Lus10031111 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10031111 pacid=23151022 polypeptide=Lus10031111 locus=Lus10031111.g ID=Lus10031111.BGIv1.0 annot-version=v1.0
ATGAGCTTCGCCGCCTGCAAGATGATGCACTGGCCTACCGGCATCGAGTGCTGCGACTCCGGCTTTATCACACATTGCCGCGCCGACTACGTCCCTCCAA
TCTCCGGTATGCAGACCGACGACCTCGACTCCGAGTGGCCTTCCGACGGGAGCAGGAGCATTGGCGCAATCCCTAACCTCGTCGTCACCGCAGGCAACGT
CCTTGAGCTCTACGTAGTTAGGGTGCAGCACGAATCCAATAGGAATTCAGCTGAATTGAAACGTGGCGGTATCGTCGACGGCATTTCCGGTGCTTCTCTC
GAGCTCGTGTGCCATTACAGGTTGCACGGTAACGTTGAATCAATGGCTGTACTGTCTACGGACGGAGGTGATAGTTCGAGGAGAAGAGATGCTATTGTAT
TGGCATTTAAAGATGCTAAGATTTCGGTGTTGGAGTTTGACGATTCGATACATGGGCTCCGTACTAGCTCTATGCATTGTTTTGAAGGTCCAGAATGGTG
CCATTTGAAAAGAGGTAGGGAGTCTTTTGCTCGAGGGCCATTAGTGAAGGTTCATCCACGAGGCAGATGTGGTGGCGTTCTTGTTTATGATTTACAGATG
ATAATACTTAGTGCTGCTGAGGTCGGTTCTGGTTTGATTGGAGATGATGATGCTTTTAGCTCTGGAGGAGCCATATCTGCTAGTATCAAGTCATCTTACA
TTATCAATCTTAGAGACTTGGATGTGAAGCATGTAAAAGACTTTGTATTTATACATGACTATATGGAGCCTGCAGTGGTTATCCTTCATGAGCAAGAGCT
TACTTGGGCTGGTCGCGTAGCATGGAAGCATCACACATGTGTCATATCTGCTTTTAGCATTAGCACAACGTCAAAGCCACCAACCCTGATATTTTCATCT
GTGAATCTCCCGCATGATGCTTACAAGCTACTTGCTGTCCCTCTCCCAATTGGAGGCGTTCTTGTGATCGGCACAAATTCAATACATTATTACAATCAGT
CAGCCTCGTGTACTTTGGCTTTGAACCATTATGCAGTTCCTATTGACAGCAGCCAAGAGCTACCAAGAGCAAGCTTTAATGTTGAACTTGATGCTGCCAA
TGCAACTTGGTTACTCAATGATGTGGCCTTATTGTCGACAAAAACTGGGGAACTACTTTTGCTAACCCTGGTTTATGATGGAAGGGTTGTGCAGAGGCTT
GATCTTTCAAAGTCTAAAGCCTCAGTTCTAACTTCGGACATTGCCATGATTGGAAAATCATTGTTCTTTCTGGCAAGTCGTCTGGGAGACAGCTTGCTTG
TACAATTTACCAGCGGATCAGTGTCCGATGTTTCTCCTGGCTTGAAAGAAGAGGTTGGTGACATTGATGAAGTTCATTCAGCTAAGAGGTTGAAGAGACT
GTCGTCCGATGCTTTACAAGATGGGATGAGCGTTGATGAGCTTTCTTTGTATGGTTCAAACAATACTGAATCAGCACAGAAAACCTTCTCATTTGCAGTG
AGAGATTCATTGGTCAATGTTGGTCCTTTGAAGGACTTCTCCTATGGCTTCCGAATCAATGCTGATGCAAATGCAACTGGAATTGCAAAACAGAGCAACT
ATGAATTGGTCTCCTGTTCAGGTCATGGTAAAAATGGTGCTCTATCTGTTCTTCGGCAGTCAATTCGTCCTGATACGATTACAGAGGTAGAGTTACCTGG
TTGCAAAGGGATCTGGACTGTCTACCATAAAAATTCCCGTGGTCACAATGCAGATTCATCTAAAATAGCAGCCTATGATGATGAATTTCATGCGTATTTA
ATTATAAGTATGGAGGCCCGTACGATGGTTCTCGAAACTGGTGACCTTTTGACAGAGGTTACTGAAAGCGTTGACTACTTTGTTCAAGGACGGACAATTG
CTGCAGGGAATTTGTTTGGAAGGCGTCGAGTTGTTCAGGTCTATGAACGCGGTGCACGAATCCTTGATGGTTCTTTTATGACTCAAGATTTGAGCATCTG
TGCTGCTAACACTGAATTGGGTCCGGGTGTCGAGACCTGTACTGTACTGTCTGTCTCCATTGCTGATCCATATGTGCTAATAAAAATGACAGACGGGAGC
ATAAGACTTCTTGTTGGAGATACTTCTACATGTACCGTTTCTGTGAAAACTCCACCTGCCTTTCAAATCTCGAAGAAGGCAGTATCTGCATGCACCCTGT
ATCATGATAAAGGGCCAGAGCCGTGGATCCGAAAGACAAGTACTGATGCATGGCTTTCAACAGGGATTTCTGAGCCCATAGATGGTACTGATGGTGGCTC
GCATGATCATGGTGACATATATTGCATTGTTTGTTACAAAACCGGTGCCCTAGAGATATTCGATGTACCAAACTTCAGTTCGGTGTTTTATGTGGATAAA
TTTGTTTCTGGGAAAAGCCACCTTGCTGATAACAACCTGCAAGAATCGTCTGATGAGTCACTGGAAGCAACAAGTACAAGTTCTGGTGTAGATGCAGCCC
TTGGTAGCAAAGAGGGCACACATAACCTAAAAGTCGTAGAGTTGGCTATGCAAAGATGGTCTGGGCACCACAGTCGTCCTTTTCTGTTGGGAATATTGAC
TGATGGGACAATACTTTGTTATCATGCTTACCTGTTTGAAGGTCCAGACAGTACCATAAAGCCTGAAGCTTCTGCATCTAGGCTTAGAAATTTGAGATTT
GCACGTGCCCCATTAGATACTTACATGAGGGAAGAAACATCAAGTGGAAGCAATAGTCAAAGAATGACGATTTTCACAAATATTAGTGGTCATCAAGGAT
TTTTCCTATCAGGGTCAAGACCAGCTTGGTTTATGGTTCTACGAGAGCGTTTACGTATTCATCCACAGCTGTGTGATGGTTCTATTGTTGCTTTCACTGT
TCTTCACAATGTTAACTGCAATCATGGCTTCATCTATGTGACATCCCAGGGAATTTTGAAGATTTGCCAACTACCATCTGACTGCAACTATGATAATTAT
TGGCCTGTTCAAAAGGTACCACTGAGAGGAACACCCCATCAGGTCACTTATTTCAACGAGAAAAACTTGTACCCTCTTATAGTTTCCGTTCCTGTTCAAA
AGCCACTCAATCAAGTTCTTTCTTCACTAGTTGATCAAGAAGGCAACCATCAGATCGAGAGTCAGAACATGACTTCAGATGAACTTCATAACACCTACAC
TGTAGAAGAATATGAAGTGCGGATCTTGGAACCTGAGAAATCGGGTGGCCCTTGGCAAACAAAGGCTACAATCCCAATGCATACTTCAGAAAATGCACTT
ACAGTGCGAATGGTCACACTGTTTAACACAACCACGAAGGAGAATGAAACCCTTTTGGCCATTGGGACTGCGTATGTACAAGGGGAGGATGTCGCTGCAA
GAGGACGAATACTTCTGTTTTCTCTCGGAAAGAACACAGAGAACTCTCAGGTGTCGGTCTCTGAAGTTTATTCCAAGGAACTGAAGGGAGCCATATCTGC
TTTAGCCTCACTGCAAGGTCATCTGTTACTAGCTTCTGGCCCAAAAATTATTCTACACAAGTGGACTGGTACAGAACTGAATGGGGTCGCCTTTTTCGAT
GCTCCACCATTATACGTGGTCAGCTTAAACATTGTCAAGAATTTTATCCTCTTGGGTGATATTCACAAGAGTATCTATTTCCTTAGTTGGAAGGAGCAAG
GCGCACAGCTAAGCCTATTAGCCAAGGACTTCAGTTCCCTCGATTGCTTCGCTACTGAGTTCTTAATCGATGGCAGTACATTGAGCCTTGTCGTTTCGGA
CGCACAGAAGAATGTTCAGATATTCTACTACGCTCCAAAAATGTCAGAGAGCTGGAAAGGACAGAAGCTGCTCTCCAGGGCCGAGTTCCACATTGGCTCT
CATGTGACCAAGTTCATGCGGATGCAAATGCTTTCAACAGCATCAGACCGTGGCGGTTCTGCTCCAGTGTCTGACAAAACCAATCGATTCGCCTTACTGT
TTGCAACGCTGGATGGAAGCATTGGCTGCATCACACCTCTGGACGAGATCACTTTCCGTAGGTTACAGTCCGTACAGAAGAAGCTCGTCGAAGCTGTTCC
TCACGTTGCCGGGTTAAACCCCAGATCCTTCCGCCAGTTCCACACAGATGGGAAAGCTCACCGACCTGGCCCCGAAAGCATTGTGGATTGCGAACTGCTA
TCCCATTACGAAATGCTGCCGCTCGAGGAACAGCTCGAGATCGCTCAACAGGTTAGGACAACTCGTGAACAGATGGTAGAGGGAGCAGATGGTCTCGACG
AGCAACGAGAAGAGCTAGCGTATGCAATGGGCAGTACGATAGTCGCCTCCGAGGCTCGTAAACTGTGGAAAGACGAGAATGGGAGTATGATCAAATTTAG
CTCGAGGATTGAGAACTAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10031111 pacid=23151022 polypeptide=Lus10031111 locus=Lus10031111.g ID=Lus10031111.BGIv1.0 annot-version=v1.0
MSFAACKMMHWPTGIECCDSGFITHCRADYVPPISGMQTDDLDSEWPSDGSRSIGAIPNLVVTAGNVLELYVVRVQHESNRNSAELKRGGIVDGISGASL
ELVCHYRLHGNVESMAVLSTDGGDSSRRRDAIVLAFKDAKISVLEFDDSIHGLRTSSMHCFEGPEWCHLKRGRESFARGPLVKVHPRGRCGGVLVYDLQM
IILSAAEVGSGLIGDDDAFSSGGAISASIKSSYIINLRDLDVKHVKDFVFIHDYMEPAVVILHEQELTWAGRVAWKHHTCVISAFSISTTSKPPTLIFSS
VNLPHDAYKLLAVPLPIGGVLVIGTNSIHYYNQSASCTLALNHYAVPIDSSQELPRASFNVELDAANATWLLNDVALLSTKTGELLLLTLVYDGRVVQRL
DLSKSKASVLTSDIAMIGKSLFFLASRLGDSLLVQFTSGSVSDVSPGLKEEVGDIDEVHSAKRLKRLSSDALQDGMSVDELSLYGSNNTESAQKTFSFAV
RDSLVNVGPLKDFSYGFRINADANATGIAKQSNYELVSCSGHGKNGALSVLRQSIRPDTITEVELPGCKGIWTVYHKNSRGHNADSSKIAAYDDEFHAYL
IISMEARTMVLETGDLLTEVTESVDYFVQGRTIAAGNLFGRRRVVQVYERGARILDGSFMTQDLSICAANTELGPGVETCTVLSVSIADPYVLIKMTDGS
IRLLVGDTSTCTVSVKTPPAFQISKKAVSACTLYHDKGPEPWIRKTSTDAWLSTGISEPIDGTDGGSHDHGDIYCIVCYKTGALEIFDVPNFSSVFYVDK
FVSGKSHLADNNLQESSDESLEATSTSSGVDAALGSKEGTHNLKVVELAMQRWSGHHSRPFLLGILTDGTILCYHAYLFEGPDSTIKPEASASRLRNLRF
ARAPLDTYMREETSSGSNSQRMTIFTNISGHQGFFLSGSRPAWFMVLRERLRIHPQLCDGSIVAFTVLHNVNCNHGFIYVTSQGILKICQLPSDCNYDNY
WPVQKVPLRGTPHQVTYFNEKNLYPLIVSVPVQKPLNQVLSSLVDQEGNHQIESQNMTSDELHNTYTVEEYEVRILEPEKSGGPWQTKATIPMHTSENAL
TVRMVTLFNTTTKENETLLAIGTAYVQGEDVAARGRILLFSLGKNTENSQVSVSEVYSKELKGAISALASLQGHLLLASGPKIILHKWTGTELNGVAFFD
APPLYVVSLNIVKNFILLGDIHKSIYFLSWKEQGAQLSLLAKDFSSLDCFATEFLIDGSTLSLVVSDAQKNVQIFYYAPKMSESWKGQKLLSRAEFHIGS
HVTKFMRMQMLSTASDRGGSAPVSDKTNRFALLFATLDGSIGCITPLDEITFRRLQSVQKKLVEAVPHVAGLNPRSFRQFHTDGKAHRPGPESIVDCELL
SHYEMLPLEEQLEIAQQVRTTREQMVEGADGLDEQREELAYAMGSTIVASEARKLWKDENGSMIKFSSRIEN
|
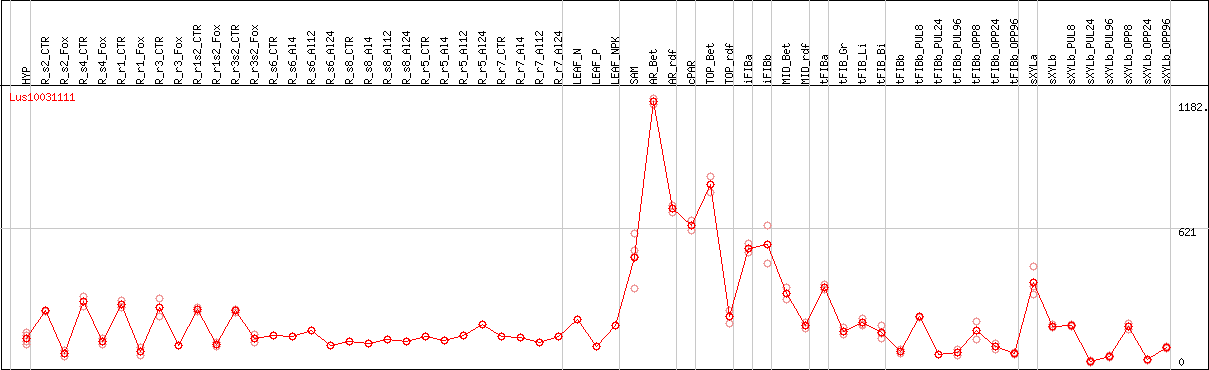
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031111 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
