Lus10030921 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10030921 pacid=23155236 polypeptide=Lus10030921 locus=Lus10030921.g ID=Lus10030921.BGIv1.0 annot-version=v1.0
ATGGCGGCTTTCGTTAATCTGGACGATTCTCCGATGTTCCAGAAAGAGATAGCAGCTATAGGGAAGATGGCAGATGAGTTGAAGGAACGCTGCCAAAGGC
TTTTTAACGGGTCTCAGAAATTTTTGACAGCTATTGGAGAGGCATGCAATGCAGACACAGTACTAGCTGATTCACTAGAAGCATTTGGAGGAGGACAAGA
TGATCCAATCAGTGTATCCATAGGAGGCCCTGTCATGTCCATGTTCGTTAATTCACTTCGCGAGCTAGCGTCCTATAAGGAGCTCCTTCGTTCTCAAGTA
GAGCATGTGCTAGTAGATCGTTTGACACAGTTTGTGGATGTTGATTTGCAAGATGCACTGGAGTCTCGGAAGCGACTGGACAAAGCAGTGTATGCTTATG
ATCAGTCACGGGATAAGTATGTATCTTTAAAGAAAAATGCCCGTGGAGACATAGTTGAAGAATTGGAAGAGAATCTACAGAACTCAAAGTCTGCATACGA
GAGAAGCCGCTTTAATCTAGTTAGTGGCCTCATGAATATTGAAGCTAAGAAGAAATACGAATTCTTGGAATCAATGAGTGCAGTTATGGATGCCCATCTG
AGATACTTTAAGCAGGGATATGAGTTGTTCCGTCAAATGGAGCCATTTGTTCACCAGGTACTGACATATGCTCAACAATCAAAAGAACATGCTAGTATTG
AGCAAGATAAACTTGCCAAGAGGATCCAGGAATTTAGGACCCAATCTGAGTTAGACAGCATACAAGCTTCAAGTAATTTAGAACCTTCAACAAGTGGTGA
GAGTTTTGCTGGTCATGGCATGAGCTCCTACAAAAATATAGAAGGAATCGTGCAATCTTCCTCAAATGGAGAGGTTCAGATTCTCAAACAAGGGTATCTA
CTAAAACGTTCCTCTAGCTCAAGAGGAGACTGGAAGAGAAGGGTGTTCAACATCGTTCAACTGCCTCAGCAGATTAAATACGGACTTACGACTTTGCTTC
AGGATAATCTCTCCATCAAAAACTTTCACACTGCAGCAGATAGGATGGATTGGGTAAAAAAAATCAGTGGAGCAATTGCTTCACTCCTTAACTTTCAGCT
CTTCCGACAGTCCAATTCAGGGAAAAGAACTCTCGAAGCCAAAGAAGCTGCTGCTTCTCATAAATCTCAAGAACTAGAAAATTATCAAAATTCTGAAGAT
GATTCAGTTTCTGGAATCCTAAGAGAAATTCCAGGAAATGAGCTTTGTGCTGAATGCAGTGCTCACGAACCTGATTGGGCATCTCTTAATCTTGGCATAT
TATTATGCATCGAGTGCTCTGGTGTTCACCGAAACCTTGGCGTTCATATTTCAAAGGCTCAAGATAGAAAACATGCGGAAACTTGTGGAATCTCTTTCAC
TTTTGTGCGGTCATTAACCTTCGATGTCAAGGTTTGGGAGCCAACAGTCTTAGACTTATTCCGTGCACTGGGAAACACCTATAGTAACTCTTTGTGGGAA
GGACTGCTGCTTCTTGAAAATAGGAGGATAAAAGAATCCAACGTCACCATATCAGTTACTAAACCCGGACCCAAAGATTCCATCTACTCTAAGGAGATGT
ATATTCATGCCAAGTATGTTGAAAAGGCTCTAGTTGTTAGAGACGCAACAGAATCCAGCTCTCTACCGAACAACAGAACCATCTGGCTAGCTGTGAAGAC
GAACAATCTACGAGAAGTATATCGATGCATTGTGATTTCCGACAAAAACATTGTCAACACCATATTCGACGAGATTGCTCCAGTCGATCTACACCACCAG
ATAAACGATCCAGAGGATCATACATCCGACTCCTCTGCAATAGACAAAATATTCTGTGATCCGGAATCATGCCAGCGAATCAAAGATTCCAATGATCCAA
GGAACTGTTTCCAAGGCTGCTCGTTACTCCACCTGGCATGTTACTATGGTAACATAGTCATGCTCGAATTGCTGCTGCAGTTCGGTGCTGATATAAACTG
GAGGGACTTCCATGGGAGGACTCCCTTGCACCATTGCATTGCCAAAGGGGATTATCCATTGGCCAAGTTCCTACTCAGAAGAGGAGCGTCACCGGCGGTG
AAAGATGGCCGGGGTCTAAGTGTGCTGGAAAGGGCAATGGAGATTGGTGCAATATCTGATGAGGAACTATTCATATTGCTAGCTGAAACTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10030921 pacid=23155236 polypeptide=Lus10030921 locus=Lus10030921.g ID=Lus10030921.BGIv1.0 annot-version=v1.0
MAAFVNLDDSPMFQKEIAAIGKMADELKERCQRLFNGSQKFLTAIGEACNADTVLADSLEAFGGGQDDPISVSIGGPVMSMFVNSLRELASYKELLRSQV
EHVLVDRLTQFVDVDLQDALESRKRLDKAVYAYDQSRDKYVSLKKNARGDIVEELEENLQNSKSAYERSRFNLVSGLMNIEAKKKYEFLESMSAVMDAHL
RYFKQGYELFRQMEPFVHQVLTYAQQSKEHASIEQDKLAKRIQEFRTQSELDSIQASSNLEPSTSGESFAGHGMSSYKNIEGIVQSSSNGEVQILKQGYL
LKRSSSSRGDWKRRVFNIVQLPQQIKYGLTTLLQDNLSIKNFHTAADRMDWVKKISGAIASLLNFQLFRQSNSGKRTLEAKEAAASHKSQELENYQNSED
DSVSGILREIPGNELCAECSAHEPDWASLNLGILLCIECSGVHRNLGVHISKAQDRKHAETCGISFTFVRSLTFDVKVWEPTVLDLFRALGNTYSNSLWE
GLLLLENRRIKESNVTISVTKPGPKDSIYSKEMYIHAKYVEKALVVRDATESSSLPNNRTIWLAVKTNNLREVYRCIVISDKNIVNTIFDEIAPVDLHHQ
INDPEDHTSDSSAIDKIFCDPESCQRIKDSNDPRNCFQGCSLLHLACYYGNIVMLELLLQFGADINWRDFHGRTPLHHCIAKGDYPLAKFLLRRGASPAV
KDGRGLSVLERAMEIGAISDEELFILLAET
|
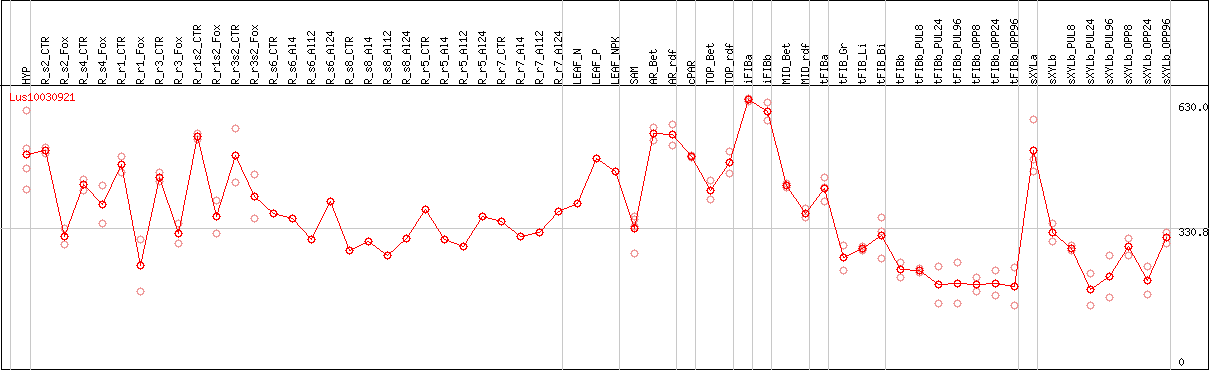
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10030921 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
