External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G20820 224 / 6e-73
Leucine-rich repeat (LRR) family protein (.1)
AT5G12940 223 / 2e-72
Leucine-rich repeat (LRR) family protein (.1)
AT3G12610 167 / 1e-50
DRT100
DNA-DAMAGE REPAIR/TOLERATION 100, Leucine-rich repeat (LRR) family protein (.1)
AT3G47570 91 / 2e-21
Leucine-rich repeat protein kinase family protein (.1)
AT3G05660 91 / 3e-21
AtRLP33
receptor like protein 33 (.1)
AT3G12145 88 / 5e-21
FLOR1, FLR1
FLOR1, Leucine-rich repeat (LRR) family protein (.1)
AT4G20140 89 / 2e-20
GSO1
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
AT5G44700 87 / 5e-20
GSO2, EDA23
GASSHO 2, EMBRYO SAC DEVELOPMENT ARREST 23, Leucine-rich repeat transmembrane protein kinase (.1)
AT3G24240 87 / 5e-20
Leucine-rich repeat receptor-like protein kinase family protein (.1)
AT5G07280 87 / 6e-20
EXS, EMS1
EXTRA SPOROGENOUS CELLS, EXCESS MICROSPOROCYTES1, Leucine-rich repeat transmembrane protein kinase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031824
314 / 4e-108
AT3G20820 527 / 0.0
Leucine-rich repeat (LRR) family protein (.1)
Lus10002551
236 / 3e-77
AT3G20820 476 / 1e-168
Leucine-rich repeat (LRR) family protein (.1)
Lus10031377
200 / 1e-63
AT3G20820 433 / 3e-152
Leucine-rich repeat (LRR) family protein (.1)
Lus10010949
197 / 2e-62
AT3G20820 440 / 8e-155
Leucine-rich repeat (LRR) family protein (.1)
Lus10015700
188 / 8e-60
AT3G12610 343 / 3e-118
DNA-DAMAGE REPAIR/TOLERATION 100, Leucine-rich repeat (LRR) family protein (.1)
Lus10037705
189 / 2e-59
AT3G12610 438 / 6e-154
DNA-DAMAGE REPAIR/TOLERATION 100, Leucine-rich repeat (LRR) family protein (.1)
Lus10021028
88 / 1e-20
AT5G06860 362 / 2e-125
polygalacturonase inhibiting protein 1 (.1)
Lus10014996
89 / 2e-20
AT5G07280 1264 / 0.0
EXTRA SPOROGENOUS CELLS, EXCESS MICROSPOROCYTES1, Leucine-rich repeat transmembrane protein kinase (.1)
Lus10033924
88 / 3e-20
AT1G75640 1332 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0022
LRR
PF00560
LRR_1
Leucine Rich Repeat
Representative CDS sequence
>Lus10031254 pacid=23150953 polypeptide=Lus10031254 locus=Lus10031254.g ID=Lus10031254.BGIv1.0 annot-version=v1.0
ATGCACCTCGATCTGCGTAACAACAGGTTCTACGGTCCGATCCCATTCGATTTCGGCCGGCTCAGGATGCTGAGCAGAGCGCTCCTGAGCCGAAACTACC
TATCCGGTTCGATCCCCGATTCGGTTTCCAAGATCTACCGCCTGGCGGATCTGGATCTCTCCCTCAACAGGCTGACCGGTACAATCCCGCTGTCGCTCGG
AAAGATGGCGGTGCTAGCGACGCTGAACCTCGACGGGAACATGATCACCGGGACGATCCCGGCGGATCTCCTGAGCTCGTCGGTGAACAATCTGAACCTG
AGCAGGAACATGCTGGACGGGAAGCTGCCGGAGTTCGGGCCGAGGTCGTACTTCACGGCGCTGGATCTGTCGAGGAACCGCCTGGTCGGAGGGATCCCGG
CGTCGATCGGGAGCGCGTCGTTCATCGGGCATTTGGATTTGAGCCACAACCATCTGTGCGGGAAGATTCCGTTGGGGTCGCCGTTTGACCACCTCGAAGC
GGCGTCGTTTGAGTACAATGACTGCCTTTGCGGCCGGCCGCTTAGACCTTGTTTGACTAAACAGTAA
AA sequence
>Lus10031254 pacid=23150953 polypeptide=Lus10031254 locus=Lus10031254.g ID=Lus10031254.BGIv1.0 annot-version=v1.0
MHLDLRNNRFYGPIPFDFGRLRMLSRALLSRNYLSGSIPDSVSKIYRLADLDLSLNRLTGTIPLSLGKMAVLATLNLDGNMITGTIPADLLSSSVNNLNL
SRNMLDGKLPEFGPRSYFTALDLSRNRLVGGIPASIGSASFIGHLDLSHNHLCGKIPLGSPFDHLEAASFEYNDCLCGRPLRPCLTKQ
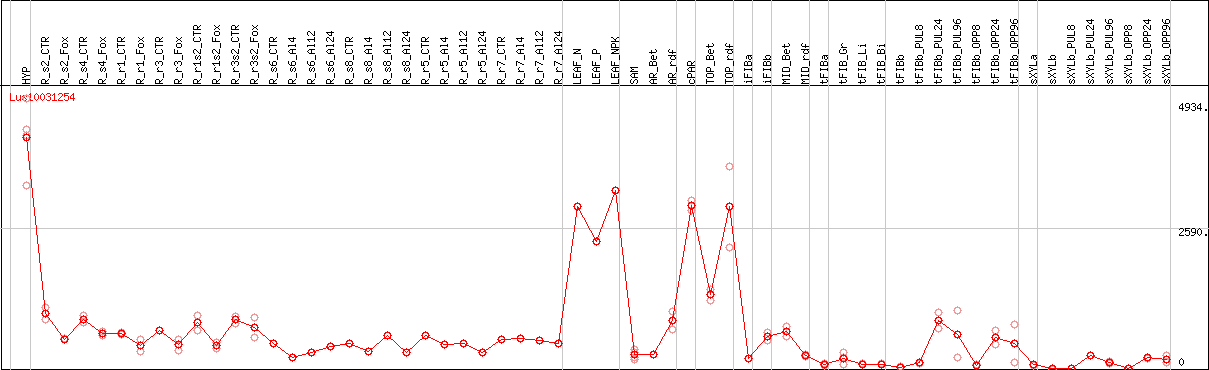
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031254 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.