External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G56090 233 / 6e-76
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G25230 72 / 4e-14
ROF1, ATFKBP62
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
AT4G30480 66 / 4e-12
TPR1, AtTPR1
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
AT5G48570 63 / 6e-11
ROF2, ATFKBP65
FKBP-type peptidyl-prolyl cis-trans isomerase family protein (.1)
AT3G54010 62 / 2e-10
DEI1, PAS1
PASTICCINO 1, FKBP-type peptidyl-prolyl cis-trans isomerase family protein (.1.2)
AT1G62390 61 / 3e-10
CLMP1, Phox2
Phox2, CLUMPED CHLOROPLASTS 1, Octicosapeptide/Phox/Bem1p (PB1) domain-containing protein / tetratricopeptide repeat (TPR)-containing protein (.1)
AT1G58450 57 / 5e-10
TPR6
tetratricopeptide repeat 6, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G17970 59 / 1e-09
ATTOC64-III
translocon at the outer membrane of chloroplasts 64-III (.1)
AT5G09420 54 / 5e-08
MTOM64, ATTOC64-V, AtmtOM64
outer membrane 64, ARABIDOPSIS THALIANA TRANSLOCON AT THE OUTER MEMBRANE OF CHLOROPLASTS 64-V, translocon at the outer membrane of chloroplasts 64-V (.1)
AT3G58620 54 / 6e-08
TTL4
tetratricopetide-repeat thioredoxin-like 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031774
449 / 8e-161
AT1G56090 259 / 2e-86
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10008024
373 / 2e-131
AT1G56090 218 / 8e-71
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10038216
73 / 3e-14
AT3G25230 870 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10025888
72 / 8e-14
AT3G25230 870 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10035117
65 / 6e-12
AT4G30480 291 / 3e-99
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Lus10002338
65 / 2e-11
AT3G25230 874 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10003171
64 / 3e-11
AT3G25230 871 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10031983
64 / 5e-11
AT4G30480 222 / 9e-76
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Lus10039592
63 / 7e-11
AT1G62390 888 / 0.0
Phox2, CLUMPED CHLOROPLASTS 1, Octicosapeptide/Phox/Bem1p (PB1) domain-containing protein / tetratricopeptide repeat (TPR)-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G097300
275 / 5e-92
AT1G56090 260 / 2e-86
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G248300
74 / 1e-14
AT3G25230 857 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Potri.018G100700
71 / 7e-14
AT4G30480 303 / 8e-104
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.014G149400
68 / 1e-12
AT5G48570 741 / 0.0
FKBP-type peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.004G002900
67 / 3e-12
AT1G62390 926 / 0.0
Phox2, CLUMPED CHLOROPLASTS 1, Octicosapeptide/Phox/Bem1p (PB1) domain-containing protein / tetratricopeptide repeat (TPR)-containing protein (.1)
Potri.011G021000
67 / 4e-12
AT1G62390 885 / 0.0
Phox2, CLUMPED CHLOROPLASTS 1, Octicosapeptide/Phox/Bem1p (PB1) domain-containing protein / tetratricopeptide repeat (TPR)-containing protein (.1)
Potri.002G117200
65 / 2e-11
AT5G48570 553 / 0.0
FKBP-type peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.006G178900
63 / 2e-11
AT4G30480 275 / 1e-92
tetratricopeptide repeat 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Potri.001G205300
57 / 5e-09
AT5G09420 724 / 0.0
outer membrane 64, ARABIDOPSIS THALIANA TRANSLOCON AT THE OUTER MEMBRANE OF CHLOROPLASTS 64-V, translocon at the outer membrane of chloroplasts 64-V (.1)
Potri.010G007000
54 / 5e-08
AT3G16760 267 / 6e-84
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF00515
TPR_1
Tetratricopeptide repeat
Representative CDS sequence
>Lus10031195 pacid=23151083 polypeptide=Lus10031195 locus=Lus10031195.g ID=Lus10031195.BGIv1.0 annot-version=v1.0
ATGGCGTCGTCTCCGCAATCTCCAGCGAACAAGATCGAGTCAGCTCACCAGCTCTACAGAGACGGCAAGTACGCCGTCGCGCTTGGATTCTACACGGAAG
CGCTTTCGATGATGGCCAACACCAGCCCTCAGAAGATCTCCCTCCACAGCAACCGAGCTGCTTGCTATCTCAAGCTCCACGATTTCAAGAAGGCGGCTGA
AGAGTGTACATCCGTGCTTGAGCTTGATCAAAACCACGCTGGAGCGTTGATGCTGCGGGCGCAGACATTAGTTACTCTCAAAGACTATCACTCTGCGCTC
TTTGATGTCAATCGGCTGTTGGAGCTGGATCCTGATTCTGAAGTTTACCAGAACCTTGAAACTCGTTTGAGGACACAGTTGTCCCTAGCTTCAATACCTG
AGTCTGAAGCTGAACTGGAAGAAGAGGACGAGAGTGAATCGGAAGACGAACAAGACGGCATAGAGCAGACCAGTAACCTCGAGCCTGATGAAAGTAGGAA
CGAAAGTACTGTCGATGCTGGAGAAGCTTTTGGTGAAGAACAGAGATCTGAGTCCAAGAAAAGCAGCATCGCGGCCGAAGTGATTGCACAAGCACAGAAA
AAGGCGGCCGAGCCAAGGACAACCATTGCATCCGAAGTCATTGCACAGGCACAGAAGGCGAAGCAATCGCCCGAGCAACAACAGTCGGAAGGATGGCAGG
CGATTCCGAAACCAAAGGGACACTCGACGCTGAATTACAGGAGATGGGACAGGGTTGTGGTTGATGACTCTAGCGATGACGACGACGACGATGGAGAAGA
GTGTCGGCCGCAGTATAGGTTCCGTGTTCGAACTGTTGGCGTGCAACCTGTAAAGTGA
AA sequence
>Lus10031195 pacid=23151083 polypeptide=Lus10031195 locus=Lus10031195.g ID=Lus10031195.BGIv1.0 annot-version=v1.0
MASSPQSPANKIESAHQLYRDGKYAVALGFYTEALSMMANTSPQKISLHSNRAACYLKLHDFKKAAEECTSVLELDQNHAGALMLRAQTLVTLKDYHSAL
FDVNRLLELDPDSEVYQNLETRLRTQLSLASIPESEAELEEEDESESEDEQDGIEQTSNLEPDESRNESTVDAGEAFGEEQRSESKKSSIAAEVIAQAQK
KAAEPRTTIASEVIAQAQKAKQSPEQQQSEGWQAIPKPKGHSTLNYRRWDRVVVDDSSDDDDDDGEECRPQYRFRVRTVGVQPVK
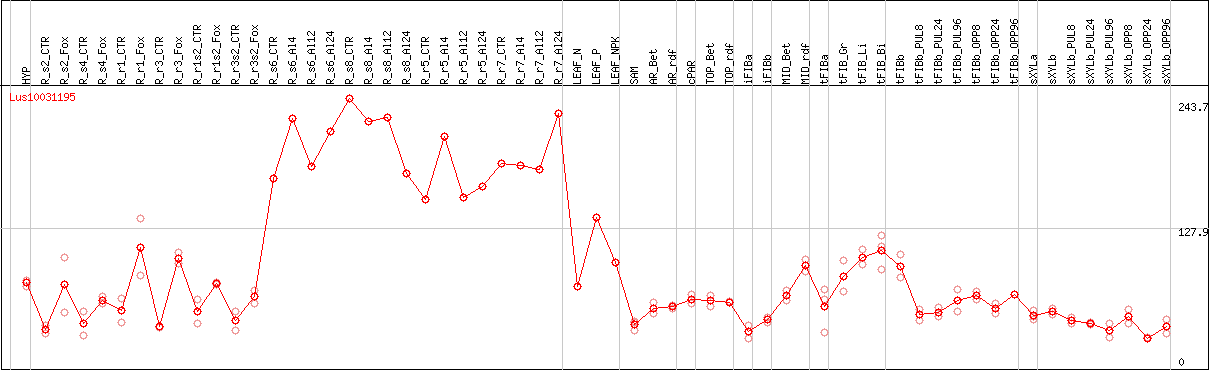
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031195 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.