAATCTG
TCAGAGAACGGAACCTTAAGAATGTTAGATCTGGCCGGTGGAACAAACTACCACTTGCAACATCACCCAAACATCAATGCAGTTTGGGCATCACTTATAG
TAGAAGAATGCACCCGCCTTGGTCTGACGTTCTTTTGTATAGCTCCTGGGTCAAGATCTTCACCACTTGCAATTGCTGCTTCATCTCACAACAAAACCAC
TTGCATTGCTTGCTTTGATGAGCGTGCACTTGCTTTTCATGCTGTCGGTTACTCAAAGGCATCTCAGAAACCAGCAGTCATCATAACATCATCAGGCACT
GCAGTTTCGAATCTTCATCCTGCTGTTGTGGAAGCTAGTCAGGAATTTGTTCCATTACTATTGTTAACTGCTGACCGTCCTCCAGAGCTGCAGGATACTG
GGGCAAATCAAACAATTAATCAAGTGAACCATTTCGGTTCTTTTGTCAGGTTCTTCTTCAGTCTTCCTGCACCCAGTGATGATATTCCTGCTCGGATGGT
ACTGACAACACTCGACTCTGCTGTTAATTGGGCTACTTCTCCACCATATGGTCCGGTTCATATTAACTGTGCATTCAGAGAACCTCTAGATGATAGTCCA
AAAAAATGGATGTTGAGCTGTTTGACAGGATTGGATAGTTGGATGTGTAAAAGTGAACCGTTCACTAAATACATCAAAGAACATTACCGTGTCACCTCAA
ATATTCCCATGTTGACAGAAGTGATTGAGATAATTCAAGGAACTAATAGAGGGCTGCTTCTAATATGTGCTTTGGACGACGAAGACGAGATATGGGCTGC
TCTTATATTGGCAAAACATCTCCATTGGCCAGTTGTAGCTGACATTTTATCTGGCTTGCGTCTAAGAAAAGTTTTGGCAACTTGTCCTGAAGTTCAGGAG
ACTGTTGTATTCCTTGACCATCTAGACCATGTTCTGCTCTCAAAGTCGGTCAAGAGTTCCATTCAATTCAATGTGATTTTGCAGATTGGAACACGGATAA
CGAGCAAGCGTATCTCTCAGATGTTGGAGGAATGTAATCCACACTCTTATATCATAGTTGACAATCATCCATGTCGCCACGATCCATCACATGTTGTCTC
ACACAGAATCCAAAGCTCAATCATTCAGTTTGTTGATAACTTAATGAAAGTGCAGTTTCCTCATGCAACCAGGAACTTTTGTTCTTATTTACAAGCTTTG
AACGAGATGGTGGCATGGGATATATCGTTTCAGATTGCAGCCGAGCGCTCCTTAACTGAGCCTCATGTTGCACAAGTAATTTCAGAAATACTTGATGCAG
AGTTTGTGCTTTTTGTTGGAAATAGCATGCCAATACGTGATTCTGATATGTATGGCCGTAGTTGGAAACATCAAACCTGCACAGCTGCAGACATCCTGTC
TTGCTCTGAACTTCCATGTCTGGGGATTCGGGTTTCTGGAAACAGAGGAGCTAGTGGTATTGATGGTTTGCTTAGCACGGCAACTGGTTTTGCTACTGGA
TGCAATAGACGGGTTCTTTGCGTGATTGGAGATGTTTCTTTCCTTCATGATACAAATGGCTTGTCATTACTGAACTGCAGGACGTTGCGCAAACCAGTGA
CTGTACTTGTGATTAACAATCATGGTGGAGCAATCTTTGGCCTTCTTCCTATTGCAGATAGGACTAGTCCAACAGTTTTGAGTCAGTACTTCTACACCAC
ACACGACATATCCATCCAGAAGTTGTGCCTGGCTCACAGAATCCATTGGGCGGACTTACAGCTCGTATATCTAATGATATTCTGGGACAATGATCTTTCT
AATCTTGTGTCTGATCTTTCCGTGGAGTCAAGTGTCAGACATTTGCTAGTGAAGACAAAAGCAGAACTTCAAGACGCTTTGGTCTCATCTCGGGATGATG
CATCTAATTGTGTAATTGAAGTGGAGAGTGGCATTGATGCCAATACTACTTTCCACAGTAGTTTAAGACAATCTGCCCGTGGAGCAGCAGATAATGCTCT
AAGCAGTCTTTCAAGGCTTTCTGTTCCCAGTTCAGATGGATTCCTTTTCTGCAAGATGGAGTTTTCCTTTTACAGAATTCAACTATGTGCACCAACTACA
GCATCCTCTCTGTGTGATAGAAGTGAGTTCCATAAAGAAGGATATGTCTTGTCTTTGTTTCTGGAAGACGGAACTGTTGGTTATGGTGAGGTTGCACCTC
TCGAAATCCACAAAGAAAGTCTGGCAGATGCAGAAGAACAACTTCGGTTTCTTTGTCATGTTATTGAAGGGGCAAAAATATCATGCACTCTTCCTTTGCT
TAAGGGAACATTTACTTCTTGGTTGTGGAACAGCTTGGGAATCCCGGAATGTTCGATCTTTCCAAGTGTCAGATGTGGTCTGGAAATGGCTATCCTCAAT
GCCATGGCTGCAAGAACAGGTTCCAGCTTGTTAGACATTCTTCTACCTCAGAAAGCAACAGAAGAAATGCCTGGAATGCCAGAACATGTTAGATTATGCG
CCCTCCTTGATTCCAATGGCATGCCCCATGAAGTTGCTTATATTGCTGCATCCCTAGTTGCAGAAGGGTTCACAGCAATAAAGCTTAAAGTGGCTCGGCG
GAAGAATCCAATGGAAGATGCTGCAGTATTGCAGGAAGTGAGAAAGGCGGTTGGTTGGGAAATCCAACTCCGTGTGGATGCTAATAGAAAATGGACCTTT
GAAGAGGCCATTCTCTTTGGTTCTCTTGTAAAACATTCCGACCTACAGTATATTGAGGTGGAATGCGATGCTCAACATTGTTTTGCTGCTTGGATGCCTT
TGCTGGAACCAGTTCAAAACGAAGAAGAAATAATAAAATACTGCAATGAAAGTGGCTTGCCGGTAGCACTTGATGAAACGATCGACAATATCTCTGGAAA
CCCTCTTGATATTCTTGCGAAGTATGCTCATCCAAACATAGTTGCTGTGGTTATTAAACCAGGTGTAGTTGGTGGCTTTGAGAAGGCAGCGCTAATTGCA
CGTTGGGCACAGAAGCAAGGGAAGATGGCCGTTGTGAGTGCTGCTTTTGAAAGTGGACTCTCTTTGTCCACATATGCTGTGTTCTCTTATTACCTGGAGA
TGCACCATGCTGATCTCTACAAGGTTGGCCCCCGCCCGCCGGTAGCTCATGGTCTTGGAACTTACCGATGGCTTAAGCAAGATACAAGCACTCGTCAACT
TCGAGTAAGTCCCGATCCATGCAGTGGGTCAATGGGAGTATTCACGGCAGATGCAATTCAACACTTGCAGCAGTTTCAGCTTGATGACAACATTATCCGC
CGAACGTTTAGTGGAGAGGAAGTCCGCACGTATACCTTGGCTGTGAAAACCAATGAGCTTTCTTCTTGCTCCATCAGAATTCGTGAAGTTGGGCCGACAA
ATTGTTGTCTCGACGGTAAAGTTGTGGTATTTCTGCACGGATTTCTTGGTACCGGTGAAGACTGGATCCCCATCATGAAGGCCATCTCCGGATCTGCAAG
ATGCATTTCAGTTGATCTACCCGGGCACGGGGAGACGAAGATTCAAACACAAGACCGGACTTGGTCAGTCAAAAGAGTTTCCAAGCTACTTTACAAGTTG
ATCGAAGATGTAGCTCCTGGAAGCGTCACACTTGTTGCATATTCAATGGGAGCAAGGATTGCATTACATATGGCATTGCGACATGACGATAAGATTAATG
GAGCCGTCATAATATCCGGTAGCCCTGGGCTAAAGGACGGAATGGGTAGGAAAATTCGGAGGGCTAAAGATGATTCGAGAGCTCGCAATCTCATCGCATA
TGGTTTGCAGACCTTTCTGAGCTCTTGGTACGCTGGGAAACTGTGGAAAAGCTTGAGATGCCATCCTTGTTACAAGGAAGTCGTGGTGAGTCGGTCACGA
TCTGGCGATGCTCGGAGCCTTGCAAAAGCTCTGACCGACCTAAGCATAGGGAGACAAGAACCATTGTGGGATGACTTGAAGCATTGCAAGCTGCCCCTAC
TGTTCGTTGTGGGAGCACAAGATGACAAGTTCAAGGCAATTGCTCGGAAAATGCTCGAAGAAATCGAGACAGAGAGTTGTTGCAAGAGTGAGATAGTCGA
GGTTCCAGTAAGTGGTCATGCTGTCCATTTGGAGAACCCTCTGCATCTCATCCATGCTCTTAGCCAATTTGTTGCCAGAATAAGGAACGATGGTGAAGAC
TAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030915 pacid=23155418 polypeptide=Lus10030915 locus=Lus10030915.g ID=Lus10030915.BGIv1.0 annot-version=v1.0
MEAGLQSIKDVVEELKRNPPSSSSGVLRFQVALPPSPKALNWFCAYPPLSGVFPRFFISKETADPSCKTLNLNRTHGVLGIGAAIYFTHSSCSSPGDRSL
IRRYISSDAVGMTAYGFEDVNFQSSFKLETGSVYFCIPEIELDELGDTSILSATLAWNDSTSNIIRQAIQSLELSLREANYHFWTATCQSITSALGKANL
SENGTLRMLDLAGGTNYHLQHHPNINAVWASLIVEECTRLGLTFFCIAPGSRSSPLAIAASSHNKTTCIACFDERALAFHAVGYSKASQKPAVIITSSGT
AVSNLHPAVVEASQEFVPLLLLTADRPPELQDTGANQTINQVNHFGSFVRFFFSLPAPSDDIPARMVLTTLDSAVNWATSPPYGPVHINCAFREPLDDSP
KKWMLSCLTGLDSWMCKSEPFTKYIKEHYRVTSNIPMLTEVIEIIQGTNRGLLLICALDDEDEIWAALILAKHLHWPVVADILSGLRLRKVLATCPEVQE
TVVFLDHLDHVLLSKSVKSSIQFNVILQIGTRITSKRISQMLEECNPHSYIIVDNHPCRHDPSHVVSHRIQSSIIQFVDNLMKVQFPHATRNFCSYLQAL
NEMVAWDISFQIAAERSLTEPHVAQVISEILDAEFVLFVGNSMPIRDSDMYGRSWKHQTCTAADILSCSELPCLGIRVSGNRGASGIDGLLSTATGFATG
CNRRVLCVIGDVSFLHDTNGLSLLNCRTLRKPVTVLVINNHGGAIFGLLPIADRTSPTVLSQYFYTTHDISIQKLCLAHRIHWADLQLVYLMIFWDNDLS
NLVSDLSVESSVRHLLVKTKAELQDALVSSRDDASNCVIEVESGIDANTTFHSSLRQSARGAADNALSSLSRLSVPSSDGFLFCKMEFSFYRIQLCAPTT
ASSLCDRSEFHKEGYVLSLFLEDGTVGYGEVAPLEIHKESLADAEEQLRFLCHVIEGAKISCTLPLLKGTFTSWLWNSLGIPECSIFPSVRCGLEMAILN
AMAARTGSSLLDILLPQKATEEMPGMPEHVRLCALLDSNGMPHEVAYIAASLVAEGFTAIKLKVARRKNPMEDAAVLQEVRKAVGWEIQLRVDANRKWTF
EEAILFGSLVKHSDLQYIEVECDAQHCFAAWMPLLEPVQNEEEIIKYCNESGLPVALDETIDNISGNPLDILAKYAHPNIVAVVIKPGVVGGFEKAALIA
RWAQKQGKMAVVSAAFESGLSLSTYAVFSYYLEMHHADLYKVGPRPPVAHGLGTYRWLKQDTSTRQLRVSPDPCSGSMGVFTADAIQHLQQFQLDDNIIR
RTFSGEEVRTYTLAVKTNELSSCSIRIREVGPTNCCLDGKVVVFLHGFLGTGEDWIPIMKAISGSARCISVDLPGHGETKIQTQDRTWSVKRVSKLLYKL
IEDVAPGSVTLVAYSMGARIALHMALRHDDKINGAVIISGSPGLKDGMGRKIRRAKDDSRARNLIAYGLQTFLSSWYAGKLWKSLRCHPCYKEVVVSRSR
SGDARSLAKALTDLSIGRQEPLWDDLKHCKLPLLFVVGAQDDKFKAIARKMLEEIETESCCKSEIVEVPVSGHAVHLENPLHLIHALSQFVARIRNDGED
|
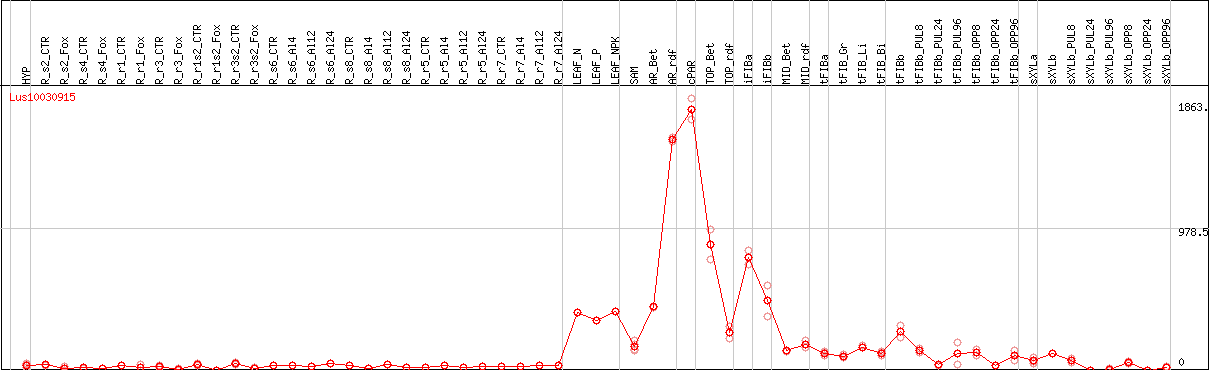
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030915 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
