Lus10031050 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10031050 pacid=23166741 polypeptide=Lus10031050 locus=Lus10031050.g ID=Lus10031050.BGIv1.0 annot-version=v1.0
ATGGCACCGATCCGGCGCACGGTAGCGCATCTTGACGCTTCCCACCGTCTGTCTACACCACAGCCCTTCTTCCCTGCATCTTATCCTACGCACCTTTCTT
CTGATCACTCTCCTCCACTCCATCTCCTACTCGACGACGCTGCTCTTACTCAATGGATCGCAGGAGAGAAGCTAACGACTCGCTTTCTTACTCCAACCTC
TTCAACCTCGAGGTTTGTGGATTTGTCTTTGATGAACTTCAAAGTCCCACAACCAGATGATGAATTTGATTATTATGGGAATAGTAGTCAGGATGAGAGC
AGAGGTAGCCAAGGTGGATCAATGACAAAATATGGCAATGGGAGTGTGACAGAGAAGGATCCAAGTTCAGGGAAGAGAAGGAAGCTGGCTGTGGACAGTG
ACGGTGAGGAGGAAGATAGGTCTTACGGGGCTGGTATTACCGAGGAACGATACAGATCGATGTTGGGAGAGCACATTAAGAAGTATAAGAGGAGGTTTAG
AGACTCTCCTAGTCCTGCACCTGCTCCTTCTCGAACTGCAGCCATCCCCAAGAATGGTTTGGGAAGTTCGAAAACTAGAAAACTGGGAAACGAGCAGCGA
GGTGTACATGATATGGATGTCACATCCGAATGGAAGAATGATGCCACAATCCCTAAGCGTGGGGACCGCACTCGAGATCTTACTTCCAAAATCTTTTACG
AGCCTGCTTATCTAGATATTGGGGAGGGTATAACTTATAGGATCCCTCCTTCTTATGACAAGATGGCTGCATCCTTGAATTTGCCTAGCTTATCAGATAT
CCGGGTAGATGAAGTTTACTTGGAAGGTACACTTGATTTGGGGTCTTTGGCGCAAATGATGGCCAATGATAAAAGATTTGGTTCTAAGAGCCAAGCAGGA
ATGGGAGATCCTCACCCCCAGTATGAATCTCTGCAAGGGAGGTTGAAGGCATTGGCAGCATCTAGTTCTATGCCAAAGTTTAGTCTTAAGATATCTGAGG
ATGCTTTGAAGTCTGCCATTCCCGAGGGGGCAGCTGGAAATATAGCACGCTCCATTTTGTCAGATGGTGGTGTGTTGCAGGTTTATTATGTAAAAGTTTT
AGAGAAAGGAGACACTTATGAGATTATTGAGCGAAGTTTGCCAAAGAAACCAAAGTTGAAAAAAGATCCCTCTGTGATTGAGAGAGAGGAAATGGAGAGG
ATTGGTAAGGTTTGGGTCAACATTGTAAGAAGAGACATACCCAAGCATCATAGGATATTCACTACTTTTCATCGTAAACAATTAATCGATGCGAAGAGGT
TTTCAGAAAACTGTCAAAGAGAGGTGAAGTTGAAGGTTAGCAGGTCCCTCAAGCTCATGCGGAATGCTGCCCTACGAACTAGGAAGTTGTCCAGAGATAT
GCTGCTATTTTGGAAACGAGTAGATAAGGAGATGGCGGAGGTAAGGAAGAAGGAGGAAAAAGAAGCTGCAGAAGCGTTAAAGCGTGAGCAGGAGCTCCGG
GAAGCAAAGAGGCAGCAGCAACGCCTTAATTTCCTCATACAACAAACCGAGCTTTACAGACATTTCATGCAGGACAAGCAAAGTTCACAACCTTCTGAAC
CCTTGCCTTTGGAAGATGAGAAACTGGATGCCGAACAAATGGGATTAACCTCTTCCGAAGTTCAACCCGGTGAAGAATCTCCAGAGGATGCTGAGCTGAG
AAAGGAAGCTCTGAAGGCTGCTCAAGATGCAGTTTTTAAGCAGAGAAAGTTGACAAGTGCTTTTGATACTGAATGTTCACGGTTGCGGGAGTCAGCTGAT
ACTGAAGTAATGCCAACTGACGACTCTGTTGCAGGAGCTAGCCAAATAGATTTACATAACCCGTCGACCATGCCAGTCACATCAACTGTTCAGACACCAA
AACTCTTCAACGGTAGTCTAAAAGAATATCAGTTGAAGGGTCTGCAGTGGCTGGTTAACTGTTATGAGCAGGGTTTAAATGGTATACTAGCAGACGAGAT
GGGCCTGGGAAAGACTATTCAAGCAATGGCATTCTTAGCTCACCTCGCTGAGGAGAAAAATATATGGGGTCCCTTTTTGGTCGTTGCACCAGCTTCTGTC
TTGAATAACTGGGCTGATGAAATAAGTCGTTTTTGCCCTGAGTTGAAAACCTTACCTTACTGGGGAGGCCTCCAGGAGCGTACTATACTTAGGAAGAACA
TTAATCCCAAGCGTCTTTATCGAAGAGATGCTGGGTTTCATATTCTCATTACAAGCTATCAATTACTTGTGCTGGATGAGAAGTATTTCCGTCGTGTGAA
GTGGCAGTACATGGTACTAGATGAAGCGCAGGCTATTAAAAGCTCCACCAGCATAAGATGGAAGACGCTTCTCAGTTTTAACTGCCGTAACCGCCTACTT
CTGACTGGCACACCTATCCAGAATAGTATGGCAGAATTATGGGCACTATTGCATTTTATTATGCCTACATTATTTGATAGTCACGAACAGTTCAACGAGT
GGTTTTCAAAAGGGATCGAGAATCATGCAGAACACGGGGGTACTGTAAATGAACACCAGCTTAACAGATTGCATGCAATTTTAAAGCCCTTTATGCTACG
TAGACTTAAAGCGGATGTGACTCAAGAGCTCACTAGGAAAACAGAGGTTACAGTTCACTGTAAGTTGAGCTCGCGACAGCAGGCCTTTTATCAAGCTATC
AAGAATAAGATATCTCTTGCGGAGTTGTTTGATGGCTCTCGGGGATATATTAACGAGAAGAAACTTATGAACTTGATGAATATAGTGATTCAGCTCCGGA
AAGTATGTAACCATCCAGAACTGTTTGAGAGGAATGAGGGAAGTTCGTATCTTTATTTTGGACAGCTTCCAAATACTCTTCTACCTCCTCCATTTGGGGA
GTTGGATGACATACACTATCCAGGTGGCCAGAATCCTATTAAGTATAAGATACCAAAGCTGATCAACGACGAGACTGGCCAAAGTTCCAGGGTGCTCCTC
TCGACAGCTGCTCCTGGTTTTTACAGAGAATTATTTCTGAAAAAATTTAATATTTTCTCCCCCGAAAATGTTCAAGCTTCTTTACCCGCAGAGACGAACA
ATTTGCCTGGATTGTTTTCCAGGAGTGGGGCATTTGGCTTTACCCGTCTGATGGACTTGTCCCCCTCGGAAGTTGCTTTTCTCGCAGCTAGCTCGCTTAT
GGAGAAACTGTTGTTTTCTATGATGCGATGGGATCGTCAGTTTCTAGATGGAATCCTCGATGACATTGTAGAAGAGGCGGATGCCAACCTGCTTTGCACT
CAGCTTGAAGCAGGAAAAGTAAGGGCCATTTCACGGATGTTACTAATGCCATCCAGATCTCAGACTAGTGTACTAAGAAGGAAACTTGCAACTGGCCCTA
CTGATGCCCCATTTGAGGCTTTGATAGTCTCTCATCAAGACAGGCTTTTGTCGAATATTGGAGTCCTTCATTCGACATACACTTTCATCCCAAAAAGTCG
AGCTCCACCGATTTCAATCCAATGCTCAGATAGGAATTTCTCTTACAAGATGACTGAAGAACTGCATCAGCCTTGGCTGAAGAGGTTATTAATTGGGTTT
GCGCGGACATCTGAACTTAACGGTCCCAGAAAACCAACAGGGCCACATCCCTTAATAGAAGAGATTGATTCTGAAGTGCCAGTTTCTCAACCTGCCCTTC
AAGTAATACACGAGATTTTTGGATCGTACCCCCCTATGCAGAGCTTTGATCCAGCCAAGCTACTTACGGATTCTGGGAAGCTACAGACCCTTGATATATT
ATTGAAACGCTTGCGTGCGGAGAACCATCGAGTTTTATTGTTTGCACAAATGACGAAAATGTTGAATATTCTTGAGGACTACATGAACTATAGAAAGTAT
AGATATCTAAGACTTGATGGGTCATCCACCATTATGGACCGCCGAGACATGGTCAGAGACTTCCAGGACAGGAATGATATTTTCGTGTTCCTGCTGAGTA
CTAGAGCTGGTGGAGTGGGTATAAATTTGACAGCTGCAGACACGGTTATCTTCTATGAAAATGATTGGAATCCCACGTTAGATTTGCAGGCAATGGATAG
AGCCCATCGGTTGGGTCAGACAAAAGATGTGACTGTCTACCGTCTCATCTGTAAAGAGACGATTGAGGAGAAGATACTGCATCGGGCTAGCCAGAAGAAC
ACAGTCCAGCAACTTGTCATGACTGGTGGCCATGTGCAGGGTGATCTCTTAGCTCCTGAAGATGTTGTGTCATTGCTTCTTGATGATCCACAACTGGAGC
AGAAACTAAAAGAAGTTCCACTGCAGATGAAAGATAATAAACAAAAGAAGAAACCTACAAGGGCCATCGTGATTGATGCTGAAGGTGACGCAACTCTTGA
AGATTTGACTGAACTTGCGACCCAGAATGCGGCAGCAACCGACCCGTCCCAAGATGCTGATAAATCTAAAACAAGTAGCAAAAAGAGGAAAGCTGCAACC
GATAAACCGACTCAATCGAGGCCAAGGAACTCAGCAAGGACGACAGAGGCAGGTTCCGCATCCATTGATAATGACACGGATGGTGGGGTATCTGGTTCCG
ATCCACTATCAGAAAGAGGCAAGAGGCCCAAGAGACCCAAGAAGAGCGTAAATGAAGCCCTTGAGCCGGCATATGTTGCTTCCCCTAGAGTTGATTCAGC
AGCACATACTCTAACCAACAACCTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10031050 pacid=23166741 polypeptide=Lus10031050 locus=Lus10031050.g ID=Lus10031050.BGIv1.0 annot-version=v1.0
MAPIRRTVAHLDASHRLSTPQPFFPASYPTHLSSDHSPPLHLLLDDAALTQWIAGEKLTTRFLTPTSSTSRFVDLSLMNFKVPQPDDEFDYYGNSSQDES
RGSQGGSMTKYGNGSVTEKDPSSGKRRKLAVDSDGEEEDRSYGAGITEERYRSMLGEHIKKYKRRFRDSPSPAPAPSRTAAIPKNGLGSSKTRKLGNEQR
GVHDMDVTSEWKNDATIPKRGDRTRDLTSKIFYEPAYLDIGEGITYRIPPSYDKMAASLNLPSLSDIRVDEVYLEGTLDLGSLAQMMANDKRFGSKSQAG
MGDPHPQYESLQGRLKALAASSSMPKFSLKISEDALKSAIPEGAAGNIARSILSDGGVLQVYYVKVLEKGDTYEIIERSLPKKPKLKKDPSVIEREEMER
IGKVWVNIVRRDIPKHHRIFTTFHRKQLIDAKRFSENCQREVKLKVSRSLKLMRNAALRTRKLSRDMLLFWKRVDKEMAEVRKKEEKEAAEALKREQELR
EAKRQQQRLNFLIQQTELYRHFMQDKQSSQPSEPLPLEDEKLDAEQMGLTSSEVQPGEESPEDAELRKEALKAAQDAVFKQRKLTSAFDTECSRLRESAD
TEVMPTDDSVAGASQIDLHNPSTMPVTSTVQTPKLFNGSLKEYQLKGLQWLVNCYEQGLNGILADEMGLGKTIQAMAFLAHLAEEKNIWGPFLVVAPASV
LNNWADEISRFCPELKTLPYWGGLQERTILRKNINPKRLYRRDAGFHILITSYQLLVLDEKYFRRVKWQYMVLDEAQAIKSSTSIRWKTLLSFNCRNRLL
LTGTPIQNSMAELWALLHFIMPTLFDSHEQFNEWFSKGIENHAEHGGTVNEHQLNRLHAILKPFMLRRLKADVTQELTRKTEVTVHCKLSSRQQAFYQAI
KNKISLAELFDGSRGYINEKKLMNLMNIVIQLRKVCNHPELFERNEGSSYLYFGQLPNTLLPPPFGELDDIHYPGGQNPIKYKIPKLINDETGQSSRVLL
STAAPGFYRELFLKKFNIFSPENVQASLPAETNNLPGLFSRSGAFGFTRLMDLSPSEVAFLAASSLMEKLLFSMMRWDRQFLDGILDDIVEEADANLLCT
QLEAGKVRAISRMLLMPSRSQTSVLRRKLATGPTDAPFEALIVSHQDRLLSNIGVLHSTYTFIPKSRAPPISIQCSDRNFSYKMTEELHQPWLKRLLIGF
ARTSELNGPRKPTGPHPLIEEIDSEVPVSQPALQVIHEIFGSYPPMQSFDPAKLLTDSGKLQTLDILLKRLRAENHRVLLFAQMTKMLNILEDYMNYRKY
RYLRLDGSSTIMDRRDMVRDFQDRNDIFVFLLSTRAGGVGINLTAADTVIFYENDWNPTLDLQAMDRAHRLGQTKDVTVYRLICKETIEEKILHRASQKN
TVQQLVMTGGHVQGDLLAPEDVVSLLLDDPQLEQKLKEVPLQMKDNKQKKKPTRAIVIDAEGDATLEDLTELATQNAAATDPSQDADKSKTSSKKRKAAT
DKPTQSRPRNSARTTEAGSASIDNDTDGGVSGSDPLSERGKRPKRPKKSVNEALEPAYVASPRVDSAAHTLTNNL
|
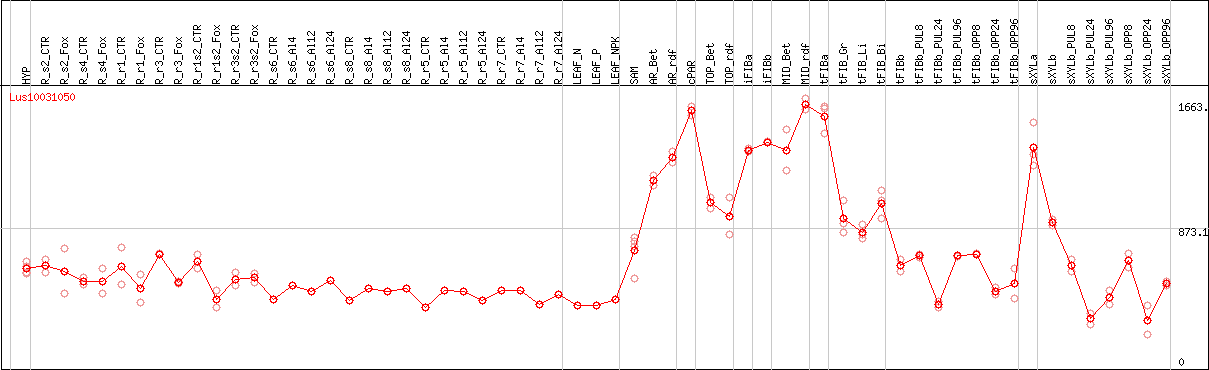
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031050 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
