External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G62350 211 / 1e-69
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT3G47380 196 / 1e-63
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25260 187 / 2e-60
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G12390 164 / 2e-51
PME1
pectin methylesterase inhibitor 1 (.1)
AT1G62770 162 / 1e-50
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G62360 158 / 6e-49
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT2G01610 139 / 2e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G62760 140 / 8e-41
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G14890 135 / 6e-40
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25250 127 / 5e-37
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031713
343 / 7e-122
AT5G62350 202 / 4e-66
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038914
226 / 5e-76
AT5G62350 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10027198
225 / 2e-75
AT5G62350 191 / 3e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10032230
153 / 9e-47
AT1G62770 166 / 9e-52
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038915
149 / 3e-45
AT5G62360 174 / 5e-55
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031138
142 / 2e-42
AT5G62360 172 / 3e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10027199
142 / 2e-42
AT5G62360 169 / 3e-53
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031711
139 / 2e-41
AT5G62360 145 / 4e-44
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10024593
131 / 2e-38
AT1G62770 162 / 2e-50
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G127500
244 / 5e-83
AT5G62350 220 / 2e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128700
243 / 2e-82
AT5G62350 221 / 7e-74
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.003G113700
183 / 1e-58
AT4G12390 178 / 1e-56
pectin methylesterase inhibitor 1 (.1)
Potri.015G128200
176 / 4e-56
AT5G62360 172 / 1e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128100
175 / 1e-55
AT5G62360 221 / 1e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128300
173 / 6e-55
AT5G62360 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128400
152 / 2e-46
AT5G62360 134 / 2e-39
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.002G145800
149 / 2e-45
AT4G00080 164 / 4e-51
unfertilized embryo sac 11, Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127400
147 / 1e-44
AT4G25250 150 / 7e-46
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.001G119300
147 / 2e-44
AT1G62760 167 / 3e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04043
PMEI
Plant invertase/pectin methylesterase inhibitor
Representative CDS sequence
>Lus10031133 pacid=23151155 polypeptide=Lus10031133 locus=Lus10031133.g ID=Lus10031133.BGIv1.0 annot-version=v1.0
ATGGCAAACCCTAAATTTCCCCTTCTCCTAATCTCCATCCTCACCATCACCATCACCCTAGCCTCAGCATCCAACCCGGCGACATCGAGCACCGTCGCAG
CGACAAACTTCATAAAGTCCTCCTGCAGTACCACGACCTACCCAGCACTCTGCGTGCAGTCTCTCCTGTCTTACGCCCCGACCATCCAGCGCAGCCCGCG
TCAGCTGGCGCAGACTGCGTTGGCAGTAAGTCTGGCCAGGGCGCAGTCCACGAGCTCCTTCGTCAGGAAGCTGGCCAGGTTCAAGGGTCTGAGGCCCAGG
GAGGTGGCCGCCATCAAAGACTGCAAGGAAGAGATCGAGGACACAGTTGACCGGCTCAGCAAGTCGATCAAGGAGCTGAAGATGGTCGGGACGGGTGGCG
GGCCAGGGTTCGAGTGGCACGTGAGCAATGTGGTGACGTGGGTCAGCGCGGCGTTGACGAACGAGAACACGTGTGTGGACGGGTTTGGTGGGAAGGCGTT
GAATGGGAGGATGAAGGATGGGATTAGAGGGAGGTTTAGGAACACTGTTCAGGTTACTAGTAATGCTTTGGCTTTGATTAATAAGTACGCTAATAAGCAC
TGA
AA sequence
>Lus10031133 pacid=23151155 polypeptide=Lus10031133 locus=Lus10031133.g ID=Lus10031133.BGIv1.0 annot-version=v1.0
MANPKFPLLLISILTITITLASASNPATSSTVAATNFIKSSCSTTTYPALCVQSLLSYAPTIQRSPRQLAQTALAVSLARAQSTSSFVRKLARFKGLRPR
EVAAIKDCKEEIEDTVDRLSKSIKELKMVGTGGGPGFEWHVSNVVTWVSAALTNENTCVDGFGGKALNGRMKDGIRGRFRNTVQVTSNALALINKYANKH
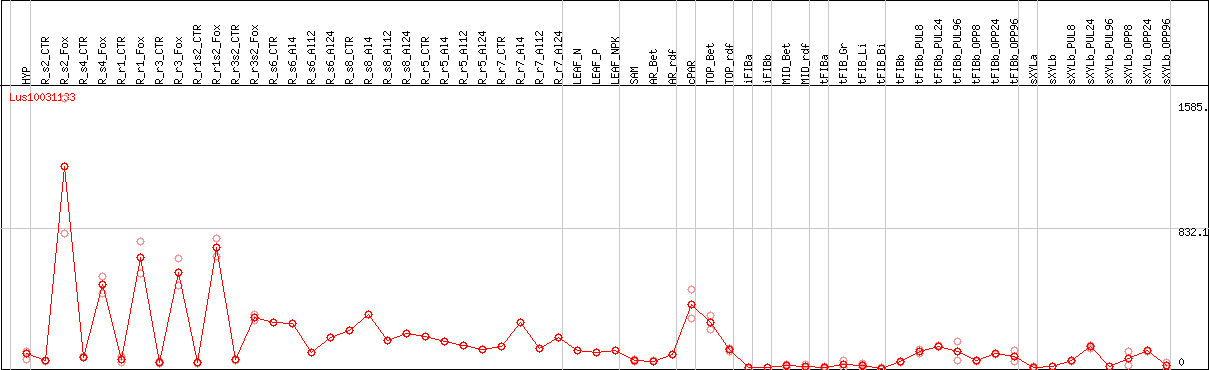
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031133 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.