External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G56070 475 / 2e-163
LOS1, AT1G56075.1
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
AT3G12915 424 / 3e-144
Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
AT3G22980 175 / 3e-49
Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
AT1G06220 144 / 2e-38
GFA1, CLO, MEE5
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
AT5G25230 143 / 4e-38
Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
AT5G13650 96 / 6e-22
SVR3
SUPPRESSOR OF VARIEGATION 3, elongation factor family protein (.1.2)
AT5G39900 96 / 7e-22
Small GTP-binding protein (.1)
AT2G31060 89 / 2e-19
EMB2785
EMBRYO DEFECTIVE 2785, elongation factor family protein (.1.2.3)
AT5G08650 86 / 2e-18
Small GTP-binding protein (.1)
AT1G45332 75 / 6e-15
Translation elongation factor EFG/EF2 protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031201
491 / 1e-169
AT1G56070 1415 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10031763
488 / 2e-169
AT1G56070 1418 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10031779
487 / 8e-169
AT3G12915 760 / 0.0
Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10004749
173 / 2e-48
AT3G22980 1376 / 0.0
Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Lus10043477
143 / 3e-39
AT1G06220 828 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10031129
144 / 2e-38
AT1G06220 1751 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10034111
140 / 3e-37
AT1G06220 1761 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Lus10007250
96 / 4e-22
AT5G39900 1080 / 0.0
Small GTP-binding protein (.1)
Lus10015364
94 / 3e-21
AT5G39900 1082 / 0.0
Small GTP-binding protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G065700
469 / 5e-161
AT1G56070 1623 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.007G065600
468 / 1e-160
AT1G56070 1620 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.005G098100
466 / 9e-160
AT1G56070 1648 / 0.0
LOW EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 1, Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.009G152500
186 / 5e-53
AT3G22980 1521 / 0.0
Ribosomal protein S5/Elongation factor G/III/V family protein (.1)
Potri.018G114900
148 / 6e-40
AT1G06220 1710 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Potri.006G190600
143 / 2e-38
AT1G06220 1744 / 0.0
MATERNAL EFFECT EMBRYO ARREST 5, GAMETOPHYTE FACTOR 1, CLOTHO, Ribosomal protein S5/Elongation factor G/III/V family protein (.1.2)
Potri.017G079800
90 / 4e-20
AT5G39900 999 / 0.0
Small GTP-binding protein (.1)
Potri.006G053000
88 / 3e-19
AT5G13650 1123 / 0.0
SUPPRESSOR OF VARIEGATION 3, elongation factor family protein (.1.2)
Potri.001G305300
86 / 9e-19
AT5G08650 1063 / 0.0
Small GTP-binding protein (.1)
Potri.013G123700
86 / 2e-18
AT2G31060 1084 / 0.0
EMBRYO DEFECTIVE 2785, elongation factor family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00009
GTP_EFTU
Elongation factor Tu GTP binding domain
Representative CDS sequence
>Lus10031185 pacid=23150972 polypeptide=Lus10031185 locus=Lus10031185.g ID=Lus10031185.BGIv1.0 annot-version=v1.0
ATGGTGAAGTTCACAGCTGAGGAGCTTCGTCGCATTATGGACTTCAAGCACAACATCCGTAATATGTCTGTCATTGCCCATGTCGACCATGGGAAATCCA
CACTTACTGATTCTCTAGTGGCTGCTGCGGGTATTATTGCCCAAGAGGTTGCTGGAGATGTGCGGATGACAGATACCCGTGCAGATGAAGCAGAGCGTGG
TATTACAATCAAATCCACTGGAATCTCTCTTTTCTATCAGATGACTGATGAGAGCATCAAGAGTTTCACGGGAAAGAGGGATGGGAGTGAGTACCTTATC
AATCTCATCGATTCCCCTGGACACGTTGACTTCTCATCTGAAGTCACTGCTGCTCTCCGTATCACTGATGGTGCTCTTGTGGTCGTGGATTGTGTTGAGG
GTGTCTGTGTGCAGACTGAGACTGTTCTTCGTCAAGCTTTGGGTGAAAGGATCAGGCCTGTGCTCACTGTGAACAAGATGGACAGGTGTTTCCTTGAGCT
CCAAGTGGATGGAGAGGAAGCCTATCAGACGTTCCAGAGGGTTATTGAGAATGCTAATGTCATCATGGCAACCTATGAAGATCCTCTTCTCGGTGATGTC
CAGGTGTACCCGGAGAAAGGAACTGTTGCCTTTTCCGCTGGTCTGCATGGATGGGCTTTTACGTTGACCAACTTTGCCAAGATGTATGCTTCTAAGTTTG
GAGTGGACGAGGCCAAGATGATGGAAAGGCTGTGGGGTGAGAACGTCTTTGACCCTGCCACTTGTCATGCAAACCTGGCTTCCAGCTGCAGATGCTCTCC
TAGAAATGATGATCTTTCACCTTCCGTCTCCTGCTACTGCCCAGAAGTACCGTGTGGAGAACTTGTATGA
AA sequence
>Lus10031185 pacid=23150972 polypeptide=Lus10031185 locus=Lus10031185.g ID=Lus10031185.BGIv1.0 annot-version=v1.0
MVKFTAEELRRIMDFKHNIRNMSVIAHVDHGKSTLTDSLVAAAGIIAQEVAGDVRMTDTRADEAERGITIKSTGISLFYQMTDESIKSFTGKRDGSEYLI
NLIDSPGHVDFSSEVTAALRITDGALVVVDCVEGVCVQTETVLRQALGERIRPVLTVNKMDRCFLELQVDGEEAYQTFQRVIENANVIMATYEDPLLGDV
QVYPEKGTVAFSAGLHGWAFTLTNFAKMYASKFGVDEAKMMERLWGENVFDPATCHANLASSCRCSPRNDDLSPSVSCYCPEVPCGELV
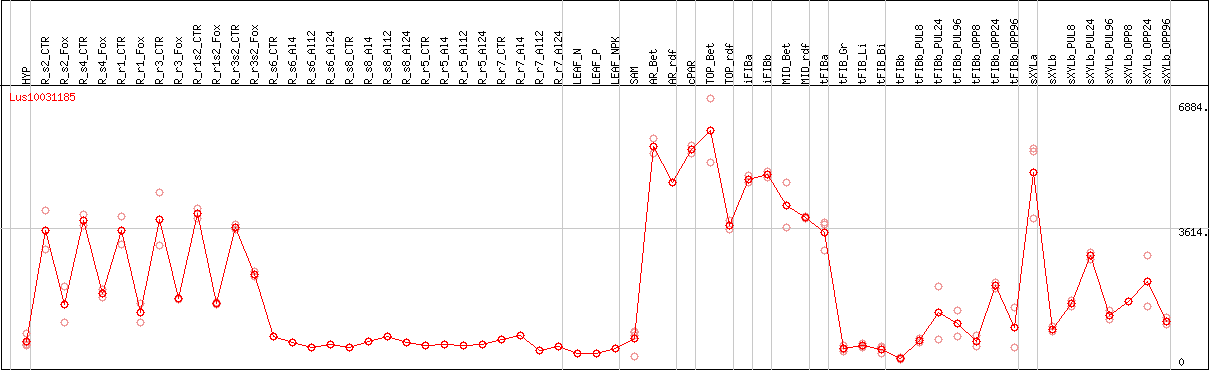
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031185 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.