External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G05010 114 / 2e-29
ACO4, EAT1, EFE
ethylene forming enzyme, ethylene-forming enzyme (.1)
AT2G30830 114 / 2e-29
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G36690 110 / 8e-28
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10490 109 / 1e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G62380 108 / 1e-27
ATACO2, ACO2
ACC oxidase 2 (.1)
AT1G06620 109 / 2e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G77330 108 / 2e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G19000 107 / 1e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT1G06645 106 / 3e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G49630 105 / 3e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024230
211 / 5e-67
AT3G19000 149 / 3e-42
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10023602
203 / 8e-64
AT3G11180 158 / 5e-45
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10000857
122 / 3e-32
AT1G77330 440 / 2e-156
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10028678
113 / 5e-29
AT1G77330 435 / 1e-154
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10022196
113 / 1e-28
AT5G59530 420 / 5e-147
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10022191
112 / 2e-28
AT1G06620 456 / 3e-161
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10022415
111 / 4e-28
AT1G06620 432 / 6e-152
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10022192
109 / 1e-27
AT5G43440 372 / 1e-128
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10021203
108 / 3e-27
AT1G06620 416 / 2e-145
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G069200
201 / 6e-63
AT3G19000 147 / 2e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.002G078600
132 / 2e-36
AT1G77330 455 / 1e-162
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.005G182700
132 / 3e-36
AT1G77330 461 / 4e-165
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.004G003000
115 / 4e-30
AT1G05010 467 / 6e-167
ethylene forming enzyme, ethylene-forming enzyme (.1)
Potri.014G159000
113 / 3e-29
AT1G05010 464 / 8e-166
ethylene forming enzyme, ethylene-forming enzyme (.1)
Potri.002G224100
112 / 4e-29
AT1G05010 440 / 2e-156
ethylene forming enzyme, ethylene-forming enzyme (.1)
Potri.011G020900
110 / 3e-28
AT1G05010 473 / 1e-169
ethylene forming enzyme, ethylene-forming enzyme (.1)
Potri.017G048700
108 / 3e-27
AT2G36690 255 / 3e-82
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.012G006300
108 / 4e-27
AT5G24530 506 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451600
107 / 6e-27
AT4G10500 406 / 5e-142
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10031150 pacid=23151146 polypeptide=Lus10031150 locus=Lus10031150.g ID=Lus10031150.BGIv1.0 annot-version=v1.0
ATGGCCTACAACTACCCGGTGATCGATCTGGCCCCATTCCTCACCAGCTCCGGGGATGCCAAGAAGAAGGCGGAGGTGATTGAGAAGCTGAGGGAAGCTT
GCTCCGACTATGGCTTCTTCGTGGCGGTCAACCACGGCATCCCGGACGAGACCATGGACAAAACCATGGACGTGACCATAGATTTCTTCAACCTCGATTT
GGACGAGAAACTCAAAATCACCACCCCGCCGGGCGCTCCTCTCGCCGCCGGCTACCGTCACAAGGAGACAATGGAGGAAATGTGGTGGAAATTGGATGAA
GCGGCGCAGGTGGTACAGAAGTTGTTGAACGAAGCTCTGAGACTCCCAGAAGGGACGTTGGCCAAGTACAACGACAACAGGGGGACCGACATTCTGTTGG
GATTCCACTACTTGCCGGCGACGGAGACCGAAAAGACGGCGGTCAACGCACACAGGGACAACGGTCTCTTTAGTTTGGTGTTCGAGAATGAAGTTGAAGG
ACTCGAGTTTCTAAAAGACGGCGAGTGGATTCCCATCGATCCCATCCCCCATTCCCTCGTTATCAACGTCGGCGACGCCCTCCAGGCATTGAGCAATGAC
AAGATGAAGAGCCCGACCCACAGGATTTTGGCGCAGCCGGGGAAAAGCCGATATGCATTTGTGTACGGGTACATGGTAGCAGAAGAGAAGTGGGTTGAGC
CGATGCCTGAGTTGATCGGTCAGACCAACGAGTCGCCCAAGTTCAAGAAGTTCAACATCAAAGAGCACATGGCGCTCAAGCAAGTGTTCAAGAACCCTCC
TCCTCACTGGAACGACGTCGTCGGTGTTCATTACCATGCCATCACTCCGTCCGCCGCCGCCGCCGCCAACTGA
AA sequence
>Lus10031150 pacid=23151146 polypeptide=Lus10031150 locus=Lus10031150.g ID=Lus10031150.BGIv1.0 annot-version=v1.0
MAYNYPVIDLAPFLTSSGDAKKKAEVIEKLREACSDYGFFVAVNHGIPDETMDKTMDVTIDFFNLDLDEKLKITTPPGAPLAAGYRHKETMEEMWWKLDE
AAQVVQKLLNEALRLPEGTLAKYNDNRGTDILLGFHYLPATETEKTAVNAHRDNGLFSLVFENEVEGLEFLKDGEWIPIDPIPHSLVINVGDALQALSND
KMKSPTHRILAQPGKSRYAFVYGYMVAEEKWVEPMPELIGQTNESPKFKKFNIKEHMALKQVFKNPPPHWNDVVGVHYHAITPSAAAAAN
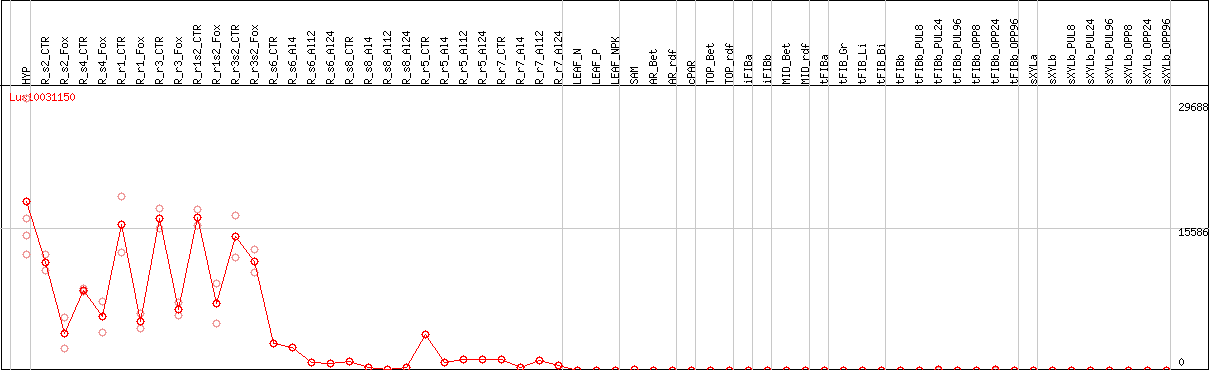
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031150 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.