External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G30690 271 / 1e-88
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT1G72160 219 / 7e-69
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT4G09160 220 / 7e-68
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
AT3G51670 190 / 2e-58
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
AT1G22530 173 / 6e-50
PATL2
PATELLIN 2 (.1)
AT1G72150 166 / 4e-48
PATL1
PATELLIN 1 (.1)
AT1G22180 48 / 1e-06
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT4G08690 46 / 6e-06
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT4G36640 44 / 4e-05
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT1G75170 44 / 6e-05
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031752
473 / 4e-166
AT1G30690 412 / 4e-137
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10038435
277 / 8e-91
AT1G30690 513 / 2e-178
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10009453
218 / 1e-70
AT1G72160 389 / 9e-134
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10001542
219 / 2e-68
AT1G72160 450 / 8e-155
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10001539
210 / 3e-67
AT1G72160 490 / 1e-173
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10003552
206 / 7e-64
AT1G30690 290 / 4e-92
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10012219
185 / 2e-57
AT3G51670 564 / 0.0
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10001269
186 / 8e-57
AT3G51670 602 / 0.0
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10009458
148 / 3e-42
AT4G09160 275 / 2e-87
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G461400
282 / 5e-93
AT1G30690 497 / 4e-172
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Potri.005G224800
223 / 9e-71
AT1G30690 333 / 4e-109
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Potri.019G079500
223 / 3e-69
AT1G72160 439 / 1e-148
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.013G105900
221 / 7e-69
AT1G72160 574 / 0.0
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.T011100
221 / 2e-68
AT1G72160 454 / 6e-155
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.T011101
221 / 2e-68
AT1G72160 452 / 4e-154
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.004G138200
219 / 9e-68
AT1G72160 457 / 2e-156
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.013G106000
219 / 1e-67
AT1G72160 457 / 6e-156
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.019G079300
218 / 1e-67
AT1G72160 545 / 0.0
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.016G127700
188 / 1e-57
AT3G51670 641 / 0.0
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0512
CRAL_TRIO
PF00650
CRAL_TRIO
CRAL/TRIO domain
Representative CDS sequence
>Lus10031175 pacid=23150968 polypeptide=Lus10031175 locus=Lus10031175.g ID=Lus10031175.BGIv1.0 annot-version=v1.0
ATGGAGAAATGTGTGATTGAACAGCTGGATTTCAATCCTGGTGGGACTTGCGAGATTCTTCAGATCAATGATCTCAAGAACACTCCTTTGCCTACTAAGA
AAGAAGTCCGGCTTGCCACCAGGAGAGTTGTTGAGGTTCTTCAGGACAACTACCCTGAATTTGTCTATACCAACATATTTATCAATGTACCATTGTGGTA
CTACGCGTTCAATACAGTTCTGTCCCCTTTCCTAACCCCAAGGACTAAGAACAAGTTCATTACAGCCAGGCCATCCAACGTTACAGAAATTCTACTGAAA
TATATTGCTGCGGAGCAACTTCCAGTAAGGTATGGAGGCCTGAAGATGGACAATGATCCAGATTTCTGTCCCGATGATGTCGCGAAAGAGGTGTGGGTGA
AGTCAGCATCCACCGAAATCATAGAGATTCCTGTGCCAGAGGGAGGCAAAACAATGTTGTGGGACCTCACAGTGTCGAGCTGGGAAGTGAGCTACAAGGA
AGAATTCGTACCGACAAATGAAAAGTCGTACACAATAATAGTTCAGAAGACAAAGAAGATAAATTGGCAGGAAGGAACAATCCGGAATTCATTCGGGAAT
CCAGAGCCGGGGAAGATTCTAGTCACAATTGACAATGCTTCCTTCAGGAAGAAACGCATTCTTTACAGATACAAGGCCAAGAATGCCTCATCTTCATGA
AA sequence
>Lus10031175 pacid=23150968 polypeptide=Lus10031175 locus=Lus10031175.g ID=Lus10031175.BGIv1.0 annot-version=v1.0
MEKCVIEQLDFNPGGTCEILQINDLKNTPLPTKKEVRLATRRVVEVLQDNYPEFVYTNIFINVPLWYYAFNTVLSPFLTPRTKNKFITARPSNVTEILLK
YIAAEQLPVRYGGLKMDNDPDFCPDDVAKEVWVKSASTEIIEIPVPEGGKTMLWDLTVSSWEVSYKEEFVPTNEKSYTIIVQKTKKINWQEGTIRNSFGN
PEPGKILVTIDNASFRKKRILYRYKAKNASSS
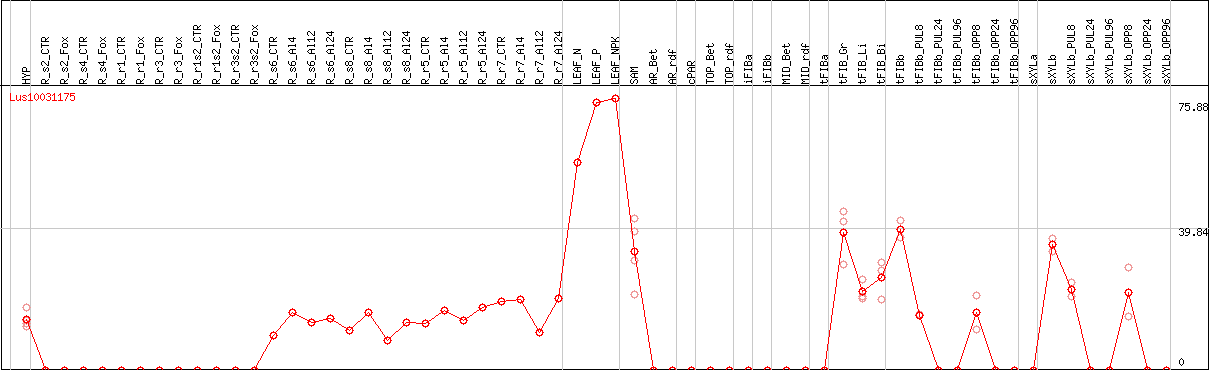
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031175 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.